* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Design of Tight-Binding Human Immunodeficiency
Drug discovery wikipedia , lookup
Photosynthetic reaction centre wikipedia , lookup
Peptide synthesis wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Biochemistry wikipedia , lookup
Proteolysis wikipedia , lookup
Biosynthesis wikipedia , lookup
Catalytic triad wikipedia , lookup
Discovery and development of neuraminidase inhibitors wikipedia , lookup
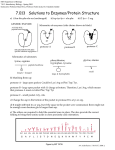
Sathaporn Prutipanlai DESIGN OF PROTEASE INHIBITORS AND RITONAVIR SYNTHESIS Introduction Acquired Immune Deficiency Syndrome (AIDS) appeared as a dangerous disease to mankind. Until this day, human immune deficiency virus, HIV, has caused the death of many humans and infected patients have little chance to become healthy. In the beginning, people become exposed to HIV mainly by unsafe sex with HIV positive person of the same sex. But the situation has changed. Transmission can occur by many ways such as unsafe sex with either sex, sharing contaminated needles during recreational drug use, transfusions with virally contaminated blood, or mother to child. If we do not stop the spread of HIV, it will cause a terrible effect to mankind. For that reason, researchers are trying to understand the life cycle of HIV and develop new drugs, this, however, is a difficult task. From 1981 until today, the spreading of AIDS is a very serious problem especially in Asia. In Thailand, AIDS is a critical problem in health care because there are about 700,000 infected people in this year and the number is expected to increase further. Nowadays, scientists have found that HIV-1 and HIV-2 are the causative agent of AIDS and try to discover new drugs in order to treat or protect humans from AIDS. However, many drugs cause serious side effects and alternative treatment are in clinical trials. AIDS is a perfect module to study and perform the processes of establishing a pharmaceutical factory, analyzing the clinical aspects, and studying the ethical implication. The aim of the production group is to synthesize drugs used in the treatment of AIDS such as ritonavir, vaccines, and Interleukin 2. In addition, the environment, the location of the plant and the cost are other point that our team must consider. My topic is the design of new protease inhibitor and the synthesis of Ritonavir. Ritonavir, HIV protease inhibitor, is one of the most effective drug that is used nowadays and for the treatment of HIV. Function of HIV- Protease Enzyme The human immune deficiency syndrome virus encodes an aspartic protease which is responsible for the posttranslational proteolytic processing of the gag and gag-pol polyprotein gene products into mature, functional proteins. Accompanying these processing events, the nascent viral particles undergo a morphological transformation from immature, noninfectious form into mature, infectious virions. The critical role of functional HIV protease in the HIV replication cycle was initially demonstrated by site directed mutagenesis wherein mutation of the catalytic aspartate residues to either asparagine or alanine generated viral particles which remained in the immature form. Importantly, these particles were incapable of establishing a new round of infection in susceptible T-lymphocytes. Similar behavior was subsequently observed through the blockade of HIV protease by small-molecule inhibitors, thus validating HIV protease as a target for drug design for acquired immune deficiency syndrome Molecular Structure of HIV-protease Enzyme Acquired immune deficiency syndrome (AIDS) appeared as a clinical entity in 1981 and, until nowadays it is estimated that in excess of 22 million people are HIV infected. During 19 years, many researchers have been attempt to synthesis novel drugs that can inhibit replication of this retrovirus. The HIV protease has emerged as 1 Sathaporn Prutipanlai an attractive therapeutic target for the treatment of acquired immune deficiency syndrome (AIDS) because its inhibition interrupts the life cycle of the retrovirus. HIV protease belong to the general class of aspartic acid protease (AAP) enzyme, same as renin and angiotensin. HIV proteases are unique within this class in that functional protease exists as a C2-symmetric homodimer with a single active site(Figure 1). Each monomeric unit contains 99 amino acid and contributes one of the conserved catalytic triads (Asp-Thr-Gly). Two chains assemble to form a long tunnel that is covered by 2 flexible regions of the protein called “flaps”. The flaps open up and the enzyme wraps around a protein chain, closing and holding it tightly in the tunnel. The active site is at the center of the tunnel, where a water molecule is used to break the peptide bond. HIV protease has the job of cutting this long polypeptide into the proper protein – sized pieces. The intact polyprotein is necessary early in the life cycle, when it assembles the immature form of virus. Then, the polyprotein must be cut into proper pieces to form the mature virus, which can then infect a new cell. Because of its sensitive and essential function, HIV-1 protease is an excellent target for drug therapy. Drugs bind tightly to the protease, blocking its action, and the virus perishes because it is unable to mature into its infectious form. Fig1. Molecular structure of HIV- protease (From http://www.rcsb.org/pdb/) The Mechanism of Action of HIV Protease Enzyme Figure. 2 shows a proposed chemical mechanism for HIV-protease that provides mechanistic detail. This mechanism outlined shares many common features with the general acid-base mechanism proposed for the aspartic proteases. The active site of the free HIV protease (EH-) is symmetric and contains a single negative charge at the oxygen of the water bound between the carboxyls. When the substrate binds to form the Michaelis complex, EH-S, the negative charge of the water is engaged in nucleophilic attack on the carbonyl carbon in the substrate peptide bond. The substrate carbonyl oxygen is further polarized by one of the carbonyl hydrogens (HA). At the transition state of the catalysis, ET, the carbonyl carbon of the substrate takes on a tetrahedral conformation with the negative charge transferred to the carbonyl oxygen. The hydrogen bond to another carbonyl hydrogen, HC, facilitates the breaking of the peptide bond of the substrate, which results in the formation of enzyme-product complex, EP. The diffusion away by the substrate returns the enzyme to its free form 2 Sathaporn Prutipanlai and restarts the catalytic cycle (Figure 2). The catalytic mechanism of HIV protease similar to that proposed for eukaryotic aspartic proteases. An important difference is that in free enzyme of HIV protease, the structure and charge distribution are completely symmetrical, as they are contributed by two identical subunits. Once the substrate binds, however, this symmetry no longer exists. The rest of the catalytic cycles of HIV and eukaryotic proteases should be basically similar. The homodimer of HIV protease have two flaps which almost completely close the entrance to the active site cleft. On the other hand, eukaryotic proteases have a single flap that also needs to open up in order to permit the entrance of the substrate and exit of the hydrolytic products. Fig.2 Proposed mechanism of hydrolysis of HIV- protease. Approaches Used to Rationally Design Inhibitors of Protease Enzyme Human immune deficiency virus type1 (HIV-1) protease processes the gag/pol polyprotein, generating important viral enzymes and structural proteins. It is essential for viral replication and offers an attractive strategy for halting the pathogenecity of HIV-1. The approach for designing enzyme inhibitors has involved various methods. When a researcher is confronted with a new proteolytic enzyme, the initial question explored is the sequence and optimal size of the substrate. Once this is known, a medicinal chemist designs possible inhibitors based on the optimal