Steven Bhutra - Proteomics of Alzheimer's Disease
... incredibly difficult to control and almost impossible to degrade. The events leading to its formation involve proteolytic cleavage by two enzymes, β-secretase, which fold the amino and carbonyl terminal. The excessive accumulation of this 42 amino acid long protein could be explained by a ...
... incredibly difficult to control and almost impossible to degrade. The events leading to its formation involve proteolytic cleavage by two enzymes, β-secretase, which fold the amino and carbonyl terminal. The excessive accumulation of this 42 amino acid long protein could be explained by a ...
Section Abstracts: Structural Biology, Biochemistry, and Biophysics
... Richmond, Virginia 23298-0032. Dependent binary data can be simply simulated using the multinomial sampling method. W e extend this method to simulate dependent binary data with clustered random effect structures. Several distributions are considered for constructing random effects among cluster-spe ...
... Richmond, Virginia 23298-0032. Dependent binary data can be simply simulated using the multinomial sampling method. W e extend this method to simulate dependent binary data with clustered random effect structures. Several distributions are considered for constructing random effects among cluster-spe ...
Classification of Protein
... Oligopeptides have 10 or fewer amino acids. Polypeptides and proteins are chains of 10 or more amino acids, but peptides consisting of more than 50 amino acids are classified as proteins. ...
... Oligopeptides have 10 or fewer amino acids. Polypeptides and proteins are chains of 10 or more amino acids, but peptides consisting of more than 50 amino acids are classified as proteins. ...
Lecture 3
... Proteins are the most abundant organic molecules in living cells. They may be monomeric with one polypeptide chain or multimeric having more than one chain. In case of a homomultimer the chains are of one kind whereas for a heteromultimer two or more different chains form the protein. (e.g. Hemoglob ...
... Proteins are the most abundant organic molecules in living cells. They may be monomeric with one polypeptide chain or multimeric having more than one chain. In case of a homomultimer the chains are of one kind whereas for a heteromultimer two or more different chains form the protein. (e.g. Hemoglob ...
Essential Amino Acids
... Tryptophan - a relaxant, alleviates insomnia, prevents migraine; reduces anxiety and depression; promotes proper immune system function. It reduces the risk of cardiovascular spasms. Works in conjunction with Lysine to lower cholesterol levels. ...
... Tryptophan - a relaxant, alleviates insomnia, prevents migraine; reduces anxiety and depression; promotes proper immune system function. It reduces the risk of cardiovascular spasms. Works in conjunction with Lysine to lower cholesterol levels. ...
ppt-file
... amino acid metabolism in E. coli are considered. The carbon source 2-phosphoglycerate (PG), CO2, NH3, the produced amino acids and all cofactors such as ATP, ADP etc. are considered “external”. For this scheme, we computed, by the program METATOOL, 14 elementary modes, four of them producing lysine ...
... amino acid metabolism in E. coli are considered. The carbon source 2-phosphoglycerate (PG), CO2, NH3, the produced amino acids and all cofactors such as ATP, ADP etc. are considered “external”. For this scheme, we computed, by the program METATOOL, 14 elementary modes, four of them producing lysine ...
PLoS Pathogens
... of reactive oxygen species (ROS), which are formed in response to the bacterial flagellin peptide flg22. We suggest that NopM promotes nodule initiation by reducing the levels of harmful ROS during the infection process. tobacco and Lotus japonicus, NopL suppressed expression of defense genes [17]. ...
... of reactive oxygen species (ROS), which are formed in response to the bacterial flagellin peptide flg22. We suggest that NopM promotes nodule initiation by reducing the levels of harmful ROS during the infection process. tobacco and Lotus japonicus, NopL suppressed expression of defense genes [17]. ...
Prezentace aplikace PowerPoint
... What is in the Sample Buffer? *Tris buffer to provide appropriate pH *SDS (sodium dodecyl sulphate) detergent to dissolve proteins and give them a negative charge *Glycerol to make samples sink into wells *Bromophenol Blue dye to visualize samples ...
... What is in the Sample Buffer? *Tris buffer to provide appropriate pH *SDS (sodium dodecyl sulphate) detergent to dissolve proteins and give them a negative charge *Glycerol to make samples sink into wells *Bromophenol Blue dye to visualize samples ...
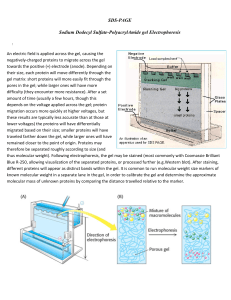
SDS-PAGE Sodium Dodecyl Sulfate
... An electric field is applied across the gel, causing the negatively-charged proteins to migrate across the gel towards the positive (+) electrode (anode). Depending on their size, each protein will move differently through the gel matrix: short proteins will more easily fit through the pores in the ...
