Top down - The Fenyo Lab
... RED: triplicate experiments, FAl treated grindate BLACK: duplicated experiments, FAl treated cells (then ground) SCORE: Log Ion Current / Log protein abundance ...
... RED: triplicate experiments, FAl treated grindate BLACK: duplicated experiments, FAl treated cells (then ground) SCORE: Log Ion Current / Log protein abundance ...
PrionPPSatBlack
... Determining evolutionary relationships among the various organisms examined. Investigating how amino acid sequence may be linked to the overall structure of the protein Examining the role of repetitive elements in prion homologies. ...
... Determining evolutionary relationships among the various organisms examined. Investigating how amino acid sequence may be linked to the overall structure of the protein Examining the role of repetitive elements in prion homologies. ...
Method for recognizing local descriptors of protein
... Predicting the three dimensional structure of a protein from its amino acid sequence is a very important unsolved problem in bioinformatics [4]. Enabling science to predict proteins threedimensional structures accurately would help scientists understand a variety of different hereditary diseases and ...
... Predicting the three dimensional structure of a protein from its amino acid sequence is a very important unsolved problem in bioinformatics [4]. Enabling science to predict proteins threedimensional structures accurately would help scientists understand a variety of different hereditary diseases and ...
Conserved Positions for Ribose Recognition: Importance of Water
... govern ribose recognition by proteins is sketchy because tertiary and even secondary structural elements involved in nucleotide recognition are very diverse among proteins having different folds.1 Furthermore, although some sequence motifs have been identified within ligand binding folds, in general ...
... govern ribose recognition by proteins is sketchy because tertiary and even secondary structural elements involved in nucleotide recognition are very diverse among proteins having different folds.1 Furthermore, although some sequence motifs have been identified within ligand binding folds, in general ...
Disulfide bridge assignment in complex proteins - HES
... using mass spectrometry, in particular, to enable the study of 'challenging' proteins such as venom proteins, which fail simple disulfide bridge assignment methods. The disulfide assignment strategy is highly dependent on the protein sequence and disulfide bonding pattern. Thus to study a variety of ...
... using mass spectrometry, in particular, to enable the study of 'challenging' proteins such as venom proteins, which fail simple disulfide bridge assignment methods. The disulfide assignment strategy is highly dependent on the protein sequence and disulfide bonding pattern. Thus to study a variety of ...
Evolution of b-type cytochromes in prokaryotes
... The alignment, including 22 eukaryotic cytochrome b561 (K08360) and 61 prokaryotic cytochrome b561 (K12262) sequences (dataset as in Figure 4), was generated using Muscle, and visualized in Jalview. The four helices (pink) predicted by PSIpred for the E. coli sequence, along with the 6 helices based ...
... The alignment, including 22 eukaryotic cytochrome b561 (K08360) and 61 prokaryotic cytochrome b561 (K12262) sequences (dataset as in Figure 4), was generated using Muscle, and visualized in Jalview. The four helices (pink) predicted by PSIpred for the E. coli sequence, along with the 6 helices based ...
Structure of the FHA1 Domain of Yeast Rad53 and Identification of
... two fragments of slightly different size due to a minor thrombin-cleavage site near residue 165. Subsequently, the fragment 2-164 was constructed. However, the ®rst 15 residues were shown to exist in random coil conformations by NMR. These residues were best deleted in order to avoid interfering wit ...
... two fragments of slightly different size due to a minor thrombin-cleavage site near residue 165. Subsequently, the fragment 2-164 was constructed. However, the ®rst 15 residues were shown to exist in random coil conformations by NMR. These residues were best deleted in order to avoid interfering wit ...
Attributes Tutorial
... Apply. The coloring shows that many positions in or near the binding site are highly conserved. The Render by Attribute dialog also lists the residue attribute mavPercentConserved. Whereas mavConservation changes along with the Conservation calculation method, mavPercentConserved is always the perce ...
... Apply. The coloring shows that many positions in or near the binding site are highly conserved. The Render by Attribute dialog also lists the residue attribute mavPercentConserved. Whereas mavConservation changes along with the Conservation calculation method, mavPercentConserved is always the perce ...
Slides
... • Use of astral database to obtain protein domain sequences of the SCOP database. • Retained only superfamilies having atleast two families that have atleast 10 members in each family. • A dataset with1639 domains in 23 superfamilies & 56 families was yielded. • Protein sequences annotated with EC n ...
... • Use of astral database to obtain protein domain sequences of the SCOP database. • Retained only superfamilies having atleast two families that have atleast 10 members in each family. • A dataset with1639 domains in 23 superfamilies & 56 families was yielded. • Protein sequences annotated with EC n ...
