* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download 1.14 Polynomial regression
Survey
Document related concepts
Transcript
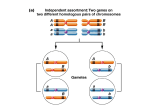
16 Exercises 1.14 Polynomial regression A quite flexible class of models for the mean of a real valued random variable X given a real valued covariate y is EX = β0 + β1 y + β2 y 2 + . . . + βd y d , thus the mean is a d’th order polynomial in the covariate y. Let y1 , . . . , yn be given, real numbers – the covariates – and Xi = β 0 + β 1 y + β 2 y 2 + . . . + β d y d + ε i where the εi ’s are iid with the N (0, σ 2 )-distribution. Then we can estimate the d + 1 parameters β0 , . . . , βd by least squares linear regression. Exercise 1.14.1. Download the dataset for this exercise and load it into R using read.table. You have a data frame with an x column and an y column. Fit polynomial regression models to the dataset, and find out how large d should be for the model to fit the data. Report the estimates for the final model including the estimate of σ 2 . Support your conclusion with graphs etc. Advice on lm: You can either add additional columns to the data frame with the values of y 2 , y 3 etc. before you do the linear regression estimation, or you can directly in the formula specify that you want to regress upon y 2 , y 3 etc. In the last case you need to write something like lm(x ∼ y+ I(y^2)...). The use of I() tells R that you want this to be taken literally as an arithmetic operation on y before regression. The formula x∼y + y^2 has a different interpretation (actually being the same as x∼y for reasons we are not going to explain here). Likelihood functions, genetics and MLE 1.15 17 Likelihood functions, genetics and MLE In genetics one is often interested in estimating the recombination fraction between two loci (genes or markers, say) on the genome. If the loci both are found in two alleles, denoted A and a and B and b, respectively, then if we cross two individuals with allele combinations AaBb and aabb 1 , respectively, the progeny can only get two allele combinations, AaBb and aabb, if there is no recombination (why). We will always assume that they are equally likely, that is, the probability of either of the allele combinations without recombination is 1/2. We introduce the recombination fraction as a parameter θ ∈ Θ = [0, 1], such that the probability of the gamete allele combination Ab from the first individual (which can only occur, if we have recombination) is θ/2. Exercise 1.15.1. Find the probability of all the four possible progeny allele combinations AaBb, Aabb, aaBb, and aabb when the recombination fraction is θ. Write down the likelihood function, the minus-log-likelihood function, and the likelihood equation for estimating θ when observing the allele combinations for n crosses of individuals with combination AaBb and aabb, AaBb n1 Aabb n2 aaBb n3 aabb n4 Total . n = n1 + n2 + n3 + n4 Find the maximum likelihood estimator, find its mean and variance. Exercise 1.15.2. If we observe AaBb 18 Aabb 4 aaBb 6 aabb 27 Total , 55 plot the minus-log-likelihood function and compute the maximum-likelihood estimate. In the F2-cross, we cross AaBb with AaBb 2 . With recombination rate θ ∈ [0, 1] the probability of gamete allele combination Ab is θ/2. Gamete allele combination aB has, likewise, probability θ/2. Exercise 1.15.3. Argue that all 9 (distinguishable) allele combinations are possible when we cross AaBb with AaBb. Assuming the gamete allele combinations are independent in the two gamete cells that fuse, compute the corresponding probabilities when the recombination fraction is θ. Write down the likelihood function, the minuslog-likelihood function, and the likelihood equation for estimating θ when observing the allele combinations for n crosses of individuals with allele combination AaBb and AaBb. Hint: For the computation of the 9 point probabilities, you may start by computing the 16 point probabilities corresponding to the 16 possible combinations of gamete alleles and then sum out over the indistinguishable ones. 1 This is a backcross – starting from two homozygotes AABB and aabb, the progeny will always be a heterozygote, AaBb, for both loci, and then we cross this heterozygote back with its parent homozygote aabb 2 Starting from homozygotes, AABB and aabb, we cross the progeny AaBb with itself 18 Exercises Exercise 1.15.4. Implement a Newton-Raphson algorithm for estimating θ in the F2-cross. If we observe AABB 21 AABb 10 AAbb 2 AaBB 11 AaBb 42 Aabb 6 aaBB 3 aaBb 5 aabb 12 Total , 112 plot the minus-log-likelihood function and compute the maximum-likelihood estimator of θ. Exercise 1.15.5. Using the estimated recombination rate from the previous exercise, simulate B = 200 replications of the F2-cross and (re)estimate the recombination rate for each of the simulated crosses. Investigate, empirically, the distribution of the maximum-likelihood estimator for θ.