* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Bioinfo primer - part 6/6
RNA interference wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Pharmacogenomics wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Non-coding RNA wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Nutriepigenomics wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Transposable element wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Epitranscriptome wikipedia , lookup
Genomic imprinting wikipedia , lookup
Whole genome sequencing wikipedia , lookup
Ridge (biology) wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Oncogenomics wikipedia , lookup
Genome (book) wikipedia , lookup
Metabolic network modelling wikipedia , lookup
Microevolution wikipedia , lookup
Designer baby wikipedia , lookup
Primary transcript wikipedia , lookup
History of genetic engineering wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Gene expression profiling wikipedia , lookup
Human genome wikipedia , lookup
Human Genome Project wikipedia , lookup
Helitron (biology) wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Genomic library wikipedia , lookup
Genome editing wikipedia , lookup
Minimal genome wikipedia , lookup
Non-coding DNA wikipedia , lookup
Metagenomics wikipedia , lookup
Pathogenomics wikipedia , lookup
Genome evolution wikipedia , lookup
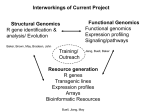
Christophe Roos - MediCel ltd [email protected] High throughput data acquisition New issues in storage and analysis Annotating genomes with functional information: automatic but without errors? Genome annotation • Annotations is the sum of all non-sequence information that can be connected to any sequence Phylogenetic inference Metabolic profiles Sequence homologs in other genomes Connectors to other maps Cofactors and metabolites Metabolic map locator Gene Sequence Functional chemistry Experimental data Genome location Expression info Raw images Numerical values Cluster genes Christophe Roos - 6/6 Functional genomics Structure Raw data Electron density Structure annotation Spring 2002 SS assignments Genome annotation • Primary sources of information about what genes do are laboratory experiments. It may take several experiments for one data point. • All that data should ideallically be associated – hyperlinked among DBs. – Magpie is an environment for genome annotation • Compare genomes to learn how their structure affects function – Bacteria have modules of genes functioning together organised in ‘operons’ – Higher organisms need to pack the DNA to fit it in the nucleus. Activating a gene means unpacking and is not efficient if it is done for each gene separately Christophe Roos - 6/6 Functional genomics Spring 2002 Functional genomics • High throughput technologies give us long lists of the parts of systems (chromosomes, genomes, cells, etc). We can now analyse how they work together to produce the complexity of the organisms. • The function of the genome is – Metabolism: metabolic pathways convert chemical energy derived from food into useful work in the cell. – Regulation: regulatory pathways are biochemical mechanisms that control what genomic DNA does. It switches genes on and off in a controlled way. – Signalling: signalling pathways control the movement of information (chemicals) from one component to another on many levels – Construction • Functional genomics tries to map these pathways Christophe Roos - 6/6 Functional genomics Spring 2002 Analysing the activity of the genome • Genomics: look at transcriptional activity of genes – Transcription: When a gene is transcriptionally active, it means that messenger RNA (mRNA) is synthesised. The amount of mRNA from each active gene varies over time. – Turnover: Different mRNA species have different half-lives. – Translation: