Chapter 19 Nucleic Acids
... RNA Polymerase • RNA polymerase (RNA pol) catalyzes DNAdirected RNA synthesis (transcription) • RNA pol is core of a larger transcription complex • Complex assembles at one end of a gene when transcription is initiated ...
... RNA Polymerase • RNA polymerase (RNA pol) catalyzes DNAdirected RNA synthesis (transcription) • RNA pol is core of a larger transcription complex • Complex assembles at one end of a gene when transcription is initiated ...
Regulation of Transcription
... of a group of genes (i.e. heat shock proteins) A single gene may be regulated by a number of independent transcription factors (i.e. metallothionine) Eukaryotic regulation does not seem to involve repression To achieve high levels of expression, several different transcription factors binding to dif ...
... of a group of genes (i.e. heat shock proteins) A single gene may be regulated by a number of independent transcription factors (i.e. metallothionine) Eukaryotic regulation does not seem to involve repression To achieve high levels of expression, several different transcription factors binding to dif ...
Promoter Regions
... Transcription Start Site: The beginning of RNA transcription. Downstream of binding sequences. Activator: A protein that binds DNA and stabilizes the binding of transcription factors. Activator Site: The region of DNA an activator binds to. Repressor: A protein that binds DNA and destabilizes the bi ...
... Transcription Start Site: The beginning of RNA transcription. Downstream of binding sequences. Activator: A protein that binds DNA and stabilizes the binding of transcription factors. Activator Site: The region of DNA an activator binds to. Repressor: A protein that binds DNA and destabilizes the bi ...
Slide 1
... and multiple genes in a pathway are expressed in a coordinated fashion because such genes are often transcribed as part of a polycistronic mRNA. In eukaryotes, the expression of each gene is typically controlled by multiple transcription factors, and the coordinated expression of different genes dep ...
... and multiple genes in a pathway are expressed in a coordinated fashion because such genes are often transcribed as part of a polycistronic mRNA. In eukaryotes, the expression of each gene is typically controlled by multiple transcription factors, and the coordinated expression of different genes dep ...
biological sciences 354
... Course Content: The goal of this course is to provide students with a comprehensive understanding of transcriptional and post-transcriptional regulation of eukaryotic genes. This goal will be achieved using a combination of lecture, textbook readings, recent reviews, and readings from the current li ...
... Course Content: The goal of this course is to provide students with a comprehensive understanding of transcriptional and post-transcriptional regulation of eukaryotic genes. This goal will be achieved using a combination of lecture, textbook readings, recent reviews, and readings from the current li ...
Gene Regulation - public.iastate.edu
... » these enzymes Ï x103 in 15 min after supply of lactose (host drinking milk) ie ‘inducible enzymes’ ...
... » these enzymes Ï x103 in 15 min after supply of lactose (host drinking milk) ie ‘inducible enzymes’ ...
Gene Section BACH2 (BTB and CNC homology 1, basic leucine
... Comparison of the mRNA profile of a CML cell line, BV173, before and after imatinib treatment revealed an accumulation of BACH2 mRNA upon BCR-ABL kinase inhibition (Vieira et al., 2001). This upregulation of BACH2 by imatinib was seen in lymphoid BCR-ABL1-positive cell lines, as well as in CD34+ cel ...
... Comparison of the mRNA profile of a CML cell line, BV173, before and after imatinib treatment revealed an accumulation of BACH2 mRNA upon BCR-ABL kinase inhibition (Vieira et al., 2001). This upregulation of BACH2 by imatinib was seen in lymphoid BCR-ABL1-positive cell lines, as well as in CD34+ cel ...
Chapter 11 Lecture PowerPoint - McGraw Hill Higher Education
... Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. ...
... Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. ...
Discussion of control of the lac operon and mutational analysis
... Lac Operon: Negative Control and Induction ...
... Lac Operon: Negative Control and Induction ...
TRANSCRIPTION AND PROCESSING OF RNA
... (translational open reading frames (ORF) or cistrons). In prokaryotes polycistronic mRNAs are common. In eukaryotes, monocistronic mRNAs are the general rule, but some transcription units encode more than one polypeptide as a consequence of alternative transcriptional start sites and/or alternative ...
... (translational open reading frames (ORF) or cistrons). In prokaryotes polycistronic mRNAs are common. In eukaryotes, monocistronic mRNAs are the general rule, but some transcription units encode more than one polypeptide as a consequence of alternative transcriptional start sites and/or alternative ...
