* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Protein Basics
Interactome wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Western blot wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Peptide synthesis wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Genetic code wikipedia , lookup
Structural alignment wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Biosynthesis wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Point mutation wikipedia , lookup
Proteolysis wikipedia , lookup
Protein Chemistry
Basics
• Protein function
• Protein structure
– Primary
• Amino acids
• Linkage
• Protein conformation framework
– Dihedral angles
– Ramachandran plots
• Sequence similarity and variation
Protein Function in Cell
1. Enzymes
•
Catalyze biological reactions
2. Structural role
•
•
•
Cell wall
Cell membrane
Cytoplasm
Protein Structure
Protein Structure
Model Molecule: Hemoglobin
Hemoglobin: Background
• Protein in red blood cells
Red Blood Cell (Erythrocyte)
Hemoglobin: Background
• Protein in red blood cells
• Composed of four subunits, each
containing a heme group: a ring-like
structure with a central iron atom that
binds oxygen
Heme Groups in Hemoglobin
Hemoglobin: Background
• Protein in red blood cells
• Composed of four subunits, each
containing a heme group: a ring-like
structure with a central iron atom that
binds oxygen
• Picks up oxygen in lungs, releases it in
peripheral tissues (e.g. muscles)
Hemoglobin – Quaternary Structure
Two alpha subunits and two beta subunits
(141 AA per alpha, 146 AA per beta)
Hemoglobin – Tertiary Structure
One beta subunit (8 alpha helices)
Hemoglobin – Secondary Structure
alpha helix
β-Hairpin Motif
• Simplest protein motif involving two beta
strands [from Wikipedia]
– adjacent in primary sequence
– antiparallel
– linked by a short loop
• As isolated ribbon or part of beta sheet
• a special case of a turn
– direction of protein backbone reverses
– flanking secondary structure elements
interact (hydrogen bonds)
Xin Zhan
CS 882 course project
14
Types of Turns
• β-turn (most common)
– donor and acceptor residues of hydrogen bonds are separated by 3
residues (i i +3 H-bonding)
• δ-turn
– i i +1 H-bonding
• γ-turn
– i i +2 H-bonding
• α-turn
– i i +4 H-bonding
• π-turn
– i i +5 H-bonding
• ω-loop
– a longer loop with no internal hydrogen bonding
Xin Zhan
CS 882 course project
15
Structure Stabilizing Interactions
• Noncovalent
– Van der Waals forces (transient, weak electrical
attraction of one atom for another)
– Hydrophobic (clustering of nonpolar groups)
– Hydrogen bonding
Hydrogen Bonding
• Involves three atoms:
– Donor electronegative atom (D)
(Nitrogen or Oxygen in proteins)
– Hydrogen bound to donor (H)
– Acceptor electronegative atom (A) in close
proximity
D–H
A
D-H Interaction
• Polarization due to electron withdrawal from
the hydrogen to D giving D partial negative charge
and the H a partial positive charge
• Proximity of the Acceptor A causes further charge
separation δ- δ+
δ-
D–H
A
D-H Interaction
• Polarization due to electron withdrawal from the
hydrogen to D giving D partial negative charge
and the H a partial positive charge
• Proximity of the Acceptor A causes further charge
separation
δ-
• Result:
δ+
δ-
D–H
A
– Closer approach of A to H
– Higher interaction energy than a simple van der Waals
interaction
Hydrogen Bonding
And Secondary Structure
alpha-helix
beta-sheet
Structure Stabilizing Interactions
• Noncovalent
– Van der Waals forces (transient, weak electrical
attraction of one atom for another)
– Hydrophobic (clustering of nonpolar groups)
– Hydrogen bonding
• Covalent
– Disulfide bonds
Disulfide Bonds
• Side chain of cysteine contains highly reactive
thiol group
• Two thiol groups form a disulfide bond
Disulfide Bridge
Disulfide Bonds
• Side chain of cysteine contains highly reactive
thiol group
• Two thiol groups form a disulfide bond
• Contribute to the stability of the folded state by
linking distant parts of the polypeptide chain
Disulfide Bridge –
Linking Distant Amino Acids
Hemoglobin – Primary Structure
NH2-Val-His-Leu-Thr-Pro-Glu-Glu-
Lys-Ser-Ala-Val-Thr-Ala-Leu-TrpGly-Lys-Val-Asn-Val-Asp-Glu-ValGly-Gly-Glu-…..
