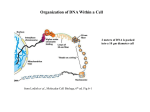
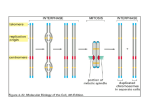
* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Inquiry into Life Twelfth Edition
Short interspersed nuclear elements (SINEs) wikipedia , lookup
DNA vaccination wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
DNA damage theory of aging wikipedia , lookup
Genome evolution wikipedia , lookup
Molecular cloning wikipedia , lookup
Epigenetics of cocaine addiction wikipedia , lookup
Genome (book) wikipedia , lookup
Gene expression profiling wikipedia , lookup
Neocentromere wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Genomic imprinting wikipedia , lookup
Deoxyribozyme wikipedia , lookup
DNA supercoil wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
Minimal genome wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Point mutation wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Epigenetics of depression wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Designer baby wikipedia , lookup
Microevolution wikipedia , lookup
Ridge (biology) wikipedia , lookup
Helitron (biology) wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
History of genetic engineering wikipedia , lookup
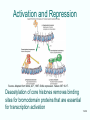
Non-coding DNA wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Transcription factor wikipedia , lookup
Epigenetics in stem-cell differentiation wikipedia , lookup
Epigenetics wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Primary transcript wikipedia , lookup
Epigenomics wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
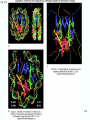
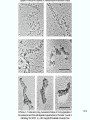
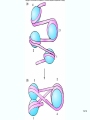
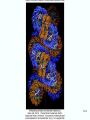
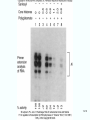
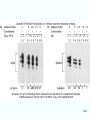
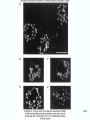
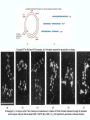
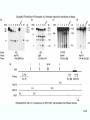
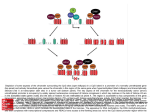
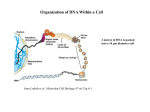
Molecular Biology Lecture 20 Chapter 13 Chromatin Structure and Its Effects on Transcription Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Histones • Eukaryotic cells contain 5 kinds of histones – – – – – H1 H2A H2B H3 H4 Source: Panyim and Chalkley, Arch. Biochem. 13-2 & Biophys. 130, 1969, f. 6A, p.343. Properties of Histones • Pronounced positive charge at neutral pH • Most are well-conserved from one species to another • Not single copy genes, repeated many times – Some copies are identical – Others are quite different – H4 has only had 2 variants ever reported 13-3 Nucleosomes • First order of folding is the nucleosome – X-ray diffraction has shown strong repeats of structure at 100Å intervals – This spacing approximates the nucleosome spaced at 110Å intervals 13-4 Histones in the Nucleosome • Chemical cross-linking in solution: – H3 to H4 – H2A to H2B • H3 and H4 exist as a tetramer (H3-H4)2 • Chromatin is composed of roughly equal masses of DNA and histones – Corresponds to 1 histone octamer per 200 bp of DNA – Octamer composed of: • 2 each H2A, H2B, H3, H4 • 1 each H1 13-5 Fig. 13.4 13-6 H1 and Chromatin • Treatment of chromatin with trypsin or high salt buffer removes histone H1 • This treatment leaves chromatin looking like “beads-on-a-string” • The beads named nucleosomes – Core histones form a ball with DNA wrapped around the outside – H1 also lies on the outside of the nucleosome 13-7 Nucleosome Structure • Central (H3-H4)2 core attached to H2AH2B dimers • Grooves on surface define a left-hand helical ramp – a path for DNA winding – DNA winds almost twice around the histone core condensing DNA length by 6- to 7-X – Core histones contain a histone fold: • 3 a-helices linked by 2 loops • Extended tail of about 28% of core histone mass • Tails are unstructured 13-8 The 30-nm Fiber • Second order of chromatin folding produces a fiber 30 nm in diameter – The string of nucleosomes condenses to form the 30-nm fiber in a solution of increasing ionic strength – This condensation results in another six- to seven-fold condensation of the nucleosome itself • Four nucleosomes condensing into the 30nm fiber form a zig-zag structure 13-9 13-10 Formation of