Regulation of Gene Expression
... identical • Differences between cell types result from differential gene expression, the expression of different genes by cells with the same genome • Abnormalities in gene expression can lead to diseases including cancer • Gene expression is regulated at many stages ...
... identical • Differences between cell types result from differential gene expression, the expression of different genes by cells with the same genome • Abnormalities in gene expression can lead to diseases including cancer • Gene expression is regulated at many stages ...
Bioinformatik - Brigham Young University
... Check if: 1. There are interactors for your protein in the literature 2. There are databases of interactions where your protein may appear 3. There are homologues of your protein in the protein interaction databases 4. You can predict interactors by other means? 5. This failing, at this point you go ...
... Check if: 1. There are interactors for your protein in the literature 2. There are databases of interactions where your protein may appear 3. There are homologues of your protein in the protein interaction databases 4. You can predict interactors by other means? 5. This failing, at this point you go ...
Ballas and Mandel 2005
... with its signature functional domains. REST harbors three functional domains: a DNA binding domain containing eight zinc-finger motifs that binds to the RE1 motif, and two independent repressor domains one www.sciencedirect.com ...
... with its signature functional domains. REST harbors three functional domains: a DNA binding domain containing eight zinc-finger motifs that binds to the RE1 motif, and two independent repressor domains one www.sciencedirect.com ...
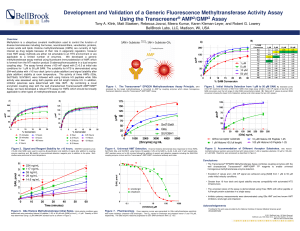
Development and Validation of a Generic
... Methylation is a ubiquitous covalent modification used to control the function of diverse biomolecules including hormones, neurotransmitters, xenobiotics, proteins, nucleic acids and lipids. Histone methyltransferases (HMTs) are currently of high interest as drug targets because of their role in epi ...
... Methylation is a ubiquitous covalent modification used to control the function of diverse biomolecules including hormones, neurotransmitters, xenobiotics, proteins, nucleic acids and lipids. Histone methyltransferases (HMTs) are currently of high interest as drug targets because of their role in epi ...
Insights into Chromatin Structure and Dynamics in Plants
... inter-nucleosome contacts and therefore the chromatin structure itself. Experiments in which tail residues were chemically modified on recombinant histone core preparations showed that acetylation of histone H4 on lysine 16 (H4-K16Ac) has a negative effect on the formation a higher order chromatin s ...
... inter-nucleosome contacts and therefore the chromatin structure itself. Experiments in which tail residues were chemically modified on recombinant histone core preparations showed that acetylation of histone H4 on lysine 16 (H4-K16Ac) has a negative effect on the formation a higher order chromatin s ...
Epigenetic Modifications - Carol Lee Lab
... genome at each generation to define cell types and patterns of gene expression in the developing embryo. These “marks” define which genes are turned on and off. • Marks from the previous generation are typically removed in the germline, to enable totipotency of cells in early embryos • Epigenetic ch ...
... genome at each generation to define cell types and patterns of gene expression in the developing embryo. These “marks” define which genes are turned on and off. • Marks from the previous generation are typically removed in the germline, to enable totipotency of cells in early embryos • Epigenetic ch ...
Epigenetic Inheritance - Carol Eunmi LEE
... -- Genomic imprinting: where methylation and histone modifications alter gene expression without altering the genetic sequence. When inherited, these “epigenetic marks” are established in the germline and are maintained throughout all somatic cells of an organism. -- Gene Silencing: could occur ...
... -- Genomic imprinting: where methylation and histone modifications alter gene expression without altering the genetic sequence. When inherited, these “epigenetic marks” are established in the germline and are maintained throughout all somatic cells of an organism. -- Gene Silencing: could occur ...
Cells Genes in Pro-B H Frequency of Individual V with the
... during lymphocyte development by assembly of Ag receptor gene segments in a process termed V(D)J recombination. In B cell precursors, there is a precise order of rearrangement of the gene segments, with DH to JH rearrangement occurring before VH to DJH rearrangement, and H chain rearranging before L ...
