* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Lecture 8 LC710- 1st + 2nd hr
Non-coding DNA wikipedia , lookup
List of types of proteins wikipedia , lookup
Gene expression wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Cell-penetrating peptide wikipedia , lookup
Messenger RNA wikipedia , lookup
Peptide synthesis wikipedia , lookup
Bottromycin wikipedia , lookup
Non-coding RNA wikipedia , lookup
Protein structure prediction wikipedia , lookup
Oligonucleotide synthesis wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Epitranscriptome wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Molecular evolution wikipedia , lookup
Biochemistry wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Transfer RNA wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
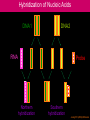
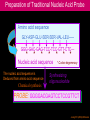
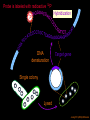
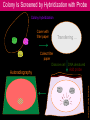
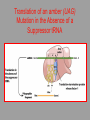
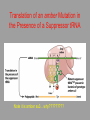
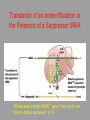
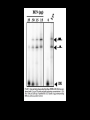
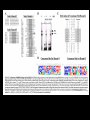
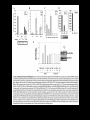
Hybridization of Nucleic Acids DNA1 DNA2 RNA Probe Northern hybridization Southern hybridization Juang RH (2004) BCbasics Preparation of Traditional Nucleic Acid Probe Amino acid sequence GLY-ASP-GLU-SER-SER-VAL-LEU----GGG-GAC-GAG-TCC-TCC-GTT-CTC--- * * * * Nucleic acid sequence The nucleic acid sequence is Deduced from amino acid sequence Chemical synthesis * * * * Codon degeneracy Synthesizing oligonucleotide PROBE: GGGGACGAGTCCTCCGTTCT Juang RH (2004) BCbasics Probe is labeled with radioactive 32P 32 P GG G GAC G AG Hybridization TC GG T C TC CG A T T G T C T C C C C T CAG G G TCC A A G G AGGCAA C C T A DNA denaturation Target gene Single colony Lysed Juang RH (2004) BCbasics Colony Is Screened by Hybridization with Probe Colony hybridization Transferring … Collect filter paper Dissolve cell Autoradiography DNA denatured Add probe Juang RH (2004) BCbasics Cover with filter paper Biochip Based on Hybridization Sample DNA Complementary DNA hybridize Biochip Each spot contains known DNA Signal appears Schena (2000) Microarray Biochip Technology, p. A31 Juang RH (2004) BCbasics The Genetic Code Initiation and termination Codons – Initiation codon: AUG – Termination codons: UAA, UAG, UGA Degeneracy: partial and complete Ordered Nearly Universal (exceptions: mitochondria and some protozoa) Key Points Each of the 20 amino acids in proteins is specified by one or more nucleotide triplets in mRNA. (20 amino acids refers to what is attached to the tRNAs!) Of the 64 possible triplets, given the four bases in mRNA, 61 specify amino acids and 3 signal chain termination. (have no tRNAs!) Key Points The code is nonoverlapping, with each nucleotide part of a single codon, degenerate, with most amino acids specified by two to four codons, and ordered, with similar amino acids specified by related codons. The genetic code is nearly universal; with minor exceptions, the 64 triplets have the same meaning in all organisms. (this is funny) Do all cells/animals make the same Repertoire of tRNAs? The Genetic Code The Wobble Hypothesis: Base-Pairing Involving the Third Base of the Codon is Less Stringent. Base-Pairing with Inosine at the Wobble Position In molecular biology, a wobble base pair is a non-WatsonCrick base pairing between two nucleotides in RNA molecules. The four main wobble base pairs are guanine-uracil, inosine-uracil, inosine-adenine, and inosinecytosine (G-U, I-U, I-A and I-C). The thermodynamic stability of a wobble base pair is comparable to that of a Watson-Crick base pair. Wobble base pairs are fundamental in RNA secondary structure and are critical for the proper translation of the genetic code. Suppressor Mutations Some mutations in tRNA genes alter the anticodons and therefore the codons recognized by the mutant tRNAs. These mutations were initially detected as suppressor mutations that suppressed the effects of other mutations. Example: tRNA mutations