substrate. In the course of designing new compounds for an enzyme, it is desirable to use information about how the enzyme and inhibitor interact in order to construct more potent and novel structure. Fortunately, HIV protease belongs to aspartic acid protease (AAP). AAP can serve as a guide for chemist approach in the design and preparation of potent protease inhibitors because of the many structures of enzyme inhibitor complexes that have been solved. These inhibitors represent all three categories 3 Sathaporn Prutipanlai i) Based on peptide isosteres. Peptide substrate is including statin and its analogs. ii) Exploitation of the symmetrical properties of the protease dimer. iii) Based on structural considerations of the enzyme having no relation to a peptide or peptide analog. Type of designing Protease Inhibitor The type of designing protease inhibitor divided into 5 groups including; Type 1. Non- Hydrolyzable Analogs of Peptide Substrates. Non hydrolyzable analog of peptide substrates are substrates that can not be hydrolyzed by the enzyme. A non hydrolyzable bond is used to replace the scissile bond of the substrate. An inhibitor designed to look like a good peptide substrate is N-acetyl-Thr-Ile-Nle-[CH2-NH]Nle-gln-Arg-NH2(Figure 3). Binding of the inhibitor introduces a substantial conformational changes in the enzyme. The inhibitors are in the direction of closing the cleft between the two monomers of the dimeric protease. N-acetyl-Thr-Ile-Nle-[CH2-NH]Nle-gln-Arg-NH2 Fig 3 Chemical structure of non hydrolyzable analogs of substrate peptides Type 2 Transition-State Analogs. Analogs of peptide substrate in which the scissile dipeptide is replaced by a mimetic of the proteolytic transition state have been shown to be effective inhibitors of the purified HIV- 1 protease. Hydroxyethylene isosteres were found to be potent inhibitors in purified enzyme assay (Figure 4) Hydroxyethylene isosteres Fig 4. Chemical structure of some of the transition-state analogs Type 3. Pepstatin-Protease Complex 4 Sathaporn Prutipanlai Different class of compounds that are known to be potent inhibitors of the acid protease are those containing a statin residue such as acetylpepstatin (Figure5). In fact, this type of compound was used to charactize acid proteases in general. Binding of the inhibitor induced conformational changes but led to less effective inhibition than symmetrical inhibitor (Figure 6). (Sta = Statin) Acetylpepstatin Fig 5 Chemical structure of some of the Pepstatin-Protease complex Fig 6. Schematic diagram of the hydrogen bond network between acetylpepstatin and the HIV- protease. Type 4.Two-Fold Symmetrical or Pseudosymmetrical Inhibitor. The concept, which led to the design of twofold symmetrical compounds as inhibitors of the HIV protease such as ritonavir is based on the observation that the protease is a symmetrical homodimer in the native state. Thus, the active site should exhibit twofold symmetry, and the S1 and S’1 subsite should be indistinguishable in the unligand retroviral enzyme. The recognition of the binding subsite on the either side of the active site to symmetrical inhibitor should be more advantageous. Figure 7 was shown the chemical structure of this inhibitor. 5 Sathaporn Prutipanlai Fig.7 Chemical structure of some of the Two-fold symmetrical or Pseudosymmetrical inhibitor Type 5. Structure-Based Inhibitor Design. This type of binding cause distinctive conformation from peptidomimetic inhibitors. It binds across the pseudotwofold axis of the enzyme. The phenyl rings bind in similar locations related by this twofold symmetry axis, but in different orientations. The five-membered thioketal ring extends into the P2 substrate binding pocket (Figure 8). The largest difference between the structure of the protease complex with this inhibitor and the complex of other inhibitors is the unique conformation of the active site flaps. The conformation of the flaps is intermediate between that in the unliganded enzyme and that of the peptide-analog-bound