* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download III. challenges in drug delivery
Survey
Document related concepts
Psychopharmacology wikipedia , lookup
Orphan drug wikipedia , lookup
Plateau principle wikipedia , lookup
Polysubstance dependence wikipedia , lookup
Neuropsychopharmacology wikipedia , lookup
Compounding wikipedia , lookup
Pharmacognosy wikipedia , lookup
List of comic book drugs wikipedia , lookup
Pharmacogenomics wikipedia , lookup
Pharmaceutical industry wikipedia , lookup
Nicholas A. Peppas wikipedia , lookup
Prescription costs wikipedia , lookup
Prescription drug prices in the United States wikipedia , lookup
Drug interaction wikipedia , lookup
Neuropharmacology wikipedia , lookup
Theralizumab wikipedia , lookup
Drug discovery wikipedia , lookup
Transcript
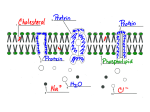
BME 1450 Midterm Paper by Sawitri Mardyani 1 Rational Development of Drug Delivery Systems through Systems Biology Sawitri Mardyani, Member, IEEE Abstract—Up to now, many drug delivery strategies have been developed through hit and miss methods. A move towards rational design requires an understanding of how the components in the human body work together as a system. Using systems biology, drug delivery strategies may be simulated, existing strategies may be optimized, and new strategies may be developed that take advantage of existing structures in the body in order to deliver the right amount of drug to the right place at the right time. Index Terms—drug delivery systems, strategy, simulation, systems biology mechanism-based I. INTRODUCTION I N the process of developing drugs to treat human diseases, it is not sufficient to merely develop a molecule that would act to produce a desired response. There must also be an effective method for delivering the drug to the target tissue. Ideally, the drug delivery strategy should safely transport the drug molecule through the body to reach the target tissue. Through this transport route, the drug should be protected so as not to lose its effectiveness and so that healthy tissues that are passed en route to the sick tissue are not adversely affected by the drug. Once the drug reaches its target site, it should be released in just the right quantity to produce the desired healing effects. This ideal drug delivery situation would optimize the effectiveness of the drug and minimize its side effects. Many past advancements towards the ideal drug delivery strategy have come out of serendipitous discoveries and hit-and-miss research methods [1]. In order to develop a more rational, systematic approach, it is necessary to have an understanding of how the human body works as a whole. Along with providing a better process for creating better drug delivery strategies, this holistic understanding of a biological system may inspire entirely new methods for administering drugs to unhealthy cells or tissue. This paper will discuss the relevance of systems biology to drug delivery research, outline some of the challenges faced in drug delivery and how a systems biology understanding can be used to overcome those challenges. It will also discuss some current drug delivery methods that were developed using knowledge from systems biology. Manuscript submitted October 28, 2003. This work was supported in part by NSERC. S. Mardyani is with the University of Toronto, Toronto, ON M5S 1A1, CANADA (email: [email protected]) II. SYSTEMS BIOLOGY A. A holistic approach Systems biology studies how the components of a biological system work together [2]. It tries to model the interactions of different components in a biological system and then use that model to predict how the system would respond to stimuli [2]. Whereas traditional biology often focused on only one particular component of a system, systems biology aims to study how they all work together [3]. It is this dynamic understanding that is necessary to design effective drug delivery strategies. The immediate appeal of such an approach from a drug delivery standpoint lies in the fact that the drug delivery process involves the interaction between many biological components. In its journey to a target tissue or cell, a drug molecule may pass by several organs and many different types of cells, receptors and proteins. In order to develop effective drug delivery methods, we need to look at the big picture of how these components interact with each other. Putting together this big picture requires an interdisciplinary effort [3]. The information collected by the biologists is compiled to produce a computer model. In this model, equations and algorithms describe the behavior of each component [3]. The complexity of biological system limits the effectiveness of a simple intuitive understanding when it comes to predicting results [2]. Thus, computers are needed to produce accurate simulations of biological systems. Through the combined efforts of biologists, mathematicians, computer scientists, and control systems experts, computer models are made that simulate the behaviors of the biological systems [3]. B. Computer simulation Computer simulation has important applications in drug delivery. A computer model of a biological system can be used to test the