... An electric field is applied across the gel, causing the negatively-charged proteins to migrate across the gel towards the positive (+) electrode (anode). Depending on their size, each protein will move differently through the gel matrix: short proteins will more easily fit through the pores in the ...
abstract form
... shown to be significantly increased in comparison with neutrophilic and mesophilic species. It is interesting to note, that the fraction of such proteins of halophilic archeae was decreased, and we did not notice similar patterns in bacterial proteomes. Next, we analyzed distribution of amyloidogene ...
... shown to be significantly increased in comparison with neutrophilic and mesophilic species. It is interesting to note, that the fraction of such proteins of halophilic archeae was decreased, and we did not notice similar patterns in bacterial proteomes. Next, we analyzed distribution of amyloidogene ...
Lecture 13
... • While exciting, these are often very difficult experiments, requiring a lot of time and good control experiments • The data one gets is often ambiguous, but no more so than solid state structures obtained in X-Ray crystallography • It’s crucial to differentiate between solution state and gas phase ...
... • While exciting, these are often very difficult experiments, requiring a lot of time and good control experiments • The data one gets is often ambiguous, but no more so than solid state structures obtained in X-Ray crystallography • It’s crucial to differentiate between solution state and gas phase ...
haemoglobin: structure, properties and biomedical functions
... in mammals and other animals. A spheroidal heme protein having four subunits each consisting of a globular protein non-covalently bound, with an embedded heme group. Hb has a molecular weight of about 64456. The globular protein units of Hb is made up of two identical pairs of polypeptide chains, i. ...
... in mammals and other animals. A spheroidal heme protein having four subunits each consisting of a globular protein non-covalently bound, with an embedded heme group. Hb has a molecular weight of about 64456. The globular protein units of Hb is made up of two identical pairs of polypeptide chains, i. ...
Genetically Modified Organism
... Protease: (proteinases, peptidases or proteolytic enzymes) are enzymes that break peptide bonds between amino acids of proteins ...
... Protease: (proteinases, peptidases or proteolytic enzymes) are enzymes that break peptide bonds between amino acids of proteins ...
PROTEIN STRUCTURE
... • The hydrophobicity of certain amino acid Rgroups tends to drive them away from the exterior of proteins and into the interior. This driving force restricts the available conformations into which a protein may fold. ...
... • The hydrophobicity of certain amino acid Rgroups tends to drive them away from the exterior of proteins and into the interior. This driving force restricts the available conformations into which a protein may fold. ...
Fundamentals of protein structure
... (1)Hydrogen bonds (between side chains) (2)Hydrophobic bonds (between the non-polar side chain of a.a.) (3)Electrostatic bonds (salt bonds)(Formed between oppositely charged group in the side chains of amino acids)e.g. epsilon-amino group of lysine and carboxyl group of aspartate, interact electrost ...
... (1)Hydrogen bonds (between side chains) (2)Hydrophobic bonds (between the non-polar side chain of a.a.) (3)Electrostatic bonds (salt bonds)(Formed between oppositely charged group in the side chains of amino acids)e.g. epsilon-amino group of lysine and carboxyl group of aspartate, interact electrost ...
ppt file
... Both proteins are either belong to the same complex or are parts of the same functional pathway. The same trend is generally true for the larger data set. By manually inspecting the top 100 pairs, the author found that in >95% of them both proteins have similar functions. ...
... Both proteins are either belong to the same complex or are parts of the same functional pathway. The same trend is generally true for the larger data set. By manually inspecting the top 100 pairs, the author found that in >95% of them both proteins have similar functions. ...
Proteins Review - kehsscience.org
... 11. Some amino acids are hydrophilic. What does that mean? It means the amino acids are attracted to ...
... 11. Some amino acids are hydrophilic. What does that mean? It means the amino acids are attracted to ...
In Vitro Reconstitution of SCF Substrate Ubiquitination with Purified
... model system for understanding the mechanisms of protein ubiquitination and degradation. In vitro reconstitution of this process has been used successfully to study how SCF recognizes phosphorylated substrates (Nash et al., 2001; Orlicky et al., 2003; Verma et al., 1997a), the mechanisms of ubiquiti ...
... model system for understanding the mechanisms of protein ubiquitination and degradation. In vitro reconstitution of this process has been used successfully to study how SCF recognizes phosphorylated substrates (Nash et al., 2001; Orlicky et al., 2003; Verma et al., 1997a), the mechanisms of ubiquiti ...
PROTEIN STRUCTURE
... • The hydrophobicity of certain amino acid Rgroups tends to drive them away from the exterior of proteins and into the interior. This driving force restricts the available conformations into which a protein may fold. ...
... • The hydrophobicity of certain amino acid Rgroups tends to drive them away from the exterior of proteins and into the interior. This driving force restricts the available conformations into which a protein may fold. ...
protein structure - MBBS Students Club
... • The hydrophobicity of certain amino acid Rgroups tends to drive them away from the exterior of proteins and into the interior. This driving force restricts the available conformations into which a protein may fold. ...