Protein and amino acids
... Interpretation of the data is, however, somewhat complicated. The values determined by this method are more correctly termed ‘ileal digestibilities’ rather than bioavailabilities because AAs are sometimes absorbed in a form that cannot be fully used in metabolism. Furthermore, unless a correction is ...
... Interpretation of the data is, however, somewhat complicated. The values determined by this method are more correctly termed ‘ileal digestibilities’ rather than bioavailabilities because AAs are sometimes absorbed in a form that cannot be fully used in metabolism. Furthermore, unless a correction is ...
Methodology for predicting semantic annotations of protein
... part comprises all the methods used in the proposed methodology for extracting and modeling of features derived of the protein sequences and the classification of proteins using those features. The third part of the thesis lies on the evaluation of the method using a reported data set on Gram negati ...
... part comprises all the methods used in the proposed methodology for extracting and modeling of features derived of the protein sequences and the classification of proteins using those features. The third part of the thesis lies on the evaluation of the method using a reported data set on Gram negati ...
Full Text
... a more robust consensus of multiple algorithms. This allows for a more detailed description of such a cavity’s characteristics than in any of our previous works. ...
... a more robust consensus of multiple algorithms. This allows for a more detailed description of such a cavity’s characteristics than in any of our previous works. ...
Pharmacophore screening of the Protein Data Bank for specific
... Step 2: Generating a library of putative pockets. An SDF-formatted virtual library is assembled by searching the PDB for pockets lined by clusters of aromatic and acidic residues. For each PDB structure, the protein is first stripped of water molecules, ligands (including nucleic acids) and bound pe ...
... Step 2: Generating a library of putative pockets. An SDF-formatted virtual library is assembled by searching the PDB for pockets lined by clusters of aromatic and acidic residues. For each PDB structure, the protein is first stripped of water molecules, ligands (including nucleic acids) and bound pe ...
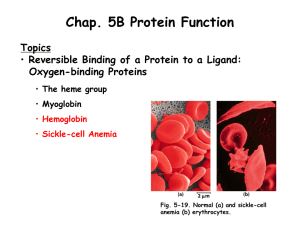
Chapter 5B Lecture
... When the normal ligand and modulator are identical as they are with Hb (i.e., O2), the modulator is termed homotropic. If the modulator is a molecule other than the normal ligand, the interaction is heterotropic. Some proteins can have two or more modulators, and therefore can have both homotropic a ...
... When the normal ligand and modulator are identical as they are with Hb (i.e., O2), the modulator is termed homotropic. If the modulator is a molecule other than the normal ligand, the interaction is heterotropic. Some proteins can have two or more modulators, and therefore can have both homotropic a ...
Overview of Rule Curation
... FATE: After review, the rule (match/exclusion conditions and propagation fields) is sent to EBI for inclusion into the UniRule tool. ...
... FATE: After review, the rule (match/exclusion conditions and propagation fields) is sent to EBI for inclusion into the UniRule tool. ...
Poon, Andy: Predicting Phosphorylation: A critique of the NetPhos program and potential alternatives
... potential phosphorylation sites, the only certainty is that the phosphorylated residue must be a serine, threonine, or tyrosine. These three amino acids have the capacity to bind phosphates because they all contain a hydroxyl group in their side chains which are deprotonated at physiological pH, su ...
... potential phosphorylation sites, the only certainty is that the phosphorylated residue must be a serine, threonine, or tyrosine. These three amino acids have the capacity to bind phosphates because they all contain a hydroxyl group in their side chains which are deprotonated at physiological pH, su ...
Breakfast of Champions
... the most popular supplements used by bodybuilders, strength trainers and health conscience individuals because it contains the most bioavailable protein of all natural food sources. In fact, it should be a staple in your diet. The nice thing is that it is easy to incorporate into your diet so it i ...
... the most popular supplements used by bodybuilders, strength trainers and health conscience individuals because it contains the most bioavailable protein of all natural food sources. In fact, it should be a staple in your diet. The nice thing is that it is easy to incorporate into your diet so it i ...
Structural characterization of L
... l-Glutamate has a flavor-enhancing activity that creates the sensation of ‘umami’; the monosodium salt of l-glutamate is widely used as a seasoning for cooking and as a food additive. The amino acid is also the principal excitatory neurotransmitter in the brain [1,2]. Furthermore, its excessive relea ...
... l-Glutamate has a flavor-enhancing activity that creates the sensation of ‘umami’; the monosodium salt of l-glutamate is widely used as a seasoning for cooking and as a food additive. The amino acid is also the principal excitatory neurotransmitter in the brain [1,2]. Furthermore, its excessive relea ...