As Powerpoint Slide
... Fig.1 Effects of target gene overexpression on lycopene production by engineered E. coli . pACLYC04 contains the Erwinia herbicola crtE , crtB and crtI genes necessary for lycopene biosynthesis in E. coli . pBAD24 were used as vectors for dxs , idi , appY , rpoS , yjiD and ycgW expression. dxs : enc ...
... Fig.1 Effects of target gene overexpression on lycopene production by engineered E. coli . pACLYC04 contains the Erwinia herbicola crtE , crtB and crtI genes necessary for lycopene biosynthesis in E. coli . pBAD24 were used as vectors for dxs , idi , appY , rpoS , yjiD and ycgW expression. dxs : enc ...
Activator Proteins
... • first level of DNA packing • histone proteins • 8 protein molecules • many positively charged amino acids • bind tightly to negatively charged DNA ...
... • first level of DNA packing • histone proteins • 8 protein molecules • many positively charged amino acids • bind tightly to negatively charged DNA ...
Developmental Biology 8/e - Florida International University
... 9.28 Expression and regulatory interactions among gap genes products High levels of Bicoid and Hunchback induce the expression of giant, while Kruppel transcript appears over the region where Hunchback begins to decline. ...
... 9.28 Expression and regulatory interactions among gap genes products High levels of Bicoid and Hunchback induce the expression of giant, while Kruppel transcript appears over the region where Hunchback begins to decline. ...
【Grant-in-Aid for Scientific Research (S)】 Biological Sciences
... the multicellular organisms. Since land plants do development in Physcomitrella. not have centrosomes and asteroid bodies, both of [Research 3] Based on the results in Researches 1 which are involved in the axis formation of and 2, functions of orthologous genes to cell metazoans, land plants should ...
... the multicellular organisms. Since land plants do development in Physcomitrella. not have centrosomes and asteroid bodies, both of [Research 3] Based on the results in Researches 1 which are involved in the axis formation of and 2, functions of orthologous genes to cell metazoans, land plants should ...
Resources: http://sciencevideos
... Explain the process of transcription in prokaryotes, including the following: promoter region, RNA polymerase, 5’-3’ direction, free nucleoside triphosphates, complementary base pairing, terminator region. ...
... Explain the process of transcription in prokaryotes, including the following: promoter region, RNA polymerase, 5’-3’ direction, free nucleoside triphosphates, complementary base pairing, terminator region. ...
The Transcription Process
... In any case, upon binding, the RNA pol "core enzyme" binds to another subunit called the sigma subunit to form a holoezyme capable of unwinding the DNA double helix in order to facilitate access to the gene. The sigma subunit conveys promoter specificity to RNA polymerase; that is, it is responsibl ...
... In any case, upon binding, the RNA pol "core enzyme" binds to another subunit called the sigma subunit to form a holoezyme capable of unwinding the DNA double helix in order to facilitate access to the gene. The sigma subunit conveys promoter specificity to RNA polymerase; that is, it is responsibl ...
Leukaemia Section t(2;21)(p11;q22) Atlas of Genetics and Cytogenetics in Oncology and Haematology
... Genetics, Dept Medical Information, UMR 8125 CNRS, University of Poitiers, CHU Poitiers Hospital, F86021 Poitiers, France (JLH) Published in Atlas Database: February 2003 Online updated version: http://AtlasGeneticsOncology.org/Anomalies/t0221p11q22ID1261.html DOI: 10.4267/2042/37962 This work is li ...
... Genetics, Dept Medical Information, UMR 8125 CNRS, University of Poitiers, CHU Poitiers Hospital, F86021 Poitiers, France (JLH) Published in Atlas Database: February 2003 Online updated version: http://AtlasGeneticsOncology.org/Anomalies/t0221p11q22ID1261.html DOI: 10.4267/2042/37962 This work is li ...
Figure S4 Phylogenetic analysis of MdMYB121 and abiotic
... Figure S5. Phylogenetic analysis of MdoMYB121 and abiotic stress-related MYBs from other species. The tree was constructed using the neighbor-joining method of the MEGA5 program with 1000 bootstrap replicates. OsMYB, HvMYB, TaMYB, GmMYB, ZmMYB, CpMYB, and CmMYB protein from Oryza sativa, Hordeum vul ...
... Figure S5. Phylogenetic analysis of MdoMYB121 and abiotic stress-related MYBs from other species. The tree was constructed using the neighbor-joining method of the MEGA5 program with 1000 bootstrap replicates. OsMYB, HvMYB, TaMYB, GmMYB, ZmMYB, CpMYB, and CmMYB protein from Oryza sativa, Hordeum vul ...
