* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Document
Molecular cloning wikipedia , lookup
Gene expression profiling wikipedia , lookup
Non-coding RNA wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
United Kingdom National DNA Database wikipedia , lookup
DNA barcoding wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
DNA vaccination wikipedia , lookup
Pathogenomics wikipedia , lookup
Transposable element wikipedia , lookup
History of RNA biology wikipedia , lookup
History of genetic engineering wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Designer baby wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Genetic code wikipedia , lookup
Genomic library wikipedia , lookup
Genome evolution wikipedia , lookup
Primary transcript wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Computational phylogenetics wikipedia , lookup
Microevolution wikipedia , lookup
Human genome wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Microsatellite wikipedia , lookup
Non-coding DNA wikipedia , lookup
Point mutation wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Metagenomics wikipedia , lookup
Genome editing wikipedia , lookup
Smith–Waterman algorithm wikipedia , lookup
Helitron (biology) wikipedia , lookup
Multiple sequence alignment wikipedia , lookup
Lecture 2: Introduction to
Computational Biology
Alexei Drummond
Outline
•
•
•
•
•
CS369 2007
Sequences and sequence databases
Similarity and Homology
Sequence alignment
Dot plots
Database searches for similar sequences
2
Sequence
• Definition: A sequence S is an ordered set
of n characters (si) representing nucleotides
or amino acids. S = {s1, s2,…,sn-1 , sn}
– DNA is composed of four nucleotides or
bases: si {A, C, G, T}
– RNA is composed of four nucleotides:
si {A, C, G, U}(T is transcribed as U)
– Proteins are composed of twenty amino acids
CS369 2007
3
Biomolecular sequences
DNA
5’-ACGATCGACTGGTATATCGATGCT-3’
Xi {A,C,G,T}
RNA
Protein
CS369 2007
5’-ACGAUCGACUGGUAUAUCGAUGCU-3’
Xi {A,C,G,U}
MFINRWLFSTNHKDIGTLYLLFGAW
Xi {A,R,N,D,C,E,Q,G,H,I,L,K, M,F,P,S,T,W,Y,V}
4
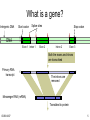
What is a gene?
Intergenic DNA
DNA
Start codon
Splice sites
Stop codon
5’
3’
3’
5’
Exon 1 Intron 1
Exon 2
Intron 2
Exon 3
Both the exons and introns
are transcribed
Primary RNA
transcript
5’
3’
The introns are
removed
Messenger RNA (mRNA)
Translated to protein
CS369 2007
5
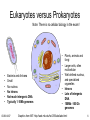
Eukaryotes versus Prokaryotes
Note: There is no cellular biology in the exam!
•
•
•
•
•
•
•
•
Bacteria and Archaea
Small
No nucleus
No introns
Not much intergenic DNA
Typically 1-10Mb genomes
CS369 2007
•
•
•
•
Plants, animals and
fungi
Larger cells, often
multicellular
Well defined nucleus,
and specialized
organelles
Introns
Lots of intergenic
DNA
100Mb -100 Gb
genomes
Graphics from MIT: http://web.mit.edu/hst.035/labs/labs.html
6
Sequence databases
• Where do biologists store their data?
– Databases
• Public, private proprietary
• General, specialist
– Hard drive
• Chromatograms/Electropherograms
• Flat file sequence formats
– Fasta, Genbank et cetera
• Flat file alignment formats
– Nexus, ClustalX, GCG et cetera
CS369 2007
7
CS369 2007
8
NCBI Nucleotide database
CS369 2007
9
Searching by accession number
CS369 2007
10
Genbank record
CS369 2007
11
Genbank headers
LOCUS
DEFINITION
ACCESSION
VERSION
KEYWORDS
SOURCE
ORGANISM
REFERENCE
AUTHORS
TITLE
JOURNAL
PUBMED
CS369 2007
X00166
711 bp
DNA
linear
PHG 10-FEB-1999
Bacteriophage lambda cI gene encoding the repressor protein for
transcriptional control of tetracycline resistance on plasmid pTR
262.
X00166
X00166.1 GI:15056
repressor; tetracycline resistance.
Enterobacteria phage lambda
Enterobacteria phage lambda
Viruses; dsDNA viruses, no RNA stage; Caudovirales; Siphoviridae;
Lambda-like viruses.
1 (bases 1 to 711)
Nilsson,B., Uhlen,M., Josephson,S., Gatenbeck,S. and Philipson,L.