beta subunit amino acid sequence
Protein Structure - Primary
• Protein: chain of amino acids joined by
peptide bonds
Protein Structure - Primary
• Protein: chain of amino acids joined by
peptide bonds
• Amino Acid
– Central carbon (Cα) attached to:
•
•
•
•
Hydrogen (H)
Amino group (-NH2)
Carboxyl group (-COOH)
Side chain (R)
General Amino Acid Structure
H
H2N
α
C
R
COOH
General Amino Acid Structure
At pH 7.0
H
+H3N
α
C
R
COO-
General Amino Acid Structure
Amino Acids
• Chiral
Chirality: Glyceraldehyde
D-glyderaldehyde
L-glyderaldehyde
Amino Acids
• Chiral
• 20 naturally occuring; distinguishing side
chain
20 Naturally-occurring Amino Acids
Amino Acids
• Chiral
• 20 naturally occuring; distinguishing side
chain
• Classification:
• Non-polar (hydrophobic)
• Charged polar
• Uncharged polar
Alanine:
Nonpolar
Serine:
Uncharged Polar
Aspartic Acid
Charged Polar
Glycine
Nonpolar (special case)
Peptide Bond
• Joins amino acids
Peptide Bond Formation
Peptide Chain
Peptide Bond
• Joins amino acids
• 40% double bond character
– Caused by resonance
Peptide bond
• Joins amino acids
• 40% double bond character
– Caused by resonance
– Results in shorter bond length
Peptide Bond Lengths
Peptide bond
• Joins amino acids
• 40% double bond character
– Caused by resonance
– Results in shorter bond length
– Double bond disallows rotation
Protein Conformation Framework
• Bond rotation determines protein
folding, 3D structure
Bond Rotation Determines
Protein Folding
Protein Conformation Framework
• Bond rotation determines protein
folding, 3D structure
• Torsion angle (dihedral angle) τ
– Measures orientation of four linked
atoms in a molecule: A, B, C, D
Protein Conformation Framework
• Bond rotation determines protein
folding, 3D structure
• Torsion angle (dihedral angle) τ
– Measures orientation of four linked atoms
in a molecule: A, B, C, D
– τABCD defined as the angle between the
normal to the plane of atoms A-B-C and
normal to the plane of atoms B-C-D
Ethane Rotation
A
D
B
C
A
D
B
C
Protein Conformation Framework
• Bond rotation determines protein
folding, 3D structure
• Torsion angle (dihedral angle) τ
– Measures orientation of four linked atoms
in a molecule: A, B, C, D
– τABCD defined as the angle between the
normal to the plane of atoms A-B-C and
normal to the plane of atoms B-C-D
– Three repeating torsion angles along
protein backbone: ω, φ, ψ
Backbone Torsion Angles
Backbone Torsion Angles
• Dihedral angle ω : rotation about the peptide bond,
namely Cα1-{C-N}- Cα2
Backbone Torsion Angles
Backbone Torsion Angles
• Dihedral angle ω : rotation about the peptide bond,
namely Cα1-{C-N}- Cα2
• Dihedral angle φ : rotation about the bond
between N and Cα
Backbone Torsion Angles
Backbone Torsion Angles
• Dihedral angle ω : rotation about the peptide bond,
namely Cα1-{C-N}- Cα2
• Dihedral angle φ : rotation about the bond
between N and Cα
• Dihedral angle ψ : rotation about the bond
between Cα and the carbonyl carbon
Backbone Torsion Angles
Backbone Torsion Angles
• ω angle tends to be planar (0º - cis, or 180 º trans) due to delocalization of carbonyl π electrons
and nitrogen lone pair
Backbone Torsion Angles
• ω angle tends to be planar (0º - cis, or 180 º trans) due to delocalization of carbonyl pi
electrons and nitrogen lone pair
• φ and ψ are flexible, therefore rotation occurs here
Backbone Torsion Angles
Backbone Torsion Angles
• ω angle tends to be planar (0º - cis, or 180 º trans) due to delocalization of carbonyl pi
electrons and nitrogen lone pair
• φ and ψ are flexible, therefore rotation occurs here
• However, φ and ψ of a given amino acid residue
are limited due to steric hindrance
Steric Hindrance
• Interference to rotation caused by spatial
arrangement of atoms within molecule
• Atoms cannot overlap
• Atom size defined by van der Waals radii
• Electron clouds repel each other
Backbone Torsion Angles
• ω angle tends to be planar (0º - cis, or 180 º trans) due to delocalization of carbonyl pi
electrons and nitrogen lone pair
• φ and ψ are flexible, therefore rotation occurs here
• However, φ and ψ of a given amino acid residue
are limited due to steric hindrance
• Only 10% of the {φ, ψ} combinations are
generally observed for proteins
• First noticed by G.N. Ramachandran
G.N. Ramachandran
• Used computer models of small polypeptides to
systematically vary φ and ψ with the objective of finding
stable conformations
• For each conformation, the structure was examined for
close contacts between atoms
• Atoms were treated as hard spheres with dimensions
corresponding to their van der Waals radii
• Therefore, φ and ψ angles which cause spheres to collide
correspond to sterically disallowed conformations of the
polypeptide backbone
Ramachandran Plot
• Plot of φ vs. ψ