... • The hydrophobicity of certain amino acid Rgroups tends to drive them away from the exterior of proteins and into the interior. This driving force restricts the available conformations into which a protein may fold. ...
SDS-PAGE_overview
... In our lab, we will be using gradients to 4%-15% acrylamide. Loading dye/ Sample buffer. Since most proteins in solution are transparent, it is difficult to monitor their progress during electrophoresis. For this reason, a visible "tracking dye" such as bromophenol blue is usually added to protein s ...
... In our lab, we will be using gradients to 4%-15% acrylamide. Loading dye/ Sample buffer. Since most proteins in solution are transparent, it is difficult to monitor their progress during electrophoresis. For this reason, a visible "tracking dye" such as bromophenol blue is usually added to protein s ...
new proteins
... Collagen protein is made of 3 polypeptide chains wound together rather like a rope. Hydrogen bonds hold the chains together(second picture from left) Each group of 3 chains is bonded to another group by co-valent bonds called cross links, thus fibrils are formed Many fibrils form a collagen fibre Co ...
... Collagen protein is made of 3 polypeptide chains wound together rather like a rope. Hydrogen bonds hold the chains together(second picture from left) Each group of 3 chains is bonded to another group by co-valent bonds called cross links, thus fibrils are formed Many fibrils form a collagen fibre Co ...
Lecture #2 – Review of Protein Chemistry, Enzyme Specificity
... The various forces that are involved in establishing the structure of enzymes are also clearly involved in determining their specificity (ie, ionic interactions with trypsin, hydrophobic interactions with chymotrypsin and elastase). Only substrates that will fit into the binding pocket can be effect ...
... The various forces that are involved in establishing the structure of enzymes are also clearly involved in determining their specificity (ie, ionic interactions with trypsin, hydrophobic interactions with chymotrypsin and elastase). Only substrates that will fit into the binding pocket can be effect ...
Recombinant Ebola virus VP40 matrix protein
... Novex® Sharp pre-stained protein markers. (B) Western blot detection of EBOV VP40 at 50 ng and 100 ng (lanes 1-2). EBOV VP40 was detected using IBT’s polyclonal antibody at 50 ng/mL (Cat. # 0301-010) and anti-rabbit IgG-HRP conjugate, followed by TMB membrane substrate. ELISA Data: EBOV VP40 ng/well ...
... Novex® Sharp pre-stained protein markers. (B) Western blot detection of EBOV VP40 at 50 ng and 100 ng (lanes 1-2). EBOV VP40 was detected using IBT’s polyclonal antibody at 50 ng/mL (Cat. # 0301-010) and anti-rabbit IgG-HRP conjugate, followed by TMB membrane substrate. ELISA Data: EBOV VP40 ng/well ...
Ubiquitin

Ubiquitin is a small (8.5 kDa) regulatory protein that has been found in almost all tissues (ubiquitously) of eukaryotic organisms. It was discovered in 1975 by Goldstein and further characterized throughout the 1970s and 1980s. There are four genes in the human genome that produce ubiquitin: UBB, UBC, UBA52 and RPS27A.The addition of ubiquitin to a substrate protein is called ubiquitination or ubiquitylation. Ubiquitination can affect proteins in many ways: It can signal for their degradation via the proteasome, alter their cellular location, affect their activity, and promote or prevent protein interactions. Ubiquitination is carried out in three main steps: activation, conjugation, and ligation, performed by ubiquitin-activating enzymes (E1s), ubiquitin-conjugating enzymes (E2s), and ubiquitin ligases (E3s), respectively. The result of this sequential cascade binds ubiquitin to lysine residues on the protein substrate via an isopeptide bond or to the amino group of the protein's N-terminus via a peptide bond.The protein modifications can be either a single ubiquitin protein (monoubiquitination) or a chain of ubiquitin (polyubiquitination). The ubiquitination bonds are always formed with one of the seven lysine residues from the ubiquitin molecule. These 'linking' lysines are represented by a ""K"" (which is the one-letter amino acid notation of lysine) and a number, referring to its position in the ubiquitin molecule. First, a ubiquitin molecule is bonded by its C-terminus to a specific lysine residue (e.g. K48, K29, K63,...) on the target protein. Poly-ubiquitination occurs when the C-terminus of another ubiquitin, will be linked again to a lysine residue (for example again K48 or K29) on the previously added ubiquitin molecule, forming a chain. This process repeats several times, leading to the addition of several ubiquitins. Only poly-ubiquitination on defined lysines, mostly on K48 and K29, is related to degradation with the proteasome (referred to as the ""molecular kiss of death""), while other polyubiquitinations (e.g. on K63, K11, K6) and monoubiquitinations may regulate processes such as endocytic trafficking, inflammation, translation and DNA repair.Lysine 48-linked chains have been much-studied. They are the forms of chains that signal proteins to the proteasome, which destroys and recycles proteins. This discovery won the Nobel Prize for chemistry in 2004.