An improved positive selection plasmid vector constructed by
oligonucleotide mediated mutagenesis
Nucleic Acids Res. 11 (22), 8019-8030 (1983)
6316281
12
Genbank feature table
FEATURES
source
CDS
CS369 2007
Location/Qualifiers
1..711
/organism="Enterobacteria phage lambda"
/mol_type="genomic DNA"
/db_xref="taxon:10710"
1..>711
/note="unnamed protein product; coding sequence cI gene"
/codon_start=1
/transl_table=11
/protein_id="CAA24991.1"
/db_xref="GI:15057"
/db_xref="GOA:P03034"
/db_xref="InterPro:IPR001387"
/db_xref="InterPro:IPR006198"
/db_xref="InterPro:IPR010982"
/db_xref="InterPro:IPR011056"
/db_xref="PDB:1F39"
/db_xref="PDB:1GFX"
/db_xref="PDB:1J5G"
/db_xref="PDB:1LLI"
/db_xref="PDB:1LMB"
/db_xref="PDB:1LRP"
…
13
Genbank sequence
ORIGIN
1
61
121
181
241
301
361
421
481
541
601
661
atgagcacaa
gcaatttatg
atggggatgg
tataacgccg
atcgccagag
gagtatgagt
acctttacca
gcattctggc
tttcctgacg
tgcatagcca
caggtgtttt
tccgttgtgg
aaaagaaacc
aaaaaaagaa
ggcagtcagg
cattgcttgc
aaatctacga
accctgtttt
aaggtgatgc
ttgaggttga
gaatgttaat
gacttggggg
tacaaccact
ggaaagttat
attaacacaa
aaatgaactt
cgttggtgct
aaaaattctc
gatgtatgaa
ttctcatgtt
ggagagatgg
aggtaattcc
tctcgttgac
tgatgagttt
aaacccacag
cgctagtcag
gagcagcttg
ggcttatccc
ttatttaatg
aaagttagcg
gcggttagta
caggcaggga
gtaagcacaa
atgaccgcac
cctgagcagg
accttcaaga
tacccaatga
tggcctgaag
aggacgcacg
aggaatctgt
gcatcaatgc
ttgaagaatt
tgcagccgtc
tgttctcacc
ccaaaaaagc
caacaggctc
ctgttgagcc
aactgatcag
tcccatgcaa
agacgtttgg
tcgccttaaa
cgcagacaag
attaaatgct
tagcccttca
acttagaagt
tgagcttaga
cagtgattct
caagccaagc
aggtgatttc
ggatagcggt
tgagagttgt
c
//
CS369 2007
14
Fasta format
>gi|15056|emb|X00166.1| Bacteriophage lambda cI gene encoding the…
ATGAGCACAAAAAAGAAACCATTAACACAAGAGCAGCTTGAGGACGCACGTCGCCTTAAAGCAATTTATG
AAAAAAAGAAAAATGAACTTGGCTTATCCCAGGAATCTGTCGCAGACAAGATGGGGATGGGGCAGTCAGG
CGTTGGTGCTTTATTTAATGGCATCAATGCATTAAATGCTTATAACGCCGCATTGCTTGCAAAAATTCTC
AAAGTTAGCGTTGAAGAATTTAGCCCTTCAATCGCCAGAGAAATCTACGAGATGTATGAAGCGGTTAGTA
TGCAGCCGTCACTTAGAAGTGAGTATGAGTACCCTGTTTTTTCTCATGTTCAGGCAGGGATGTTCTCACC
TGAGCTTAGAACCTTTACCAAAGGTGATGCGGAGAGATGGGTAAGCACAACCAAAAAAGCCAGTGATTCT
GCATTCTGGCTTGAGGTTGAAGGTAATTCCATGACCGCACCAACAGGCTCCAAGCCAAGCTTTCCTGACG
GAATGTTAATTCTCGTTGACCCTGAGCAGGCTGTTGAGCCAGGTGATTTCTGCATAGCCAGACTTGGGGG
TGATGAGTTTACCTTCAAGAAACTGATCAGGGATAGCGGTCAGGTGTTTTTACAACCACTAAACCCACAG
TACCCAATGATCCCATGCAATGAGAGTTGTTCCGTTGTGGGGAAAGTTATCGCTAGTCAGTGGCCTGAAG
AGACGTTTGGC
CS369 2007
15
Hepatitis C sequence database
• Specialist databases
usually refer to
sequences in the
public databases, but
have extra
information and
search criteria
specific to the
domain.
CS369 2007
16
Hepatitis C sequence database
CS369 2007
17
Problem 1: detecting sequence
similarity between two sequences
• Biologists often want to detect if two
sequences are similar
– How is sequence similarity defined?
– What is it used for?
– Are there different types of similarity?
CS369 2007
18
How is sequence similarity
defined?
• The number of matching nucleotides (when
aligned)?
• The amount of shared information?
• The “distance” between the two sequences
under some metric?
38 out of 60 sites are identical in this alignment
CS369 2007
19
How is sequence similarity
defined?
•
•
•
•
CS369 2007
A1 is 42 nucleotides long
A2 is 60 nucleotides long
So 38/42 = 90% of A1 is “explained” by A2
Whereas 38/60 = 63% of A2 is “explained”
by A1
20
What is similarity used for?
• Detecting homology (shared evolutionary history)
• Reconstructing evolutionary history to better
understand biology
• Determining the structure and function of new
sequences, by matching them with sequences of
known structure/function
• Grouping sequences together to increase
statistical power of single-sequence analyses
• Many many more uses…
CS369 2007
21
Are their different types of
similarity?