... during lymphocyte development by assembly of Ag receptor gene segments in a process termed V(D)J recombination. In B cell precursors, there is a precise order of rearrangement of the gene segments, with DH to JH rearrangement occurring before VH to DJH rearrangement, and H chain rearranging before L ...
ab115058 – Histone H3 (pan-methyl K4) Quantification Kit (Colorimetric)
... Epigenetic activation or inactivation of genes plays a critical role in many important human diseases, especially in cancer. A major mechanism for epigenetic inactivation of the genes is methylation of CpG islands in genome DNA caused by DNA methyltransferases. Histone methyltransferases (HMTs) cont ...
... Epigenetic activation or inactivation of genes plays a critical role in many important human diseases, especially in cancer. A major mechanism for epigenetic inactivation of the genes is methylation of CpG islands in genome DNA caused by DNA methyltransferases. Histone methyltransferases (HMTs) cont ...
Quantitative modeling of networks – HOG1 study
... By contrast, Sko1/Hot1 activity only alters expression at genes with OR gate logic when Msn2/4 activity is low (for example, in high glucose). The genes regulated in this manner play more generic roles in stress recovery such as neutralizing free radicals and cell wall or cell membrane repair (for ...
... By contrast, Sko1/Hot1 activity only alters expression at genes with OR gate logic when Msn2/4 activity is low (for example, in high glucose). The genes regulated in this manner play more generic roles in stress recovery such as neutralizing free radicals and cell wall or cell membrane repair (for ...
Unit 3. Basic of Biopolymers (3) Control of Protein Function
... targeted to cellular compartments by signal sequences or by attachment of a lipid tail that inserts into membranes. directed to a complex of interacting proteins by a structural interaction domain Localization is a dynamic process and a given protein may be targeted to different compartments at diff ...
... targeted to cellular compartments by signal sequences or by attachment of a lipid tail that inserts into membranes. directed to a complex of interacting proteins by a structural interaction domain Localization is a dynamic process and a given protein may be targeted to different compartments at diff ...
Lab 6
... When arabinose is present in the bacterium’s environment, arabinose binds with the AraC protein, forming a complex. This prevents the DNA loop from forming. The binding of arabinose also causes a change in the protein’s conformation (shape) resulting in the formation of a small pocket that will help ...
... When arabinose is present in the bacterium’s environment, arabinose binds with the AraC protein, forming a complex. This prevents the DNA loop from forming. The binding of arabinose also causes a change in the protein’s conformation (shape) resulting in the formation of a small pocket that will help ...
CHAPTER 18 LECTURE NOTES: CONTROL OF GENE
... The tryptophan operon (a repressible operon; regulation at the level of transcriptional initiation, feedback inhibition, and premature transcriptional termination) A. The trp operon encodes genes that are required for the synthesis of tryptophan (Trp) when it is not available in the growth medium. B ...
... The tryptophan operon (a repressible operon; regulation at the level of transcriptional initiation, feedback inhibition, and premature transcriptional termination) A. The trp operon encodes genes that are required for the synthesis of tryptophan (Trp) when it is not available in the growth medium. B ...
Control of Gene Expression
... normal cell growth and division Conversion of a proto-oncogene to an oncogene can lead to abnormal stimulation of the cell cycle ...
... normal cell growth and division Conversion of a proto-oncogene to an oncogene can lead to abnormal stimulation of the cell cycle ...
Sigma Factors & the Hrp
... Bacteriophage-encoded σ factor also used to take over cellular transcriptional machinery ...
... Bacteriophage-encoded σ factor also used to take over cellular transcriptional machinery ...
Document
... • proteins that bind sequences of DNA to control transcription • can act as activators or repressors to transcription – activating TFs - proteins that recruit the RNA polymerase to a promoter region – repressing TFs – proteins that prevent transcription in many ways • must contain a DNA binding doma ...
... • proteins that bind sequences of DNA to control transcription • can act as activators or repressors to transcription – activating TFs - proteins that recruit the RNA polymerase to a promoter region – repressing TFs – proteins that prevent transcription in many ways • must contain a DNA binding doma ...
Protein Structure
... is able to find alignments involving chain reversals and different topologies. • The algorithm uses distance matrices to represent each structure to be compared. • Application of DALI to the entire PDB produces two classifications of structures: FSSP and DDD (3D). ...
