* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Supplementary Text 2: Extensions to the prototype model
Survey
Document related concepts
Multi-state modeling of biomolecules wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Citric acid cycle wikipedia , lookup
Butyric acid wikipedia , lookup
Basal metabolic rate wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
Specialized pro-resolving mediators wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
Biochemistry wikipedia , lookup
Glyceroneogenesis wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Biosynthesis wikipedia , lookup
Lipid signaling wikipedia , lookup
Transcript
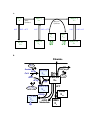
Supplementary Text 2: Extensions to the prototype model. The prototype model1 was developed in order to integrate major pathway components with other pertinent information on sphingolipid metabolism. As typical in modeling, this model design was based on abstraction and simplification. In order to facilitate the direct experimental testing of model predictions, several issues peculiar to yeast sphingolipid metabolism became important and suggested further refinements of the model. Complex sphingolipids. The complex sphingolipids inositolphosphoceramide (IPC), mannosylinositolphosphoceramide (MIPC), and mannosyl(inositol phosphate)2ceramide (M(IP)2C) are metabolized in different compartments, move between these compartments, and the accumulate over time. To capture this in the model, IPC, MIPC and M(IP)2C were elevated from independent to dependent variables, and their pools were split into plasma membrane and other compartments. The ratio of this split was determined by accounting for the contribution of membrane synthesis to cell growth, which is directly accompanied by a net positive flux of IPC, MIPC, and M(IP)2C to the plasma membrane (Fig. 1 and Supplementary Fig. 1A). It should be noted that other possible compartmentalizations were not included due to paucity of data. The relative amount of lipids in each component was determined based on a report by Patton and Lester3. A concentration of 90% of complex sphingolipids for the plasma membrane was used with the remaining 10% present in other organelles. The values for the fluxes were estimated from Wu et al. (1995, Fig. 7)4 . The flux towards the plasma membrane was based on the 30 min pulse-label experiment. The recycling flux percentage was calculated by a comparison of complex sphingolipid concentrations during exponential growth versus steady state after five generations growth (~90 min per generation). The transfer of complex sphingolipids into compartments inaccessible to IPCase is consistent with results showing that ISC1 is not in the plasma membrane5. Instead, the enzyme appears to move from the endoplasmic reticulum, where it is in early log phase, to the mitochondria, where it is found in late phase. This suggests that the calculation of percentages in plasma membrane, as taken from Patton and Lester3, is appropriate. Fatty Acid Synthesis and Elongation. Another model refinement concerned the availability and metabolism of fatty acids at their metabolic entry points into sphingolipid synthesis at serine palmitoyl transferase (SPT) and ceramide synthase. Supplementary Fig. 1B shows the expansion of fatty acid synthesis of SPT substrate and elongation to the C26-CoAs necessary for ceramide acylation. Kobayashi and Nagiec6 showed the importance of very long chain fatty acids (VLCFAs) in regulating the flow of de novo sphingolipid synthesis towards ceramide, complex sphingolipids, and free sphingoid bases and their phosphates. Modeling this observation requires the inclusion of C26-CoA as a dynamically changing (rather than constant input) variable. The elongation of fatty acids beyond C16/C18 proceeds two carbons at a time, and to model this repetitive elongation process, ELO1p alone was used to represent the independent variable for the catalysis of the elongation of fatty acids to the C26-CoA product. This decision was based on two factors. First, it has been shown in rat7,8 and swine9 that this condensation step is at least one of the rate-limiting steps in the overall elongation of very-long-chain fatty acyl-CoA. In yeast, it has been shown that mutants with disruption of ELO1 and FAS2 must be supplied with fatty acids of at least 16 carbons to grow10, indicating the importance of this initial elongation step. Second, from the modeling perspective, a linear process can be collapsed from several steps to one without penalty, except for possible dynamic inaccuracies caused by time delays between steps11. Sphingosine-phosphate flux. An equally important parameter concerns the ultimate breakdown of sphingolipids through the action of the lyase on long chain base phosphates. The computation of fluxes to and from the sphingosine-phosphate pools was accomplished by taking into consideration that, in vivo, the lyase functions as the major clearance pathway for the long chain base phosphates (LCB-Ps). Thus, the