the 30-nm Fiber • Two stacks of nucleosomes form a lefthanded helix – Two helices of polynucleosomes – Zig-zags of linker DNA • Role of histone H1? – 30-nm fiber can’t form without H1 – H1 crosslinks to other H1 more often than to core histones 13-11 13-12 13-13 Higher Order Chromatin Folding • 30-nm fibers account for most of chromatin in a typical interphase nucleus • Further folding is required in structures such as the mitotic chromosomes • Model favored for such higher order folding is a series of radial loops Source: Adapted from Marsden, M.P.F. and U.K. Laemmli, Metaphase chromosome structure: Evidence of a radial loop model. Cell 17:856, 13-14 1979. Chromatin Structure and Gene Activity • Histones, especially H1, have a repressive effect on gene activity in vitro • Two families of 5S rRNA genes studied are oocyte and somatic genes – Oocyte genes are expressed only in oocytes – Somatic genes are expressed both in oocytes and somatic cells – Somatic genes form more stable complexes with transcription factors 13-15 Transcription Factors and Histones Control the 5S rRNA • Genes active by TFIIIs preventing formation of nucleosome stable complexes with internal control region • Stable complexes require histone H1 and exclude TFIIIs once formed so that genes are repressed 13-16 Effects of Histones on Transcription of Class II Genes • Core histones assemble nucleosome cores on naked DNA • Transcription of reconstituted chromatin with an average of 1 nucleosome / 200 bp DNA exhibits 75% repression relative to naked DNA • Remaining 25% is due to promoter sites not covered by nucleosome cores 13-17 13-18 Histone H1 and Transcription • Histone H1 causes further repression of template activity, in addition to that of core histones • H1 repression can be counteracted by transcription factors • Sp1 and GAL4 act as both: – Antirepressors preventing histone repressions – Transcription activators • GAGA factor: – Binds to GA-rich sequences in the Krüppel promoter – An antirepressor – preventing repression by histones 13-19 13-20 Model of Transcriptional Activation Source: Adapted from Laybourn, P.J. and J. T. Kadonaga, Role of nucleosomal cores and histone H1 in regulation of transcription by polymerase II. Science 254:243, 1991. 13-21 Nucleosome Positioning • Model of activation and antirepression asserts that transcription factors can cause antirepression by: – Removing nucleosomes that obscure the promoter – Preventing initial nucleosome binding to the promoter 13-22 Nucleosome-Free Zones • Nucleosome positioning would result in nucleosome-free zones in the control regions of active genes • Assessment in a circular chromosome can be difficult without some type of marker 13-23 13-24 13-25 Detecting DNaseHypersensitive Regions • Active genes tend to have DNase-hypersensitive control regions • Part of this hypersensitivity is due to absence of nucleosomes 13-26 13-27 13-28 Histone Acetylation • Histone acetylation occurs in both cytoplasm and nucleus • Cytoplasmic acetylation carried out by HAT B (histone acetyltransferase, HAT) – Prepares histones for incorporation into nucleosomes – Acetyl groups later removed in nucleus • Nuclear acetylation of core histone N-terminal tails – Catalyzed by HAT A – Correlates with transcription activation – Coactivators of HAT A found which may allow loosening of association between nucleosomes and gene’s control region – Attracts bromodomain proteins, essential for transcription 13-29 Histone Deacetylation • Transcription repressors bind to DNA sites and interact with corepressors which in turn bind to histone deacetylases – Repressors • Unliganded nuclear receptors • Mad-Max – Corepressors • NCoR/SMRT • SIN3 – Histone deacetylases - HDAC1 and 2 13-30 Ternary Protein Complexes • Assembly of complex brings histone deacetylases close to nucleosomes • Deacetylation of core histones allows – Histone basic tails to bind strongly to DNA, histones in neighboring nucleosomes – This inhibits transcription 13-31 Activation and Repression Source: Adapted from Wolfe, A.P., 1997. Sinful repression. Nature 387:16-17. Deacetylation of core histones removes binding sites for bromodomain proteins that are essential for transcription activation 13-32