effectiveness of a drug delivery strategy in simulation before they are tested in vitro or in vivo. Although in silico testing cannot replace the wet lab altogether, it can provide valuable insights into what strategies work best. It can be used to optimize delivery methods and develop new ones. C. Working with the system Besides the use of computer simulations for research, one of the most valuable fruits gained from using systems biology for development of drug delivery strategies simply comes from the new way of looking at things. By studying the system and BME 1450 Midterm Paper by Sawitri Mardyani understanding how it works, we can develop delivery methods that work with the system by capitalizing on existing structures and pathways. Mechanism based approaches can be designed to more effectively use the existing biological structures to transport drug molecules to their target. Such approaches have been used in the development of cancer treatments that are now starting to appear on the market [5]. Essentially, these new cancer treatments are designed to halt a specific cancer-causing mechanism. The difference in applying this design strategy to drug delivery is that we are not looking for ways to disable an existing mechanism but to take advantage of it. We can use the human body and its pathways of operation as a blueprint for our drug delivery strategies. Like human-made systems, biological systems exhibit patterns in the way their parts work [4]. Rather than completely re-inventing the wheel, we can mimic pre-existing systems in our attempts to deliver drug molecules to specific tissue. By understanding and then adopting existing biological systems, patterns of operation or ‘motif’, we will be able to deliver drugs by working in cooperation with the human body’s own systems and components [4]. D. A knowledge-based approach Much of the difficulty in developing effective drug delivery methods arises out of our limited understanding about how the human body works. Rational design of drug delivery methods cannot proceed without first trying to fill this knowledge gap. Because drug delivery involves the interaction between many different biological components, it is necessary to develop a holistic understanding of how the body works as a system. This knowledge-based approach can be used to tackle many of the challenges seen in drug delivery today. III. CHALLENGES IN DRUG DELIVERY A. The gastrointestinal tract Oral drug delivery is one of the simplest methods to bring a drug molecule into the body. It is especially important in developing countries where clean syringes and medical professionals may be hard to find [6]. However, especially in the case of protein-based drugs, the low pH of the gastro intestinal tract may degrade a drug molecule so that it loses its function [7]. Thus, in order to preserve its effectiveness a drug must be protected from the acid in the stomach. Once it passes through the stomach, the drug must be absorbed in the intestine. Despite numerous animal studies, there remains controversy about the efficiency and the site of intestinal absorption [7]. Many of these studies use microscopy techniques where the sample preparation process may introduce artifacts into the measurements [7]. Further study must be done to develop a better understanding of this process. A comprehensive study of absorption through the intestine can be done using a systems biology approach. 2 First, data from previous studies would be collected, keeping in mind the context in which these studies were performed [2]. Data from studies about the intestine as a whole, and individual cells in the intestine as well as components inside those cells would be put together to make a computer model of the intestinal absorption process. Predictions from this computer model would be compared to experimental results. Where predictions do not match the results, the assumptions made by the model must be examined and new experiments can be conducted to ascertain the validity of such assumptions [2]. Through this iterative process, the computer model of the intestine is refined and a predictive model can be developed [2]. With this predictive model, new oral drug delivery strategies may be tested in silico before they are tested in vivo. A similar strategy has been used to model drug delivery for cystic fibrosis. Using a supercomputer model of the lungs and fluid dynamic equations to describe the trajectory of inhaled drug particles, this model offered new insight into the deposition patterns of inhaled drugs for patients with cystic fibrosis [8]. Using the model, inhaled drugs can be designed for optimal deposition in affected areas of the lung [8]. A similar model of the intestine could provide valuable insight into what forms of drug delivery vehicles would provide the most efficient transfer of a drug into the blood stream. B. The Blood Stream The blood stream is another challenge met by drug molecules. As the drug molecule travels through the blood stream, it must be able to find its destination tissue. The blood stream poses many hazards to an unprotected drug molecule. Foreign molecules do not stay very long in the blood stream as the body is designed to identify