effluxes from the LCB-P pool were partitioned with the majority (90%) processed through the lyase and 10% through the phosphatase. Using stoichiometric arguments, the specific values of the fluxes were then derived from the kinase flux, which was based on Michaelis-Menten kinetics with ATP as secondary substrate (Supplementary Fig. 1C). A. IPC-g X8 0.02 MIPC-g X18 MIPC Synthase 0.02 0.75 M(IP)2C-g X19 M(IP)2C Synthase 0.02 0.75 0.02 0.01 PI X15 DAG X14 M(IP)2C-m X22 MIPC-m X21 IPC-m X20 B. Palmitate X12 Acetate, ACSp CoA ATP X12, X23 C26-CoA X23 G3P Acylt. Ac-CoA X25 ACCp 0.01 0.5 FAS Mal-CoA X24 Pal-CoA X12 SPT Serine X13 ELO1p ACBP, Metabolism X10 KDHS X1 C. 0.000011 0.000107 DHS-1-P DHS 0.00011 Pal-CoA CDP-Eth 0.01857 PHS-1-P PHS 0.00185 0.0167 Supplementary Figure 1. Model development. A) Fluxes between complex sphingolipid components. B) Fatty acid metabolism. C) Fluxes into and out of the LCB phosphates. Effect of CDP-DAG and PI on phosphatidate phosphatase. We included the allosteric regulation of phosphatidate phosphatase by phospholipids. Wu and Carman (1996. Figures 4A and 5A)2 report for yeast a positive effect of cardiolipin (CL), CDP-diacylglycerol (CDP-DG), phosphatidylinositol (PI), and to a minor degree of phosphatidylglycerol (PG) and phosphatidylserine (PS), on PAPPase activity. We computed the corresponding kinetic orders of the model variables CDP-DG and PI. As recommended in the literature 12, we used a log-log representation (see Supplementary Figure 2) of the phospholipid concentrations vs. PA-PPase activity at 3 mol% PA. Also, since we are interested in the response of sphingolipid metabolism to acute perturbations, we eliminated the long-term regulation of PA-PPase by inositol13,14 that was included in the prototype model1. 1.2 Log (Relative PA-PPase activity) PA-PPase activity (U/mg) 12 10 8 y = 0.3269x + 0.9877 1.1 1 0.9 0.8 y = 0.3038x + 0.7254 0.7 0.6 0.5 6 0 1 2 3 4 Phospholipid (mol%) 5 6 -0.4 -0.2 0 0.2 0.4 0.6 0.8 Log (Phospholipid (mol%)) Supplementary Figure 2: Dependence of relative PA-PPase activity on concentration of phospholipids CDP-DAG () and PI (). Left panel: Original data, redrawn for 3 mol% PA from Wu and Carman (1996) 2. Right panel: Data shown in logarithmic coordinates, where the kinetic orders are determined as slopes of the linear regression. 1 References 1. 2. 3. 4. 5. 6. 7. 8. 9. 10. 11. 12. 13. 14. Alvarez-Vasquez, F., Sims, K. J., Hannun, Y. A. & Voit, E. O. Integration of kinetic information on yeast sphingolipid metabolism in dynamical pathway models. Journal Of Theoretical Biology 226, 265-291 (2004). Wu, W. I. & Carman, G. M. Regulation of phosphatidate phosphatase activity from the yeast Saccharomyces cerevisiae by phospholipids. Biochemistry 35, 3790-6 (1996). Patton, J. L. & Lester, R. L. The phosphoinositol sphingolipids of Saccharomyces cerevisiae are highly localized in the plasma membrane. J Bacteriol 173, 3101-8 (1991). Wu, W. I. et al. Regulation of lipid biosynthesis in Saccharomyces cerevisiae by fumonisin B1. J Biol Chem 270, 13171-8 (1995). Vaena de Avalos, S., Okamoto, Y. & Hannun, Y. A. Activation and localization of inositol phosphosphingolipid phospholipase C, Isc1p, to the mitochondria during growth of Saccharomyces cerevisiae. J Biol Chem 279, 11537-45 (2004). Kobayashi, S. D. & Nagiec, M. M. Ceramide/Long-Chain Base Phosphate Rheostat in Saccharomyces cerevisiae: Regulation of Ceramide Synthesis by Elo3p and Cka2p. Eukaryot Cell 2, 284-94 (2003). Prasad, M., Nagi, M., Ghesquier, D., Cook, L. & Cinti, D. Evidence for multiple condensing enzymes in rat hepatic microsomes catalyzing the condensation of saturated, monounsaturated, and polyunsaturated acyl coenzyme A. J Biol Chem. 261, 8213-7 (1986). Bernert, J. J. & Sprecher, H. An analysis of partial reactions in the overall chain elongation of saturated and unsaturated fatty acids by rat liver microsomes. J Biol Chem. 252, 6736-44 (1977). Yoshida, S. & Takeshita, M. Comparison between condensation and overall chain elongation of arachidoyl-CoA and arachidonoyl-CoA in swine cerebral microsomes. Arch Biochem Biophys. 254, 180-7 (1987). Toke, D. A. & Martin, C. E. Isolation and characterization of a gene affecting fatty acid elongation in Saccharomyces cerevisiae. J Biol Chem 271, 18413-22 (1996). Curto, R., Voit, E. O., Sorribas, A. & Cascante, M. Mathematical models of purine metabolism in man. Math Biosci 151, 1-49 (1998). Voit, E. O. Computational Analysis of Biochemical Systems. A Practical Guide for Biochemists and Molecular Biologists. (Cambridge University Press, Cambridge, UK, 2000). Morlock, K. R., McLaughlin, J. J., Lin, Y. P. & Carman, G. M. Phosphatidate phosphatase from Saccharomyces cerevisiae. Isolation of 45- and 104-kDa forms of the enzyme that are differentially regulated by inositol. J Biol Chem 266, 358693 (1991). Carman, G. M. Phosphatidate phosphatases and diacylglycerol pyrophosphate phosphatases in Saccharomyces cerevisiae and Escherichia coli. Biochim Biophys Acta 1348, 45-55 (1997).