* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Protein Structure Determined by NMR
Survey
Document related concepts
Safety data sheet wikipedia , lookup
Western blot wikipedia , lookup
Pseudo Jahn–Teller effect wikipedia , lookup
Physical organic chemistry wikipedia , lookup
Chemical thermodynamics wikipedia , lookup
Biosynthesis wikipedia , lookup
Protein adsorption wikipedia , lookup
Spin crossover wikipedia , lookup
Mössbauer spectroscopy wikipedia , lookup
Peptide synthesis wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Atomic nucleus wikipedia , lookup
Nuclear chemistry wikipedia , lookup
Chemical biology wikipedia , lookup
Biochemistry wikipedia , lookup
Metalloprotein wikipedia , lookup
History of molecular biology wikipedia , lookup
Transcript
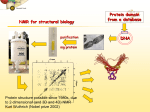
Protein Structure Determination by
NMR Spectroscopy
1D Spectrum
amide/
aromatic/
region
alpha
region
methyl
region
0.015 M glucagon 360 MHz
0.01 M Inhibitor K , 360 MHz
2D J-resolved
COSY 45. COSY 90. COSY
COSY L.R. TOCSY
Relayed COSY. (with one homon
relayed).
NOESY.
ROESY.
Protein Structure Determined by NMR
1D NMR
NOESY
TOCSY
DQF-COSY
Peak Assignment
Distance
Dihedral angle
Constraint
Constraint
Structural
Simulation
Peak Assignment Strategies
• Stage 1 Spin System Identification
• Stage 2 Sequence-Specific Assignment
In the COSY experiment, magnetization is
transferred by scalar coupling. Protons that are more
than three chemical bonds apart give no cross signal
because the 4J coupling constants are close to 0. Only
signals of protons which are two or three bonds apart
are visible in a COSY spectrum (red signals). The
cross signals between HN and Halpha protons are of
special importance because the phi torsion angle of
the protein backbone can be derived from the 3J
coupling constant between them
The TOCSY experiment correlates all protons of
a spin system. Therefore, not only the red signals
are visible (which also appear in a COSY
spectrum) but also additional signals (green)
which originate from the interaction of all protons
of a spin system that are not directly connected via
three chemical bonds.
Thus a characteristic pattern of signals results for
each amino acid from which the amino acid can
be identified
•approximate chemical shifts for various groups in a protein
•“random coil” means “not having any specific structure” e.g. helix,
sheet
•measured in unstructured tetrapeptides
•notice that none of these values is above 8.8 (except Trp sidechain
NH) or below 0.9
•all the amides come between 8 and 8.75
•alphas between 4 and 4.8
•most methyls between 0.9 and 1.4
•some amino acids have identical spin systems and therefore identical
signal patterns. They are: cysteine, aspartic acid, phenylalanine,
histidine, asparagine, tryptophane and tyrosine ('AMX systems') on
the one hand and glutamic acid, glutamine and methionine
('AM(PT)X systems') on the other hand.
Hd
Ha He
Hb
Hg
Hd
Hg
Hb
He
Ha
Sequence-Specific Assignment
the technique of making the spinsystem assignments, followed by
sequence-specific assignment
using unique fragments of
sequence, is known as sequential
assignment (Wuthrich)
main-chain directed assignment (Englander). This
technique does not focus on assigning all the spin
systems first. Rather, it focuses on the backbone and
links sizable stretches of backbone residues via
sequential (i,i+1) nOe’s and other nOe’s that are
characteristic of secondary structures. This technique is
particularly useful when there is some knowledge of
secondary structure beforehand.
Chemical Shift & Protein Secondary Structure
For helical conformation
NH & a-H move upfield from the r.c. value
For b-strand conformation
NH & a-H move downfield from the r.c. value
Wishart, Sykes & Richards, J. Mol. Biol. 1991, 222,311-333
• 化學位移差值(Chemical Shift Index;CSI)
–
以顯現與二級結構的關係,計算方法為:
C.S. difference = d測量-d無序纏捲
NH
αH
βH
α-helix
upfield
upfield
downfield
β-sheet
downfield
downfield
upfield
Temperature Coefficient
• 溫 度 每 上 升 1K ,
backbone NH 質子所感
受到周圍環境的不同,
所產生的化學位移變化,
溫度係數的單位為ppb/K。
• 溫度係數必須與氫氘交
換實驗的結果做比較,
才可以判斷背骨胺基質
子是否有氫鍵存在。
Baxter & Williamson, J. Biomolecular NMR, 1997, 9 ,359-369
Backbone H-D Exchange
2 hour
30 min
gp-41 fusion peptide in SDS micelle, 298 K
Backbone H-D Exchange
pH 5
pH 7
Amide proton resonance region
of the 23-mer fusion peptide of
HIV-gp41 in SDS micelle
Nuclear Overhauser Effect
I - I0
W2 - W0
i s =
=
I0
2W1i W2 W0
Nuclear Overhauser Effect
1
NOE 6 f c
r
Nuclear Overhauser Effect
Brownian motion and NOE
Nuclear Overhauser Effect
Distance & NOE Strength
Strong 1.8 ~ 2.7 Å; Medium 1.8 ~ 3.5 Å; Weak 1.8 ~ 5.0 Å
3J
NH-aH
& dihedral angle
Phase sensitive COSY
3J
NH-aH
& dihedral angle
DQF-COSY
3J
& dihedral angle
NH-aH
3
J HNa = 6.4 cos2 -1.4 cos 1.9