When a mRNA is produced, it does not imply that the corresponding protein is translated. Transcripts can also be produced for storage and later use. – Technically feasible: it is possible to isolate all mRNAs from cells and to quantitate it within certain limits. • Proteomics: look at proteins instead of transcripts – Limited: Presently acceptable efficiency comes at the expenses of incufficient quality – Closer to ’reality’ since the proteins are the players Christophe Roos - 6/6 Functional genomics Spring 2002 EST: Expressed sequence tags • ESTs are partial sequences of cDNA clones. cDNA clones are DNA synthesised in vitro using mRNA as template. – Why? cDNA is more stable than mRNA – How? cDNA can be made ‘en masse’ starting from total cellular mRNA isolates. cDNA libraries are specific for tissue, developmental time, stimulation etc. – Therefore, looking at cDNA is looking at mRNA is looking at active genes. – To look at cDNA means sequencing (part of) it. • Clones are picked at random (10’000-200’000) • Sequenced from one or both ends once (no proofreading) • Sequences entered into EST sequence databases Christophe Roos - 6/6 Functional genomics Spring 2002 EST: Expressed sequence tags • • • • constucting a clone by inserting a piece of DNA into a ’vector’. the vector and its insert will behave as an independent unit (’plasmid’) in the bacterial host and carries some additional genes to allow for selection (only those bacterial with the vector will survive on antibiotics) Amplify and sequence Iterate (in parallell) Christophe Roos - 6/6 Functional genomics Spring 2002 DNA hybridisation • DNA is a double-helix and can be separated by denaturing treatment into two strands. Each strand becomes ’sticky’ and attempts to renature with homologous single-strand sequences to form hybrids. • Single-strand DNA from all known genes of a given species can be attached to a matrix, then probed with labelled cDNA molecules from a given sample. Only complementary probes will hybridise and can be detected if they have been previously labelled (radioactivity, fluorescent stain, ...) • The technique can be multiplexed: – High density arrays carrying sticky probes from a full genome – Parallel hybridisation with cDNA from various sources Christophe Roos - 6/6 Functional genomics Spring 2002 The process of using microarrays Building the Chip: PCR PURIFICATION and PREPARATION MASSIVE PCR PREPARING SLIDES PRINTING Preparing RNA: CELL CULTURE AND HARVEST Hybridising the Chip: POST PROCESSING ARRAY HYBRIDIZATION RNA ISOLATION DATA ANALYSIS cDNA PRODUCTION Christophe Roos - 6/6 Functional genomics PROBE LABELING Spring 2002 The output: the image raw data cDNA is prepared from two samples (in this example) and labelled, each sample with a distinct color. Then the array is hybridised with the doubble probe and the signal is recorded as images overlay images and normalise scanning laser 2 laser 1 emission Christophe Roos - 6/6 Functional genomics analysis Spring 2002 Problems in image analysis • Noise • Spot detection and intensity • Alignment if overlay Christophe Roos - 6/6 Functional genomics Spring 2002 A set of experiments on yeast... • Each row represents one gene • Each column represents one experiment – The columns have been organised into related sets of experiments (ALPH, ELU,...) • The colors indicate gene activity (from high to absent) Christophe Roos - 6/6 Functional genomics Spring 2002 Clustering the resulting data • Looking at 10’000 genes is not easy • Group genes into clusters of genes that behave the same way over a set of several experiments – – – – Hierarchical clustering K-means clustering Self-organising maps (SOM) Etc. Christophe Roos - 6/6 Functional genomics Spring 2002 The overall process with microarrays • Microarray data has to be used in a larger frame of experimentation Christophe Roos - 6/6 Functional genomics Spring 2002 Making a model of the data Sequence Interaction Genome 1. 