structures. Fig. 8 The Example of Chemical Structure of the Structure-Based Inhibitor Design Conclusion If one considers the peptide –based HIV protease inhibitors, it is difficult to find a pattern that leads to obviously high affinity. However, the general principle that increasing the hydrophobicity of the inhibitor leads to tighter binding seems to be established. Putative transition state analogs are effective inhibitors because they add at least one strong hydrogen bond. Ritonavir(ABT-538, A-84538) , peptidomimetic inhibitor, was discovered by Abbott’s scientist laboratories in 1994. Ritonavir is chemically designated as 10Hydroxy-2- methyl-5-(1-metylyethyl)-1-[2 (methylethyl)-4-thiazolyl]-3-6-dioxo-8,11bis(phenylmethyl)2,4,7,12-tetraazatridecan-13-oic acid,5-thiazolylmethyl ester, [5S- 6 Sathaporn Prutipanlai (5R*,8R*, 10R*,11R*]. The structure of ritonavir base on C2 symmetry and substrate base inhibitor. Side effect of ritonavir and other protease inhibitor was comparable. These data shown that ritonavir have caused nausea vomiting simitar to other protease inhibitor. But indinavir, was produced by merck’s scientist emerged kidney’s side effect such as renal stone, urolithiasis. Finally, renal failure was developed. In the case of ritonavir, the most side effect occur in liver because liver play a role in this drug metabolism. Other side effect such as circumoral paraesthesia, peripheral paraesthesia and asthenia. Ritonavir used in the dose of 1200 mg/day devided into 2 time . that less than indinavir . Then, ritonavir have been potency and efficacy in order to inhibit retroviral development. Approaches Used to Rationally Design Ritonavir Approaches to the design of peptidomimetic inhibitors of human immune deficiency virus protease must be considered these following factors. i) The recognition, that HIV protease is an aspartic protease. The ramification of this classification is that strategies that had previously proved successful for designing inhibitor of other aspartic protease such as renin can be used for the design of HIV protease inhibitor. ii) C2- symmetric structure of the active HIV protease homodimer. C2 symmetric and pseudo-symmetric inhibitors have been designed in order to make high potency and selectivity of drug. iii) Structure – based inhibitor. Method used to design and preparation begins with asymmetric substrate or inhibitor Principle Destination of C2 Symmetric Inhibitor Core Units Synthesis of symmetric inhibitor core unit can be subdivided into three categories a) Linear, nonsymmetric syntheses Beginning with one half of the inhibitor core, which remainder of the carbon framework, is attached in a linear with a stepwise fashion. For example, diamine was established from Boc-phenylalanine. In the processes of syntheses, First, attachment with another molecule. Then, epoxidation was performed. Finally, epoxide was open with azide in a regioselective manner. b) Symmetric combination of identical halves This type of synthesis produce C2 symmetry by double alkylation of the three-carbon central portion with a chiral enolate introduces the side chain in a stereo controlled fashion. c) Bifunctionalization of a C2-symmetricprecursor. This type of synthesis used a highly functionalized diepoxide serve as the symmetric precursor.to synthesis diamine. The side chains are simultaneously elaborated, followed by introduction the amino groups, masked as azide, with inversion of stereochemistry. The Step of Synthesis C2-Symmetric Inhibitor Core Units. Because of protease enzyme have a structure as C2- symmetry or like butterfly wing, therefore, the core unit of inhibitor has been designed with a C2- symmetry. In addition, the flap region of protease enzyme contains 6-8 amino acid in length. In the process of synthesis, an asymmetric substrate or inhibitor should be used as precursor. The three step conceptual process by which a symmetry-based, peptidomimetic 7 Sathaporn Prutipanlai inhibitor is designed from an asymmetric