• Chance similarity
– For example: if you compare two long random
sequences of DNA you will always find some small
region containing the same sequence.
• Similarity due to a common origin, followed by
divergent/independent evolution (called
homology)
• Similarity due to convergence
– Bird wings and bat wings
– Lysozyme gut enzyme in cows and colobus monkeys
CS369 2007
22
Sequence Homology
x
• Homologous protein or DNA
sequences share common ancestry
– A statement of homology is therefore
an evolutionary hypothesis
• Homology need not imply similar
function
• Homology is a binary property, a pair
of sequences are either homologous
or not homologous.
t
a, b homologous
a
b
x
y
– No such thing as degree of homology
• Homology is often inferred by
sequence similarity
a, b not
homologous
a
CS369 2007
b
23
Origin of similar genes
• Similar genes in the same genome
arise by gene duplication
• Similar genes in different genomes
arise from common ancestry
• A copy of a gene might be inserted
next to the original
• Two copies mutate independently
• Each can take on separate functions
• All or part can be transferred from
one part of genome to another
A
Gene duplication
A
Speciation
A
B
Species I
CS369 2007
B
A’
B’
Species II
24
Orthology and paralogy
"Where the homology is a result of gene duplication so that
both copies have descended side by side during the
history of an organism, (for example, alpha and beta
hemoglobin) the genes should be called paralogous
(para=in parallel). Where the homology is the result of
speciation so that the history of the gene reflects the
history of the species (for example alpha hemoglobin in
man and mouse) the genes should be called orthologous
(ortho=exact). "
Fitch WM. Distinguishing homologous from analogous
proteins. Systematic Zoology 1970 Jun;19(2):99-113.
CS369 2007
25
Orthology and paralogy
CS369 2007
26
Orthology, paralogy and multigene
families
CS369 2007
Reproduced from NCBI education website
27
Solution 1: Pairwise sequence
alignment
• Definition: Procedure for optimizing a score
function on a pair of sequence S1 and S2 by
introducing gap characters into a subsequence of
one or both of the sequences so as to construct
aligned sequences A1 and A2. The objective is to
find the similarity regions in the two sequences.
– A1 and A2 will be the same length.
– Ai will consist only of a subsequence of Si once gap
characters are removed.
CS369 2007
28
Pairwise sequence alignment
Sequences
S1 = a c g g t
S2 = a g g c t t
Alignment
A1 = a c g g – t | || |
A2 = a – g g c t t
CS369 2007
29
Global versus Local Alignment
• We distinguish
– Global alignment algorithms which optimize
overall alignment between two sequences
– Local alignment algorithms which seek only
highly similar subsequences
• Alignment stops at the ends of regions of strong
similarity
• Favors finding conserved patterns in otherwise
dissimilar sequences
CS369 2007
30
Global vs. Local Alignment
• Global
LGPSSKQTGKGS-SRIWDN
|
| |||
| |
LN-ITKSAGKGAIMRLGDA
• Local
--------GKG-------|||
--------GKG--------
CS369 2007
31
Solution 2: The dot plot
G C T A G G A
G
A
C
T
A
G
G
C
CS369 2007
Window size = 1
Matches = 1
0/1
1/1
32
Filtering the dot plot
G C T A G G A
G
A
C
T
A
G
G
C
CS369 2007
Window size = 3
Matches = 2
0/3
1/3
2/3
3/3
33
Dot plots
1,1
2,2
The dot plot is a graphical method that can be tuned
CS369 2007
34
Dot plots
3,3
CS369 2007
5,22
35
Dot matrix analysis with Geneious
• Get phage l cI and phage P22 c2 repressor
sequences from Genbank Nucleotide
database
– Accessions X00166 and V01153 respectively
• Use Geneious 2.5.4
(http://www.geneious.com)
• Use window size of 11 and stringency of 7
• See figure 3.X in Mount
CS369 2007
36
Dot matrix analysis with Geneious
CS369 2007
37
Dot matrix analysis with
Geneious (2)
• Get human LDL receptor protein sequence
from Genbank (accession P01130)
• Make copy, and look at self-similarity
• Use window size of 1 and stringency of 1
• Use window size of 23 and stringency of 7
CS369 2007
38
Human LDL receptor self similarity
1,1
CS369 2007
23,7
39
Dot plots
• Two 100 nucleotide
fragments of the nef
gene
• Low complexity
repetitive region is
visible as dense
region of parallel lines
CS369 2007
40
Which alignment is best?
CS369 2007
41
Problem 2: finding similar sequences in
a database using query sequence
• Biologists often want to find known
sequences that are similar to a newly
obtained sequence
– How to rapidly compare the new sequence to
the hundreds of billions of bases already
sequenced?
– Pairwise align new sequence to all the
sequences in the database?
– Which database to search?
CS369 2007
42
Similarity searching
• Many heuristic algorithms
– BLAST
– FASTA
• Exact algorithms
– Pairwise alignment on all database entries
– Only possible for small databases
CS369 2007
43
BLAST
CS369 2007
44
CS369 2007
45