Overview
... Micro 201 Yuan Lecture 2, Class 24: Protein Folding and Molecular Chaperones April 20th, 2017 Overview The intracellular concentration of protein in bacterial cells can be estimated to be ~135 mg/ml. In this session, we will explore how bacteria employ a suite of molecular machines collectively know ...
... Micro 201 Yuan Lecture 2, Class 24: Protein Folding and Molecular Chaperones April 20th, 2017 Overview The intracellular concentration of protein in bacterial cells can be estimated to be ~135 mg/ml. In this session, we will explore how bacteria employ a suite of molecular machines collectively know ...
About Proteins
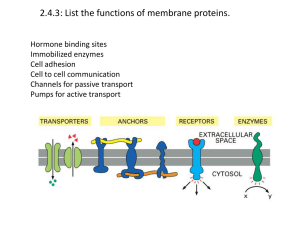
... body The order of the AAs determines the function If even one AA is out of order by mistake, the protein will not function (work) This is because proteins fold in a specific way ...
... body The order of the AAs determines the function If even one AA is out of order by mistake, the protein will not function (work) This is because proteins fold in a specific way ...
Project description
... proteins are the participants of the termination stage. For instance, besides two “canonical” and the well-known termination factors eRF1 and eRF3, in humans we have two others interesting proteins important for termination, Dbp5 and PABP. These proteins have a wide range of activities in the cells, ...
... proteins are the participants of the termination stage. For instance, besides two “canonical” and the well-known termination factors eRF1 and eRF3, in humans we have two others interesting proteins important for termination, Dbp5 and PABP. These proteins have a wide range of activities in the cells, ...
bio98a_l04
... Lambert-Beer Law: OD or Abs = log10 Io/I = -log10 I/Io = e c l e, the molar extinction coefficient, is the optical density (OD) of a material at a given concentration, c, and l is the pathlength of the cuvette. The molar extinction coefficient for Trp is 5,500 M-1 cm-1 at 280 nm, which means a 1 M s ...
... Lambert-Beer Law: OD or Abs = log10 Io/I = -log10 I/Io = e c l e, the molar extinction coefficient, is the optical density (OD) of a material at a given concentration, c, and l is the pathlength of the cuvette. The molar extinction coefficient for Trp is 5,500 M-1 cm-1 at 280 nm, which means a 1 M s ...
Ch. 5. Protein Purification and Characterization Techniques
... Salting Out • After Proteins solubilized, they can be purified based on solubility (usually dependent on overall charge, ionic strength, polarity • Ammonium sulfate (NH4SO4) commonly used to “salt out” ...
... Salting Out • After Proteins solubilized, they can be purified based on solubility (usually dependent on overall charge, ionic strength, polarity • Ammonium sulfate (NH4SO4) commonly used to “salt out” ...
Capturing denaturing proteins * Small Heat Shock Protein substrate
... sHSP chaperone action and interaction with substrates, therefore, has wide-ranging implications for understanding cellular stress and disease processes. We are studying the mechanism of sHSP substrate recognition by identifying specific crosslinking sites between sHSPs and denaturing substrates. sHS ...
... sHSP chaperone action and interaction with substrates, therefore, has wide-ranging implications for understanding cellular stress and disease processes. We are studying the mechanism of sHSP substrate recognition by identifying specific crosslinking sites between sHSPs and denaturing substrates. sHS ...
Improving Function Prediction Using Patterns of Native Disorder in
... Anna Lobley Abstract Instrinsically unstructured (disordered) proteins adopt little or no stable secondary structure in their native state. Proteins containing long disordered regions are abundant within eukaryotic genomes and can be predicted successfully from amino sequence. Disordered regions hav ...
... Anna Lobley Abstract Instrinsically unstructured (disordered) proteins adopt little or no stable secondary structure in their native state. Proteins containing long disordered regions are abundant within eukaryotic genomes and can be predicted successfully from amino sequence. Disordered regions hav ...
Document
... Formulation of hypothesis In silico mutagenesis of staphylococcal nuclease (SNase) and formulation of hypotheses. Basic laboratory technique III: preparation of buffer solutions and determination of pKa values. ...
... Formulation of hypothesis In silico mutagenesis of staphylococcal nuclease (SNase) and formulation of hypotheses. Basic laboratory technique III: preparation of buffer solutions and determination of pKa values. ...
ProteinChipâ technology is one of the most exciting advancements
... ProteinChip technology is one of the most exciting advancements in protein analysis in the last 5 years. The Protein Biology SystemTM (PBS) combines the power of mass analysis with chromatography surfaces on an integrated platform. The PBS can easily be used by biologists, biochemists, and clinicia ...
... ProteinChip technology is one of the most exciting advancements in protein analysis in the last 5 years. The Protein Biology SystemTM (PBS) combines the power of mass analysis with chromatography surfaces on an integrated platform. The PBS can easily be used by biologists, biochemists, and clinicia ...
Typical IP Protocol
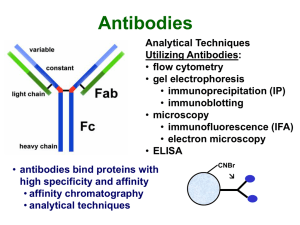
... Antibodies Analytical Techniques Utilizing Antibodies: • flow cytometry • gel electrophoresis • immunoprecipitation (IP) • immunoblotting • microscopy • immunofluorescence (IFA) • electron microscopy • ELISA • antibodies bind proteins with high specificity and affinity • affinity chromatography • an ...
... Antibodies Analytical Techniques Utilizing Antibodies: • flow cytometry • gel electrophoresis • immunoprecipitation (IP) • immunoblotting • microscopy • immunofluorescence (IFA) • electron microscopy • ELISA • antibodies bind proteins with high specificity and affinity • affinity chromatography • an ...
custom protein production service
... CUSTOM PROTEIN PRODUCTION SERVICE Highly specialized custom production service Our experience in recombinant protein production for your research! ...
... CUSTOM PROTEIN PRODUCTION SERVICE Highly specialized custom production service Our experience in recombinant protein production for your research! ...
Protein purification
Protein purification is a series of processes intended to isolate one or a few proteins from a complex mixture, usually cells, tissues or whole organisms. Protein purification is vital for the characterization of the function, structure and interactions of the protein of interest. The purification process may separate the protein and non-protein parts of the mixture, and finally separate the desired protein from all other proteins. Separation of one protein from all others is typically the most laborious aspect of protein purification. Separation steps usually exploit differences in protein size, physico-chemical properties, binding affinity and biological activity. The pure result may be termed protein isolate.The methods used in protein purification can roughly be divided into analytical and preparative methods. The distinction is not exact, but the deciding factor is the amount of protein that can practically be purified with that method. Analytical methods aim to detect and identify a protein in a mixture, whereas preparative methods aim to produce large quantities of the protein for other purposes, such as structural biology or industrial use. In general, the preparative methods can be used in analytical applications, but not the other way around.