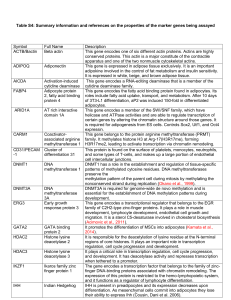
Table S4: Summary information and references on the properties of
... significant histone acetyltransferase acticity with core histones (H3 and H4), and also with nucleosome core particles. It functions as histone acetyltransferase that regulate transcription via chromatin remodeling. Histone demethylase that specifically demethylates lysine 9 and lysine 36 residues o ...
... significant histone acetyltransferase acticity with core histones (H3 and H4), and also with nucleosome core particles. It functions as histone acetyltransferase that regulate transcription via chromatin remodeling. Histone demethylase that specifically demethylates lysine 9 and lysine 36 residues o ...
Chapter 15 - jl041.k12.sd.us
... and not protected by nuclear envelope) and this DNA molecule is not bound up with histones. Thus, gene regulation in prokaryotes is unique. One of the best known pathways of gene recognition is the lac Operon, a regulatory pathway by which bacteria are able to produce the enzyme to digest lactose on ...
... and not protected by nuclear envelope) and this DNA molecule is not bound up with histones. Thus, gene regulation in prokaryotes is unique. One of the best known pathways of gene recognition is the lac Operon, a regulatory pathway by which bacteria are able to produce the enzyme to digest lactose on ...
Gene7-21
... 21.7 Binding to the response element is activated by ligand-binding 21.8 Steroid receptors recognize response elements by a combinatorial code 21.9 Homeodomains bind related targets in DNA 21.10 Helix-loop-helix proteins interact by combinatorial association 21.11 Leucine zippers are involved in dim ...
... 21.7 Binding to the response element is activated by ligand-binding 21.8 Steroid receptors recognize response elements by a combinatorial code 21.9 Homeodomains bind related targets in DNA 21.10 Helix-loop-helix proteins interact by combinatorial association 21.11 Leucine zippers are involved in dim ...
Transcription
... 2. Enhancers bind activating transcription regulators (repressing factors bind to silencers) 3. Enhancers may function – in making the promoter accessible (chromatin remodelling and modifications) – changing DNA topology (e.g.bending) – interacting with the basal transcription apparatus 4. Promoters ...
... 2. Enhancers bind activating transcription regulators (repressing factors bind to silencers) 3. Enhancers may function – in making the promoter accessible (chromatin remodelling and modifications) – changing DNA topology (e.g.bending) – interacting with the basal transcription apparatus 4. Promoters ...
CHAPTER 17 FROM GENE TO PROTEIN Learning Objectives The
... 10. Explain how RNA polymerase recognizes where transcription should begin. Describe the role of the promoter, the terminator (in bacterial cells), and define the transcription unit. 11. Explain the general process of transcription, including the three major steps of initiation, elongation, and term ...
... 10. Explain how RNA polymerase recognizes where transcription should begin. Describe the role of the promoter, the terminator (in bacterial cells), and define the transcription unit. 11. Explain the general process of transcription, including the three major steps of initiation, elongation, and term ...
Preview from Notesale.co.uk Page 4 of 14
... Describe the role of the promoter and the TATA box Eukaryotic promoters contain a sequence called a TATA box which is centred upstream from the transcriptional site. Transcription proteins bind to this promoter initiating transcription by forming a transcription initiating complex which causes t ...
... Describe the role of the promoter and the TATA box Eukaryotic promoters contain a sequence called a TATA box which is centred upstream from the transcriptional site. Transcription proteins bind to this promoter initiating transcription by forming a transcription initiating complex which causes t ...
Transcription factor
In molecular biology and genetics, a transcription factor (sometimes called a sequence-specific DNA-binding factor) is a protein that binds to specific DNA sequences, thereby controlling the rate of transcription of genetic information from DNA to messenger RNA. Transcription factors perform this function alone or with other proteins in a complex, by promoting (as an activator), or blocking (as a repressor) the recruitment of RNA polymerase (the enzyme that performs the transcription of genetic information from DNA to RNA) to specific genes.A defining feature of transcription factors is that they contain one or more DNA-binding domains (DBDs), which attach to specific sequences of DNA adjacent to the genes that they regulate. Additional proteins such as coactivators, chromatin remodelers, histone acetylases, deacetylases, kinases, and methylases, while also playing crucial roles in gene regulation, lack DNA-binding domains, and, therefore, are not classified as transcription factors.