• The computed angles which are
sterically allowed fall on certain
regions of plot
Computed Ramachandran Plot
White = sterically
disallowed
conformations (atoms
come closer than sum of
van der Waals radii)
Blue = sterically
allowed conformations
Ramachandran Plot
• Plot of φ vs. ψ
• Computed sterically allowed angles
fall on certain regions of plot
• Experimentally determined angles fall
on same regions
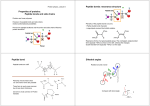
Experimental Ramachandran Plot
φ, ψ distribution in 42 high-resolution
protein structures (x-ray crystallography)
Ramachandran Plot
And Secondary Structure
• Repeating values of φ and ψ along the chain result
in regular structure
• For example, repeating values of φ ~ -57° and ψ ~
-47° give a right-handed helical fold (the alphahelix)
The structure of cytochrome C shows many segments
of helix and the Ramachandran plot shows a tight
grouping of φ, ψ angles near -50,-50
alpha-helix
cytochrome C
Ramachandran plot
Similarly, repetitive values in the region of φ = -110 to
–140 and ψ = +110 to +135 give beta sheets. The
structure of plastocyanin is composed mostly of beta
sheets; the Ramachandran plot shows values in the
–110, +130 region:
beta-sheet
plastocyanin
Ramachandran plot
Ramachandran Plot
And Secondary Structure
• White = sterically disallowed conformations
• Red = sterically allowed regions if strict
(greater) radii are used (namely righthanded alpha helix and beta sheet)
• Yellow = sterically allowed if shorter radii
are used (i.e. atoms allowed closer together;
brings out left-handed helix)
Sample Ramachandran Plot
Alanine Ramachandran Plot
Arginine Ramachandran Plot
Glutamine Ramachandran Plot
Glycine Ramachandran Plot
Note more allowed regions due to less steric hindrance - Turns
Proline Ramachandran Plot
Note less allowed regions due to structure rigidity
φ, ψ and Secondary Structure
Name
φ
ψ
Structure
------------------- ------- ------- --------------------------------alpha-L
57
47
left-handed alpha helix
3-10 Helix
-49 -26
right-handed.
π helix
-57 -80
right-handed.
Type II helices -79 150
left-handed helices
formed by polyglycine
and polyproline.
Collagen
-51 153 right-handed coil formed
of three left handed
helicies.
Sequence Similarity
• Sequence similarity implies structural,
functional, and evolutionary commonality
Homologous Proteins:
Enterotoxin and Cholera toxin
Enterotoxin
Cholera toxin
80% homology
Sequence Similarity
• Sequence similarity implies structural,
functional, and evolutionary commonality
• Low sequence similarity implies little
structural similarity
Nonhomologous Proteins:
Cytochrome and Barstar
Cytochrome
Barstar
Less than 20% homology
Sequence Similarity
• Sequence similarity implies structural,
functional, and evolutionary commonality
• Low sequence similarity implies little
structural similarity
• Small mutations generally well-tolerated by
native structure – with exceptions!
Sequence Similarity Exception
• Sickle-cell anemia resulting from one residue
change in hemoglobin protein
• Replace highly polar (hydrophilic) glutamate
with nonpolar (hydrophobic) valine
Sickle-cell mutation in
hemoglobin sequence
Normal Trait
• Hemoglobin molecules exist as single,
isolated units in RBC, whether oxygen
bound or not
• Cells maintain basic disc shape, whether
transporting oxygen or not
Sickle-cell Trait
• Oxy-hemoglobin is isolated, but deoxyhemoglobin sticks together in
polymers, distorting RBC
• Some cells take on “sickle” shape
Sickle-cell
RBC Distortion
• Hydrophobic valine replaces hydrophilic glutamate
• Causes hemoglobin molecules to repel water and be
attracted to one another
• Leads to the formation of long hemoglobin filaments
Hemoglobin Polymerization
Normal
Mutant
RBC Distortion
• Hydrophobic valine replaces hydrophilic glutamate
• Causes hemoglobin molecules to repel water and be
attracted to one another
• Leads to the formation of long hemoglobin filaments
• Filaments distort the shape of red blood cells
(analogy: icicle in a water balloon)
• Rigid structure of sickle cells blocks capillaries and
prevents red blood cells from delivering oxygen
Capillary Blockage
Sickle-cell Trait
• Oxy-hemoglobin is isolated, but deoxyhemoglobin sticks together in
polymers, distorting RBC
• Some cells take on “sickle” shape
• When hemoglobin again binds oxygen,
again becomes isolated
• Cyclic alteration damages hemoglobin
and ultimately RBC itself
Protein: The Machinery of Life
NH2-Val-His-Leu-Thr-Pro-Glu-GluLys-Ser-Ala-Val-Thr-Ala-Leu-TrpGly-Lys-Val-Asn-Val-Asp-Glu-ValGly-Gly-Glu-…..
“Life is the mode of existence of proteins, and this mode
of existence essentially consists in the constant selfrenewal of the chemical constituents of these
substances.”
Friedrich Engles, 1878