these molecules, disable them, and excrete them. The natural defense mechanisms are particularly problematic for drugs such as anti-coagulants whose targets are in the blood stream. One recent strategy to overcome this problem was to attach the drug molecule to red blood cells and then inject these cells back into the blood stream. The Muzykantov research group from the University of Pennsylvania used this strategy to deliver a drug that destroys blood clots in the heart. By disguising the drug under the cover of a red blood cell, the research group was able to increase its effectiveness. Using this Trojan horse strategy, the drug attached to the red blood cell stayed in circulation more than ten times longer than unattached red blood cells and was thus able to eliminate more clots. This delivery method also reduced side effects by keeping the drug in the vascular system away from other tissues where it may have caused excess bleeding. [9] This novel drug delivery strategy is an excellent example of rational design. By using the body’s existing vascular system to their advantage, the group was able to increase the effectiveness of the drug while reducing side-effects. Through increased understanding gained from studies in systems biology, similar solutions in even more complex environments may be designed. BME 1450 Midterm Paper by Sawitri Mardyani 3 Fig. 2. Implantable self-regulation responsive drug delivery system under development by ChipRx. [13] This kind of thinking, where known biological processes are used to inspire new delivery strategies, will only increase with the increased knowledge from systems biology. Fig. 1. Receptor-mediated endocytosis [10] C. Traversing the cell membrane The cell membrane poses a difficult challenge for a drug molecule to overcome. Its function is to protect the cell from foreign invaders while allowing cell nutrients to enter and cell waste to exit [1]. Unfortunately, because the drug molecule is a foreign invader, it has a difficult time crossing this membrane. One way to bring a drug into the cell is to hijack the cell’s own transporters. Exploiting receptor-mediated endocytosis is one example of this strategy. Fig. 1 shows a schematic of receptor-mediated endocytosis. Here, a drug carrier takes advantage of protein transport systems built into the cell membrane. Receptors in the cell membrane produce vesicles that ferry large molecules into the cell. These receptors are able to recognize different molecules and will only give passage to molecules that the cell needs. By disguising itself as one of these needed molecules, a drug molecule carrier can hijack this transport system and penetrate through the cell membrane. [1] In order to exploit receptor-mediated endocytosis for drug delivery, we must know what sorts of molecules the cell is looking for. The capsule carrying the drug can then be conjugated with that molecule so when the cell takes in its molecule of interest, it will bring the drug molecule in with it. This strategy has been successfully exploited for some new nano-particle delivery systems that are still in experimental stages [11]. A further development to this delivery scheme could be to take advantage of the changes in the environment of the drug capsule as it enters the cell through endocytosis. For example, it is known that the interior of the vesicle produced through endocytosis experiences a decrease in the pH as it enters the cell [11]. This is the effect of a proton pump on the vesicle membrane [11]. This decrease in the pH can be exploited as a mechanism to release the drug from its transportation capsule. One way to implement this would be to use polymer-based drug carriers that mimic the body’s secretory granules [12]. If these carriers are made small enough, they may be able to enter the cell through endocytosis and carry drugs to the cell. D. Regulated drug delivery Bringing the drug to the target tissue is just one aspect of drug delivery. Administering the right amount of drug to the target tissue is another challenge. This involves a feedback loop where the response of the patient to the administered drug dosage will control the subsequent dosage [14]. This feedback loop can exist on many levels. Usually, the physician will monitor the status of the patient and administer medication accordingly. In some cases, especially in the administration of analgesics, the patient may request or even administer additional doses based on the presence of pain [14]. One drawback of these systems is the delay between the body producing a signal and the administered medication taking effect in response to that signal. It takes time for a patient to detect a signal and communicate it to the physician. Then the physician must make a decision, decide what drug at what dose to deliver and then deliver the drug. In addition, more time is then taken for the drug to enter the system, find the target tissue and then take effect. Another drawback of these feedback loops is the cost of monitoring the system for responses and the tests required to measure those responses. By bringing the administration control directly to the site of the drug release, this delay and cost can be significantly reduced. The