2. 3. Elements Binary relations Networks Christophe Roos - 6/6 Functional genomics Structure Network Transcriptome Function Function Proteome Assembly Neighbour Cluster Pathway Genome Hierarchical Tree Spring 2002 Comparing networks Pathway vs. Pathway • Gain new biological information by comparison of networks • What is the metrics? • How is it done? Is it simply a problem of graph isomorphism Pathway vs. Genome Genome vs. Genome Cluster vs. Pathway Christophe Roos - 6/6 Functional genomics Spring 2002 Biological graph comparison • Search heuristically for clusters of correspondence Graph 1 A C D B G E I Correspondences K H F J A C D E I H A B C D . . . . a b c d . . Graph 2 a d i h K f j a c i j Spring 2002 k h F b g e J Christophe Roos - 6/6 Functional genomics k d G b g e Clustering algorithm B c f Example: genomic, metabolic, structural Genome-pathway comparison, which reveals the correlation of physical coupling of genes in the genome - operon structure (a) and functional coupling (b) of gene products in the pathway E. coli genome hisL hisG hisD yefM Christophe Roos - 6/6 Functional genomics hisC hisB hisH hisA hisF hisI yzzB Spring 2002 Example: genomic, metabolic, structural HISTIDINE METABOLISM Pentose phosphate cycle 5P-D-1-ribulosylformimine 3.5.1.- Phosphoribosyl-AMP PRPP 3.6.1.31 2.4.2.17 3.5.4.19 PhosphoribulosylFormimino-AICARP 2.4.2.- 5.3.1.16 PhosphoribosylFormimino-AICAR-P Phosphoriboxyl-ATP 2.6.1.- Imidazoleacetole P 2.6.1.9 4.2.1.19 ImidazoleGlicerol-3P 3.1.3.15 L-Histidinol-P 5P Ribosyl-5-amino 4Imidazole carboxamide (AICAR) 1.1.1.23 1-MethylL-histidine 3.4.13.5 Aneserine 6.3.2.11 Purine metabolism 2.1.1.- 6.3.2.11 3.4.13.3 3.5.3.5 Imidazolone acetate 3.5.2.- Imidazole4-acetate 1.14135 Christophe Roos - 6/6 Functional genomics 3.4.13.2 0 Imidazole acetaldehyde 1.2.1.3 Histamine 1.4.3.6 L-Hisyidinal 2.1.1.22 Carnosine N-Formyl-Lspartate L-Hisyidinal 1.1.1.23 6.1.1 Hercyn 4.1.1.22 4.1.1.28 L-Histidine Spring 2002 Example: genomic, metabolic, structural SCOP hierarchical tree……..NE, TYROSINE AND TRYPTOPHAN BIOSYNTHESIS 1. 2. 3. All alpha All beta Alpha and beta (a/b) 3.1 beta/alpha (TIM)-barrel 3.2 Cellulases . . . . . . . 3.74 Thiolase 3.75 Cytidine deaminase 4. Alpha and beta (a+b) 5. Multi-domain (alpha and beta) 6. Membrane and cell surface pro 7. Small proteins RNA 8. Peptides 9. Designed proteins 10. Non-protein 2.5.1.19 3-deoxyD-arabinoheptonate 1.1.1.24 1.3.1.43 4.2.1.51 4.2.1.10 4.2.1.11 1.1.9925 2.6.1.57 Pretyrosine 4.2.1.91 1.4.1.20 6.1.1.20 2.6.1.5 Phenylalanine Phenylpyruvate 2.6.1.1 2.6.1.9 2.6.1.57 4.1.3.27 Histidine 1.1.9925 2.6.1.9 2.6.1.57 4-Aminobenzoate 2.6.1.5 2.6.1.9 2.6.1.57 Prephenate 4.2.1.51 Indole 5.4.99.5 2.4.2.18 N-(5-Phosphob-v-ribosyl)anthranilate 4.2.1.20 5.3.1.24 4.1.1.48 1-(2- CarboxyPhenylamino)1-deoxy-D-ribulose 5-phosphate 4.2.1.20 (3-Indolyl)Glycerol phosphate L-Tryptophan Tryptophan metabolism Ubiquinone biosynthesis 3-Dehydro- Protocatechuate shikimate Folate biosynthesis Christophe Roos - 6/6 Functional genomics 1.4.3.2 2.6.1.1 4.2.1.91 4.1.3.- 4.2.1.10 2.6.1.5 4.2.1.20 1.4.3.2 4.6.1.4 2.6.1.1 4-Hydroxyphenylpyruvate 1.14.16.1 Shikimate 1.1.1.25 Alkaloid biosynthesis I 6.1.1.1 Tyrosine Anthranilate 4.6.1.3 3-Dehydroquinate Tyr-tRNA Chorismate 2.7.1.71 Tyrosine metabolism Spring 2002 More challenges? The list of genes being activated or inactivated or that are unaffected when comparing two samples becomes more informative if the genes can be mapped onto maps from which functions can be deduced. Christophe Roos - 6/6 Functional genomics Spring 2002 More challenges? Christophe Roos - 6/6 Functional genomics Spring 2002