... is able to find alignments involving chain reversals and different topologies. • The algorithm uses distance matrices to represent each structure to be compared. • Application of DALI to the entire PDB produces two classifications of structures: FSSP and DDD (3D). ...
Chapter 18 PPT
... • Differences between cell types result from ? • differential gene expression • Gene expression is regulated at many stages © 2011 Pearson Education, Inc. ...
... • Differences between cell types result from ? • differential gene expression • Gene expression is regulated at many stages © 2011 Pearson Education, Inc. ...
No Slide Title
... InterPro is a database of protein families, domains and functional sites in which identifiable features found in known proteins can be applied to unknown protein sequences. The aim is to provide a one-stop-shop for protein family diagnostics ...
... InterPro is a database of protein families, domains and functional sites in which identifiable features found in known proteins can be applied to unknown protein sequences. The aim is to provide a one-stop-shop for protein family diagnostics ...
Gene Section EIF4EBP1 (Eukaryotic translation initiation factor 4E binding protein 1)
... 4E-BP1 is ubiquitously expressed, although its presence is not essential to the viability of cells or the organism as a whole (Le Bacquer et al., 2007). The level of expression and state of phospho-rylation of the protein may influence cellular phenotype, with high levels of phosphorylated 4E-BP1 in ...
... 4E-BP1 is ubiquitously expressed, although its presence is not essential to the viability of cells or the organism as a whole (Le Bacquer et al., 2007). The level of expression and state of phospho-rylation of the protein may influence cellular phenotype, with high levels of phosphorylated 4E-BP1 in ...
7.2 Transcription and gene expression (HL ONLY
... (active versus inactive genes) that does NOT involve changes to the underlying DNA sequence; Epigenetic change is a regular and natural occurrence but can also be influenced by several factors: ...
... (active versus inactive genes) that does NOT involve changes to the underlying DNA sequence; Epigenetic change is a regular and natural occurrence but can also be influenced by several factors: ...
Managing Associations Between Different Chromosomes
... regulation of expression may be conInterchromosomal rendezvous. The interaction between two different gene loci on two different chromosomes is medi- trolled by interchromosomal interacated by the transcription regulatory factor CTCF and perhaps other factors. This may occur in regions of the nucleu ...
... regulation of expression may be conInterchromosomal rendezvous. The interaction between two different gene loci on two different chromosomes is medi- trolled by interchromosomal interacated by the transcription regulatory factor CTCF and perhaps other factors. This may occur in regions of the nucleu ...
Document
... antisense to Xist at the X-inactivation centre. Nat. Genet. 21, 400404. (Full Text) • Lee, J. T., (2003), X-chromosome inactivation: a multidisciplinary approach. j.semcdb 14, 311-312. ...
... antisense to Xist at the X-inactivation centre. Nat. Genet. 21, 400404. (Full Text) • Lee, J. T., (2003), X-chromosome inactivation: a multidisciplinary approach. j.semcdb 14, 311-312. ...
Histone acetylation and deacetylation

Histone acetylation and deacetylation are the processes by which the lysine residues within the N-terminal tail protruding from the histone core of the nucleosome are acetylated and deacetylated as part of gene regulation. Histone acetylation and deacetylation are essential parts of gene regulation. These reactions are typically catalysed by enzymes with ""histone acetyltransferase"" (HAT) or ""histone deacetylase"" (HDAC) activity. Acetylation is the process where an acetyl functional group is transferred from one molecule (in this case, Acetyl-Coenzyme A) to another. Deacetylation is simply the reverse reaction where an acetyl group is removed from a molecule.Acetylated histones, octameric proteins that organize chromatin into nucleosomes and ultimately higher order structures, represent a type of epigenetic marker within chromatin. Acetylation removes the positive charge on the histones, thereby decreasing the interaction of the N termini of histones with the negatively charged phosphate groups of DNA. As a consequence, the condensed chromatin is transformed into a more relaxed structure that is associated with greater levels of gene transcription. This relaxation can be reversed by HDAC activity. Relaxed, transcriptionally active DNA is referred to as euchromatin. More condensed (tightly packed) DNA is referred to as heterochromatin. Condensation can be brought about by processes including deacetylation and methylation; the action of methylation is indirect and has no effect upon charge.