that suppress amber mutations (UAG chain-termination mutations) in the coding sequence of genes. Making a (UAG) Mutation Translation of an amber (UAG) Mutation in the Absence of a Suppressor tRNA Translation of an amber Mutation in the Presence of a Suppressor tRNA Note it is amber su3…why????????? Translation of an amber Mutation in the Presence of a Suppressor tRNA If there was a single tRNATyr gene, then could one have a amber supressor of it? Fig1 Oli gonucleotide synthesis is carried out by a stepwise addition of nucleotide residues to the 5'-termi nus of the growing chain until the desired sequence is assembled. Each addition is referred to as a synthetic cycle (Scheme 6) and consists of four chemi cal reactions: * Step 1 - De-blocking (detritylation): The DMT group is removed with a solution of an acid, such as TCA or Dichloroacetic acid (DCA), in an inert solvent (dichloromethane or toluene) and washed out, resulting in a free 5' hydroxyl group on the first base. * Step 2 - Coupli ng: A nucleoside phosphoramidite (or a mi xture of several phosphorami dites) is activated by an acidic azole cataly st, tetrazole, 2-ethylthiotetrazole, 2-bezylthiotetrazole, 4,5-dicyanoimi dazole, or a number of simil ar compounds. This mi xture is brought in contact with the starting soli d support (first coupli ng) or oli gonucleotide precurs or (foll owing coupli ngs) whose 5'-hydroxy group reacts with the activated phosphorami dite moiety of the incomi ng nucleoside phosphoramidite to form a phosphite triester li nkage. This reaction is very rapid and requires, on small scale, about 20 s for its completion. The phosphorami dite coupli ng is also highly sensitive to the presence of water and is commonly carried out in anhydrous acetonitril e. Unbound reagents and by-products are removed by washing. * Step 3 - Capping: After the completion of the coupli ng reaction, a small percentage of the soli d support-bound 5'-OH groups (0.1 to 1%) remains unreacted and needs to be permanently blocked from further chain elongation to prevent the formation of oli gonucleotides with an internal base deletion commonly referred to as (n-1) shortmers. This is done by acetylation of the unreacted 5'-hydroxy groups using a mi xture of acetic anhydride and 1-methylimi dazole as a catalyst. Excess reagents are removed by washing. * Step 4 - Oxidation: The newly formed tricoordinated phosphite triester li nkage is not natural and is of lim ited stabili ty under the conditions of oli gonucleotide synthesis. The treatment of the support-bound material with iodine and water in the presence of a weak base (pyridine, lutidine, or colli dine) oxidizes the phosphite triester into a tetracoordinated phosphate triester, a protected precursor of the naturall y occurr ing phosphate diester internucleosidic li nkage. This step can be substituted with a sulf urization step to obtain oli gonucleotide phosphorothioates (see below). In the latter case, the sulf urization step is carried out prior to capping. New Base Are the proteins produced a pure reflection of the mRNA sequence???? tRNA environment, protein modifications post-translationally Good things to Know RNApol II TATAA CCATGG (Nco I site and Kozak Rule) ATG AGGT….splice GT……………A………polypyrimidine AG PolyA recog sequence AATAAA The Reasons why ATG is a single codon and TGG is a single codon. SELEX yields a functional binding site. A, COS7 cells were transfected in triplicate with either pEBG or increasing concentrations of BENwt (250, 500, and 1000 ng) and either p81TKluc (TK) luciferase reporter or p81TKluc-WT3X (WT3X). The luciferase values are reported as relative luciferase activity normalized to the amount of total protein. -Fold decrease in activity is measured relative to the basal transcriptional activity observed with pEBG empty expression vector alone. Western blot with anti-GST antibody shows dosedependent expression of GST-BEN. B, COS7 cells were transfected in triplicate with either pEBG or BENwt (1000 ng of each) and with either TK, or WT3X or Mut3X (600 ng of each) and the Renilla construct (pRL-TK).