substrate or inhibitor, is described below. Beginning with the tetrahedral intermediate for cleavage of an asymmetric dipeptide substrate, duplication of the N- terminus by a rotation about a C2 axis bisecting the carbon-nitrogen single bond led to the conceptualization of the symmetric or pseudosymmetric core diamine 1 and 2 (Fig.9). Structure-activity studies on derivative of 1 and 2 led initially to identification of A-77003 (ref 4), name was coded by abbotted’s scientist which can be changed if the final product was success, which, though displaying low oral bioavailability. Subsequently, an improved synthesis of 2 yielded large quantities of the monoprotected diamine 4. To attach heterocyclic carbamates to bind in the S2’ region of the active site, intermediates 2-4 were acylated with the p-nitrophenyl carbonates of the corresponding heterocyclic carbinols(RCH2OH) to produce monoamines 5 and/or 6 following either chromatographic separation or deprotection. For the isopropylthiazolyl and thiazolyl groups, a variety of heterocyclic N-substituted valine esters were required. Various heterocyclic alcohol or amines were prepared and acylated with L- valine methyl ester activated as either the corresponding isocyanate or p-nitrophenyl carbamate. Hydrolysis of the ester followed by carbodiimide-mediated coupling to 5 or 6 produced the desired HIV protease inhibitors with the different peripheral group (isopropylthiazolyl /thiazolyland and phenyl, respectivly)either proximal or distal to the central hydroxyl group of core unit 2.(Fig 10) Fig. 9. Imposition of C2-symmetry axes on an asymmetric substrate or inhibitor Fig.10 Step of ritonavir’s predecessor chemistry Structure- Activity Relationships of C2- Symmetric HIV Protease inhibitors. 8 Sathaporn Prutipanlai Studies of the structure- activity relationships of C2-symmetric inhibitors in order to obtain compounds with inhibitory potency against purified HIV protease in the nanomolar range. As shown in figure 11, change Boc to 2-pyr-CH2OCO-Val at “A” position should increase potency of inhibitor. A H Boc 2-Pyr-CH2OCO-Val Boc 2-Pyr-CH2OCO-Val B H Boc Boc 2-Pyr-CH2OCO-Val 2-Pyr-CH2OCO-Val IC50(nM) >1000 12 1.6 2.4 0.09 Fig 11. Structure- Activity Relationships of C2- Symmetric HIV Protease inhibitors (ref 2.) Binding of C2- symmetric Inhibitors to HIV Protease. The X-ray crystal structure of a number of symmetric and pseudosymmetric inhibitors bound to HIV protease have been reported. The hydrogen bonding interactions of inhibitor with the enzyme active site are shown in Fig 12. The central hydroxyl group of each inhibitor interact with Asp25 andAsp125, and the two carbonyl groups that flank the core unit accept a hydrogen bond from water, the ubiquitous water molecule that bridges the inhibitor and flap region of the enzyme. Fig 12. Hydrogen – bonding interactions of inhibitor bound to HIV protease. Improvment of Ritonavir’s Efficacy The discovery of protease inhibitor inhibitors with utility in the treatment of human disease has been hampered by a number of significant obstacles. Foremost among these is the identification of inhibitors which simultaneously embody potent anti-HIV activity and high oral bioavailability. A virtual necessity for the long term 9 Sathaporn Prutipanlai treatment of chronic disease, high oral bioavailability must be accompanied by long elimination half-life to yield sustained virus-suppressive drug level in the blood and infected tissue. This requirement is prescribed by the profound ability of HIV to develop resistance under selective antiviral pressure. The identification of HIV protease inhibitors with optimal oral pharmacokinetic properties therefore represents a critical milestone on the path to therapeutic efficacy with this class of agents. Unfortunately, as a protease inhibitors have pharmacokinetic profiles that are characterized by low oral absorption and rapid elimination. The major liabilities in this regard are thought to include high molecular weight, low aqueous