* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download A GLOBAL TRANSCRIPTIONAL ANALYSIS OF STREPTOCOCCUS
Survey
Document related concepts
Transcript
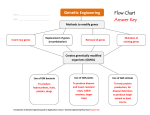
A GLOBAL TRANSCRIPTIONAL ANALYSIS OF STREPTOCOCCUS PNEUMONIAE IN RESPONSE TO LOW-SHEAR MODELED MICROGRAVITY Christopher Allen1, Cristi Galindo1, Nina Williams1, Utpal Pandya1, Ashok Chopra1, and David Niesel1 1 University of Texas Medical Branch, Galveston, TX 77555 USA Long-term habitation in space exposes humans to a variety of environmental stresses not encountered on earth such as ionizing radiation and microgravity. Decreases in immune function and increases in microbial growth and antibiotic resistance during space flight could create an increased health risk to crewmembers (Sonnenfeld et al., 1990; Mishra et al., 1992). Streptococcus pneumoniae is an upper respiratory pathogen which causes pneumonia, meningitis, and bacteremia. S. pneumoniae has previously been isolated from crewmembers and could present a potential health hazard during extended spaceflight (Cioletti et al., 1991). To facilitate groundbased microgravity analogue studies, NASA developed high aspect ratio vessels (HARVs) which model lowshear microgravity conditions (LSMMG). The LSMMG environment is generated through the continual rotation of cell cultures in fluid-filled vessels which minimize prolonged exposure to gravity through the randomization of gravity vectors. Recently, these ground-based models have been used to study the effects of LSMMG on bacteria to understand the physiological and molecular changes they undergo in the presence of microgravity as encountered during spaceflight. This study utilized S. pneumoniae as a Gram-positive bacterial model to investigate how LSMMG affects environmental stress resistance and global transcriptional activity in opportunistic bacterial pathogens. DNA microarray analysis was carried out on S. pneumoniae strain TIGR4 after cultivation in HARVs under LSMMG and compared to 1 x g and static conditions (Figure 1). Significant decreases (p ≤ 0.05) in both thermal and acid stress resistance were found in LSMMG-grown cultures after exposure for 30 minutes using a Student’s t test. Similar results were found after exposure when stress conditions were increased up to 1 hour (data not shown). These results were in contrast to previous reports which have shown increases in environmental stress resistance by Salmonella enterica serovar Typhimurium and Escherichia coli (aWilson et al., 2002; Lynch et al., 2004) in response to LSMMG. These results may be indicative of the different responses by Gram-positive and Gramnegative bacteria to LSMMG. Alternatively, the results may reflect the different phenotypes exhibited by enteric and respiratory pathogens which colonize and persist in distinct niches within the host. Figure 1. HARV Vessel Orientations for Growth. Figure 2. Thermal and Acid Stress Resistance in Response to LSMMG. Microarrays were acquired from The Institute for Genomic Research (TIGR) Pathogen Functional Genomics Resource Center (PFGRC). They were designed based on the genomic sequence of the TIGR4 strain and contain amplicons representing 2131 openreading frames. Stress analysis was carried out on TIGR4 cultures grown to mid-logarithmic phase under LSMMG and controls (1 x g rotating) followed by exposure to thermal (55°C) and acid (pH 3.5) stress conditions (Figure 2). Thermal Stress Resistance Acid Stress Resistance In order to understand what genes may be involved in bacterial LSMMG-mediated responses at the transcriptional level, global transcriptional analysis was carried out using mid-logarithmic phase cultures grown under LSMMG and control cultures (1 x g rotating, 1 x g static). A stringent analysis of the array data was carried out using multiple analysis methods (Genepix Pro 6.0, Spotfire 7.3, SAM, and ANOVA) and a threshold of ≥ 1.5-fold was used to identify differentially expressed genes. Results were confirmed by quantitative real time Gravitational and Space Biology 19(2) August 2006 143 C. Allen et al. – A Global Transcriptional Analysis of Streptococcus pneumoniae in Response to Low-Shear Modeled Microgravity RT-PCR of six representative genes differentially expressed under LSMMG. Genes were