tissue and the drug could act together in a feedback loop. If the drug delivery vehicle can sense when the tissue needs more drugs, it could respond by increasing the dose. Conversely, if the tissue has healed and is no longer in need of these drug molecules, the drug delivery system should stop administering them. Through understanding the biological system, we can find what biochemical signals a drug vehicle can monitor and use as controls for determining the dose to administer. Using a computer simulation we can find the optimal dose that will provide the desired response. An implantable self regulating drug delivery chip is currently being developed by ChipRx, a biotech company in Utah [13]. Fig. 2 shows a schematic of their implant. It uses biosensors to determine the correct amount of drug to administer and artificial muscles control the administration of the drugs. BME 1450 Midterm Paper by Sawitri Mardyani IV. DRUG DELIVERY MODELS As well as providing the inspiration for new methods of drug delivery, a systems biology approach can also be used to analyze existing drug delivery strategies. Baxter et al used a model to study the effects of physiological and antibody-related variables on their antibody mediated therapy. Their model included compartments representing organs of interest, connected in an anatomical fashion. Mass balance equations were used to represent the intra and extravascular spaces in each organ. Variables including organ volumes, blood flow rates, vascular permeability, and binding affinity of the drug molecule were included in the simulation. This model was found to be useful in determining what factors affect antibody uptake in target tissue, optimizing treatment parameters and suggesting experiments to gather data on important parameters. [15] More recently, a computer model was used to test the delivery of asthma medication. Data from MRI images was used to create a three dimensional computer model of the lung. Predicted drug deposition patterns were compared with SPECT (single photon emission computed tomography) images from human subjects. Results from the model agree with the results from human tests. [16] This model can be customized for individual patients. Because asthma is a disease with many forms and variations, a generic treatment regime may not be effective for many patients. The ability to model each patient’s lungs is important for tailoring the prescription to his or her needs. [16] V. CONCLUSION Systems biology has the potential to make many contributions in drug delivery. Using an understanding of the human body as a system, mechanism-based delivery strategies may be developed. Computer simulation may also be used to gain a better understanding of the dynamics of a system, to test delivery strategies in silico and to tailor prescriptions to individual patients according to their needs. REFERENCES [1] [2] [3] [4] [5] [6] [7] J. Alper, “Breaching the Membrane,” Science, vol. 296, pp. 839-841, May 2002. T. Ideker, T. Galitsky, and L. Hood, “A New Approach to Decoding Life: Systems Biology,” Annu. Rev. Genomics Hum. Genet., vol. 2, pp. 343-372, 2001. H. Kitano, “Looking beyond the details: a rise in system-oriented approaches in genetics and molecular biology,” Curr. Genet., vol. 41, pp. 1-10, Apr. 2002 E. J. Davidov, J.M. Holland, E.W. Marple, and S. Naylor, “Advancing drug discovery through systems biology,” Drug Discovery Today, vol. 8, pp. 175-183, Feb. 2002. J.B. Gibbs, “Mechanism-Based Target Identification and Drug Discovery in Cancer Research,” Science, vol. 287, pp. 1969-1973, Mar. 2002. C.T. Vogelson, “Advances in drug delivery systems,” Modern Drug Discovery, vol. 4, pp. 49-50, Apr. 2001. H. Chen and R. Langer, “Oral particulate delivery: status and future trends,” Advanced Drug Delivery Reviews, vol. 34, pp. 339-350, 1998. 4 [8] [9] [10] [11] [12] [13] [14] [15] [16] T. B. Martonen, D. Hwang, I. Katz, Y. Yang, and X. Guan, “Cystic fibrosis: treatment with a supercomputer drug delivery model,” Advances in Engineering Software, vol. 28, pp. 359-364, 1997. J.-C. Murciano, S. Medinilla, D. Eslin, E. Atochina, D. B. Cines, and V. R. Muzykantov, “Prohylactic fibrinolysis through selective dissolution of nascent clots by tPA-carrying erythrocytes,” Nat. Biotech,, vol. 21, pp. 891-896, Aug. 2003. M. Martin-Fernandez, “Time Resolved Microfluorimetry – a method for monitoring cell signaling and the uptake of hormones in human cells,” [Online]. Available: http://srs.dl.ac.uk/vuv/home-page/hot-topics/microfluorimeter.html. J. A. Hubbell, “Enhancing Drug Function,” Science, vol. 300, pp. 595-596. Apr. 2003. P. F. Kiser, G. Wilson, and D. Needham, “A synthetic mimic of the secretory granule for drug delivery,” Nature, vol. 394, pp. 459-462, July 1998. ChipRx, “Products”, [Online], Available: http://www.chiprx.com/products.htm. S. S. Hacisalihzadeh, “Control Engineering and Theraputic Drug Delivery,” IEEE Control Systems Magazine, pp. 44-45, June 1989. L. T. Baxter, “Antibody Mediated Therapy,” Advanced Drug Delivery Reviews, vol. 17, pp. 255-256, 1995. T. Martonen, J. Fleming, J. Schroeter, J. Conway, and D. Hwang, “In silico modeling of asthma,” Advanced Drug Delivery Reviews, vol. 55, pp. 829-849, 2003.