solubility, and susceptibility to proteolytic degradation, hepatic metabolism, and biliary extraction. A77003 (ref4) and A 80987(ref4) were discovered before Ritonavir (ABT538) but they can not used as a drug. A77003 possessed sufficient aqueous solubility for intravenous injection but low oral bioavailability. In the search for related inhibitors with improved oral bioavailability, the researcher examined the subtle effects of molecular size, aqueous solubility, and hydrogen bonding capability on pharmacokinetic behavior. ABT 538, shown in Fig 13 has additional hydrophobic interaction between the isopropyl substituent on the P3 thiazolyl group and the side chains of Pro-80 and Val-82 of HIV protease as shown in fig14 The delicate balance between solubility and oral absorption was emphasized by carbamate analogue of ABT 538, which was insoluble in the dosing vehicle. Therefore, the dissolution of the inhibitor was critical for good bioavailability. However, the improved pharmacokinetic profile of ABT-538 also correlate with the rate of metabolism by reducing the oxidation potential of the electron-rich pyridinyl groups. But pyridinyl group is significant in aqueous solubility and resulting in change oral absorption. Replacement of pyridine with thiazole pursued to balance aqueous solubility and metabolic stability as shown in figure13 Fig 13 Structures of A-77003, A-80987, and ABT-538; Ph, phenyl 10 Sathaporn Prutipanlai Fig 14. Preliminary crystal structure of ritonavir(ABT538/HIV-1 protease, Showing a hydrophobic cluster between Pro-81, Val-82 and the P3 isopropylthiazolyl and P1 phenyl groups of ritonavir Factor that influence the elaboration of inhibitors 1. the inhibitor include functionality capable of interaction with hydrophobic groups 2. the inhibitor have symmetry- related P2 subsite of the enzyme active site 3. The length of the active site cleft might be limit the extention of inhibitor. 4. Attachment of isopropylthiazolyl group substituents improves activity of inhibitor Possibility for synthesis new protease inhibitor Because of protease enzyme is an attractive target for treatment AIDS, many pharmaceutical laboratories have been targeting this enzyme. Possibility for synthesis of new protease inhibitor depends on many factors. The consideration have been divided into 2 categories; pharmacokinetis profile and pharmacodynamic properties. For pharmacokinetic profile, drug absorption is the first important factor for potency of drug, therefor chemical properties of drug should be; high aqueous solubility, high dissolution rate. Other factor to be considered is the distribution of drug. Plasma protein, especial albumin, disturbes the amount of drug at target organs in a similar manner as with other type of protease inhibitor. In addition, hepatic tissue plays a role in metabolism of drug by its enzyme. The molecular structure of protease inhibitor should be decreasing the lone pair electron in order to decrease oxidation process of drug elimination. For pharmacodynamic properties, designing of protease inhibitor must be fit to cleft area of enzyme and stabilize enzyme, while increasing its potency. Because of protease enzyme is a C2-symmetric model, the inhibitor must be designed in this way in order to block enzyme activity. Chemical processing of ritonavir synthesis 1. Synthetic Chemistry The synthetic pathway that follows is taken from the US patent (#5,846987)for Ritonavir. The chemical processes compose of 9 reaction. Each reaction is covered briefly in order to build an understanding of the reagents needed to produce Ritonavir. Anyway, each scheme shows only interested product that used to synthesiz further. 