chosen which were differentially expressed ≥ 1.5-fold in response to LSMMG, represented different functional categories, and located within different regions within the genome. Pneumococcal genes (81 total) which were found to be responsive to LSMMG represented different functional groups located throughout the genome (Table 1). The statistical analysis program, Significance Analysis of Microarrays (SAM), confirmed the statistical significance of these differentially expressed genes based on three independent experiments. external forces such as shear and rotation exert individual effects on cultured cells independent of LSMMG. Overall, these results indicate that LSMMG represents a unique environment that can alter a bacterium’s transcriptional profile and physiological state. Investigating how this unique environment can lead to new properties will allow us to better understand the impact of LSMMG on microbial virulence and the health risks these changes may create during long-term space habitation. Functional Category Sonnenfeld, G., Mandel A.D., Konstantinova I.V., Taylor G.R., Berry W.D., Wellhausen S.R., Lesnyak A.T., Fuchs B.B. 1990. Effects of Space Flight on Levels and Activity of Immune Cells. Aviat. Space Env. Med. 61: 648. LSMMG-Responsive Genes Antibiotic Resistance Cell Envelope DNA Repair/Recombination Metabolism Signaling Stress Response Transcriptional Regulation Transporters/Ion Channels Hypothetical/Unknown 1 11 4 13 3 1 4 8 36 Table 1. Functional Gene Groups Significantly Altered by LSMMG. REFERENCES Mishra, S., Pierson, D. 1992. Space Flight, Effects on Microorgansims. In: Encyclopedia of Microbiology.Vol. 4. Academic Press, Inc. pp. 53-60. Cioletti, L., Pierson, D., Mishra, S. 1991. Microbial Growth and Physiology in Space: A Review. SAE Technical Paper Series No.911512. a Interestingly, LSMMG-responsive genes were found to be predominantly down regulated in response to LSMMG. Previous studies have shown that global transcriptional profiles of Gram-negative enteric bacteria contain both up- and down-regulated genes after growth under LSMMG (bWilson et al., 2002). This consistant pattern of down-regulation seen in S. pneumoniae represents a distinct response by this Gram-positive respiratory pathogen compared with the more extensively characterized Gram-negative enteric pathogens. Hierarchical clustering analysis (Cluster/Treeview, CLUSFAVOR 6.0, and Spotfire 7.3) revealed several gene clusters displaying similar expression patterns. These clusters consisted of different genes from diverse functional groups. Currently, how these genes function in response to LSMMG remains unknown. Likewise, a common mechanism to explain these regulatory changes in response to LSMMG remains uncharacterized. Further studies will need to be carried out to identify LSMMGresponsive regulators. Controls for the LSMMG experiments included both a 1 x g rotating control and a 1 x g static control (Figure 1) as a secondary filter to identify LSMMG-responsive genes. SAM analysis of genes differentially expressed between static and rotating controls revealed 147 differentially expressed genes. Among these genes, 46 were also altered between 1 x g and LSMMG conditions while 9 were altered between static and LSMMG conditions. The results from this analysis clearly indicate that the choice of control has a significant impact on the assignment of LSMMG-responsive genes and possibly physiological properties. Further, the presence of other 144 Gravitational and Space Biology 19(2) August 2006 Wilson, J., Ott, C.M., Ramamurthy, R., Porwollik, S., McClelland, M., Pierson, D.L., Nickerson, C.A. 2002. Low-Shear Modeled Microgravity Alters the Salmonella enterica Serovar Typhimurium Stress Response in an RpoS-Independent Manner. Appl Environ Microbiol 68(11): 5408-5416. Lynch, S.V., Brodie, E.L., Matin, A. 2004. Role and Regulation of sigma S in General Resistance Conferred by Low-Shear Simulated Microgravity in Escherichia coli. J Bacteriol 186(24): 8207-8212. b Wilson, J.W., Ramamurthy, R., Porwollik, S., McClelland, M., Hammond, T., Allen, P., Ott, C.M., Pierson, D.L., Nickerson, C.A. 2002. Microarray Analysis Identifies Salmonella Genes Belonging to the Low-Shear Modeled Microgravity Regulon. Proc Natl Acad Sci USA 99(21): 13807-13812.