This JET help is useful to understand files in the folder
... an average trace value trace(j) 6= 0/ number of trees), propensity value, trace*frequency, accessibility (0 if not accessible, 1 if accessible), mixed trace for clustered residues. If iJET is run instead, with i 6= 1, then the file jet.res contains some extra columns. For each
residue in ...
... an average trace value trace(j) 6= 0/ number of trees), propensity value, trace*frequency, accessibility (0 if not accessible, 1 if accessible), mixed trace for clustered residues. If iJET is run instead, with i 6= 1, then the file
Data-driven docking for the study of biomolecular complexes
... With the presently available amount of genetic information, a lot of attention focuses on systems biology and in particular on biomolecular interactions. Considering the huge number of such interactions, and their often weak and transient nature, conventional experimental methods such as Xray crysta ...
... With the presently available amount of genetic information, a lot of attention focuses on systems biology and in particular on biomolecular interactions. Considering the huge number of such interactions, and their often weak and transient nature, conventional experimental methods such as Xray crysta ...
Facts and Fallacies
... • Better conduct de novo on all spectra. – De novo not slow, and computing is cheap. – De novo provides independent validation for DB result. # consensus AA (de novo vs. DB search) ...
... • Better conduct de novo on all spectra. – De novo not slow, and computing is cheap. – De novo provides independent validation for DB result. # consensus AA (de novo vs. DB search) ...
Efficient Estimation of Emission Probabilities in profile HMM
... residues is incorporated into the profile, whereby the analysis is able to detect structural similarities and homologies to the sequence family. In HMM models, emission probabilities of all 20 amino acids are estimated in all emitting states, and thus the number of estimated parameters can be enormo ...
... residues is incorporated into the profile, whereby the analysis is able to detect structural similarities and homologies to the sequence family. In HMM models, emission probabilities of all 20 amino acids are estimated in all emitting states, and thus the number of estimated parameters can be enormo ...
Protein Supplies for Beef Cattle Diets
... Consider cost per unit of protein and convenience of various protein supplements. Base purchasing decisions on the cost per pound of protein instead of the price per pound of supplement. Product labels indicate the protein percentage and how much protein is in the form of non-protein nitrogen. Conve ...
... Consider cost per unit of protein and convenience of various protein supplements. Base purchasing decisions on the cost per pound of protein instead of the price per pound of supplement. Product labels indicate the protein percentage and how much protein is in the form of non-protein nitrogen. Conve ...
Mutational effects on protein structure and function Jonas Carlsson Link¨
... we managed to explain the severity of all but one of the mutations. By observing the properties of these mutations we could perform good predictions on, at the time, not classified mutations. For the cancer suppressor protein p53, there are over thousand mutations with known activity. To be able to ...
... we managed to explain the severity of all but one of the mutations. By observing the properties of these mutations we could perform good predictions on, at the time, not classified mutations. For the cancer suppressor protein p53, there are over thousand mutations with known activity. To be able to ...
Structural alignment

Structural alignment attempts to establish homology between two or more polymer structures based on their shape and three-dimensional conformation. This process is usually applied to protein tertiary structures but can also be used for large RNA molecules. In contrast to simple structural superposition, where at least some equivalent residues of the two structures are known, structural alignment requires no a priori knowledge of equivalent positions. Structural alignment is a valuable tool for the comparison of proteins with low sequence similarity, where evolutionary relationships between proteins cannot be easily detected by standard sequence alignment techniques. Structural alignment can therefore be used to imply evolutionary relationships between proteins that share very little common sequence. However, caution should be used in using the results as evidence for shared evolutionary ancestry because of the possible confounding effects of convergent evolution by which multiple unrelated amino acid sequences converge on a common tertiary structure.Structural alignments can compare two sequences or multiple sequences. Because these alignments rely on information about all the query sequences' three-dimensional conformations, the method can only be used on sequences where these structures are known. These are usually found by X-ray crystallography or NMR spectroscopy. It is possible to perform a structural alignment on structures produced by structure prediction methods. Indeed, evaluating such predictions often requires a structural alignment between the model and the true known structure to assess the model's quality. Structural alignments are especially useful in analyzing data from structural genomics and proteomics efforts, and they can be used as comparison points to evaluate alignments produced by purely sequence-based bioinformatics methods.The outputs of a structural alignment are a superposition of the atomic coordinate sets and a minimal root mean square deviation (RMSD) between the structures. The RMSD of two aligned structures indicates their divergence from one another. Structural alignment can be complicated by the existence of multiple protein domains within one or more of the input structures, because changes in relative orientation of the domains between two structures to be aligned can artificially inflate the RMSD.