11 Sathaporn Prutipanlai Synthesis protected alphaaminoaldehyde Reaction 1. Coupling reaction of alpha-amino aldehyde produce diols Treatment dial with acetoxyisobutyryl bromide produce bromoacetate Reaction 2. Reaction 3 . RX: Hydrolysis produce diamine Reaction 6. Produce Oxacyclohexane Reaction 7 RX: Reduction produce Diphenyl hydrohexane Precipitation and filtration produce epoxide Reaction 5. RX:Hydrolysis Reaction 4. Acylation Reaction produce carbamate Coupling Reaction produce Retronavir Reaction 8. Reaction 9 Fig 15 shown summary of process of ritonavir synthesis. Reaction 1, protected alpha aminoaldehyde was synthesis from DMSO and oxalyl chloride as show in reaction 1. DMSO + Oxalyl Chloride Intermediate 1 Intermediate 1. + N-(((benzyl)oxy)carbonyl)-L- phenylalaninal Intermediate 2 + Triethylamine HCl + Intermediate 2 Dimethylsulfide + aminoaldehyde Reaction 1 The reaction 2, the Ritonavir synthetic pathway is the coupling reaction of protected alpha aminoaldehye ( N-(((benzyl)oxy)carbonyl)-L-phenylalaninal) with tetrahydrofuran and zinc produce a mixture of diols as show in reaction 2. 12 Sathaporn Prutipanlai alpha aminoaldehye 2-5-bis-N- ((benzyl)oxy) carboxy)amino3,4-dihydroxy-1,6-diphenylhexane Reaction2: Coupling reactionof alpha- aminoaldehyde In the reaction 3, the reaction of 2-5-bis-N- ((benzyl)oxy)carboxy)amino-3,4dihydroxy-1,6-diphenylhexane formed in reaction 2 with a alpha-acetoxy-isobutyrylbromide in hexane/dichloromethane produce bromoacetate. This reaction substitutes bromine and acetyl group to hydrogen and hydroxy group, respectively. 2-5-bis-N- ((benzyl)oxy) carboxy) amino-3,4-dihydroxy-1,6-diphenylhexane bromoacetate. Reaction 3 : Produce bromoacetate 3-Acetoxy-2,5-bis-N-(((benzyl)oxy)carboxy)amino)-3-bromo-1,6diphenylhexane was hydrolyzed with concommitant cyclization produces epoxide in reaction 4. Bromoacetate Epoxide Reaction 4.: Produce epoxide Epoxide was reduced with sodium borohydride and trifluoroacetic acid in the next step to produce 2-5-Bis(N(((benzyl)oxyl)carbonyl)amino)1,6-diphenyl-3hydroxyhaxan as show in reaction 5. 13 Sathaporn Prutipanlai 1. NaBH4 2. F3CCOOH Epoxide 2-5Bis(N(((benzyl)oxyl)carbonyl)amino)1, 6-diphenyl-3- hydroxyhaxan Reaction 5 Then, Barium hydroxide hydrolysis of this compound leads to diamine, 2, 5, diamino-1,6 diphenyl-s-hydroxylhexane, as show in reaction 6. 2-5-Bis(N(((benzyl)oxyl)carbonyl)amino) diphenyl-3- hydroxyhaxan diamine Reaction 6: Produce Diamine The reaction 7, treatment of diamine with phenylboronic acid produces 6-(1amino-2-phenylethyl)-4-benzyl-2-phenyl-3-aza-2-bora-1-oxacyclohexane. Diamine 6-(1-amino-2-phenylethyl)-4-benzyl-2phenyl-3-aza-2-bora-1-oxacyclohexane Reaction 7 14 Sathaporn Prutipanlai Acylation of product in reaction 7 by ((5-thiazolyl)methyl)-4-nitrophenyl)carbonate produces carbamate. Reaction 8. Finally, ritonavir establish by carbodiimide-mediated coupling of carbamate to N-((Nmethyl-N-((2-isopropyl-4-thiazoyl methy)amino)carbonyl-L-valine as show in reaction 9. Reaction 9. The final reaction of ritonavir systhesis Conclusions Ritonavir is a peptidomimetric protease inhibitor and has C2-symmetric model in order to increase affinity. With in this series of C2-symmetry-based inhibitors, the structure of ritonavir appears to be optimized for both anti-HIV activity and pharmacokinetics. Most pertubations in structure has led to compounds of inferior properties. Using this 9 reaction scheme and the published protocols, a process plant for the large scale production of ritonavir was investigated as described in the next report. 15 Sathaporn Prutipanlai References 1. United States Patent # 5,846,987:Kempf DJ, Norbeck DW.December 8, 1998 2. Kuo LC and Shafer JA. Method in Enzymology Vol.241:Retroviral Protease. San dieago:Academic Press;1994 3. Kempf DJ and Sham HL. Discovery of Ritonavir, a potent Inhibitor of HIV Protease with High Oral Bioavailability and Clinical Efficacy. J.Med.Chem. 1998;41:602-617 4. Kempf DJ et.al. ABT-538 is a potent inhibitor of human immunodeficiency rirus protease and has hight oral bioavailability in humans. Proc.Natl.Acad.Sci.USA. 1995;92:2484-2488. 5. Kempf DJ et al . Symmetry-Based Inhibitors of HIV Protease. Structure-Activity Studies of Acylated 2,4-Diamino-1,5-diphenyl-3-hydroxypentane and 2,5-Diamino1,6-diphenylhexane-3,4-diol.J.Med.Chem.1993;36:320-330. 16