* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Plant Cell Wall Biosynthesis
Survey
Document related concepts
Adenosine triphosphate wikipedia , lookup
Fatty acid metabolism wikipedia , lookup
Signal transduction wikipedia , lookup
Metalloprotein wikipedia , lookup
Lipid signaling wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Proteolysis wikipedia , lookup
Biochemistry wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Biosynthesis of doxorubicin wikipedia , lookup
Transcript
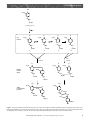
Plant Cell Wall Biosynthesis Secondary article Article Contents Stephen C Fry, The University of Edinburgh, Edinburgh, UK . Introduction Most plant cell wall polymers are synthesized from ‘activated’ precursors by the action of transferases. These enzymes are located in cell membranes (synthesizing polysaccharides and glycoproteins) or in the cell wall itself (synthesizing cutin and suberin). Lignin, in contrast, is synthesized in the wall by oxidative rather than transferase-catalysed reactions. . Mechanisms of Polymer Assembly . Synthesis of the Building Blocks of Cell Wall Polymers . Polymer Assembly in Endoplasmic Reticulum and Golgi Cisternae . Polymer Assembly at the Plasma Membrane . Polymer Assembly in the Cell Wall Introduction Plant cell walls are composed of numerous complicated polymers, principally polysaccharides, glycoproteins, polyesters and lignin. This sophisticated repertoire of polymers is essential for the wall’s ability to govern growth, morphogenesis and disease resistance. The plant invests many enzymes, and thus genes, in the business of manufacturing walls. Also, since walls often constitute the majority of a plant’s dry mass, the plant must invest much carbon and energy in wall production. The building blocks (sugars, amino acids, etc.) for wall biosynthesis are made, and often ‘activated’, in the protoplast; the polymerization process then occurs either in association with membranes or in the wall itself, depending on the polymer under consideration. Mechanisms of Polymer Assembly Transferase-catalysed polymerization Most wall polymers are assembled by transferase-catalysed, nucleophilic substitution (SN2) reactions (Table 1), in which each new bond is synthesized by transfer of a group from an ‘activated’ donor substrate (containing an electrophilic carbon atom) to a nucleophilic group (e.g. – OH or –NH2) in an acceptor substrate. The donor substrate is described as ‘activated’ because its free energy (DG8’) of hydrolysis is sufficient to drive net polymer synthesis. Glycosyl donors The donor substrates for transfer of sugar groups are nucleoside diphosphate sugars (NDP-sugars), the NDP moiety being an excellent leaving group. The most important NDP-sugars are listed in Table 2. All except UDP-Api have the sugar residue in the pyranose (sixmembered) ring form. The acceptor substrate is the nascent polymer. Linear polysaccharides (e.g. cellulose, callose, homogalacturonan) and the backbones of branched polysaccharides (e.g. xyloglucan, galactomannan) appear to be assembled by the repeated addition of single sugar residues to the polymer’s nonreducing terminus. This is described as ‘tailward growth’ and results in the most recently added sugar residue being at the nonreducing end (‘nucleophile end’) of the polymer chain. (In bacterial a-dextrans, polysaccharides synthesized by a different mechanism involving ‘headward growth’, the newest glucose residue is at the reducing end.) Acyl and alkyl donors In protein synthesis, the ‘activated’ donor substrates are peptidyl-tRNAs (Table 1). An aminoacyl-tRNA first acts as the acceptor substrate for the nascent polymer, thus becoming a peptidyl-tRNA molecule. This then acts as a donor substrate, with the next aminoacyl-tRNA as acceptor. The newest amino acid residue is therefore at the C-terminus (furthest from the ‘nucleophile end’) of the polymer – an example of ‘headward growth’. The ester bonds in cutin and suberin and in O-acetyl and O-feruloyl-polysaccharides all appear to be synthesized using coenzyme A (CoA) thioesters as donor substrate, by transesterification reactions. Methyl-ester and methylether groups are added to polysaccharides using Sadenosylmethionine (SAM) as donor. Oxidative polymerization Lignin is synthesized by homolytic reactions, not nucleophilic substitutions. The building blocks (monolignols) are not ‘activated’. Instead, the monolignols are oxidized to form free radicals, which then polymerize (see below). Synthesis of the Building Blocks of Cell Wall Polymers This section discusses the biosynthesis of ‘activated’ forms of these building blocks. ENCYCLOPEDIA OF LIFE SCIENCES / & 2001 Nature Publishing Group / www.els.net 1 Plant Cell Wall Biosynthesis Table 1 Substrates used in the biosynthesis of plant cell wall polymers Bond to be made Bond broken in donor substrate Ester Peptide bond (secondary amide) in protein Acceptor substrate (nucleophile) Example of donor substratea Peptidyl-tRNAAla 70–90 additional nucleotide residues H 2N N N O O n additional aminoacyl residues O CH 2 N O CH 3 C OH O C O UDP-Galactose O C OH O OH O O P O P O CH 2 O O O OH Thioester (a) –OH group on nascent polysaccharide O chain or (b) –OH group of NH Hyp, Ser or Thr in polypeptide or N O (c) –CONH2 in side-chain of Asn CH 2 OH HO Fatty acyl ester bond in cutin or suberin N O N H Glycosidic bond in Glycosylpolysaccharide or pyrophosphate glycoprotein P OH 10,16-Dihydroxyhexadecanoyl–CoA OH CH 2 OH O α-NH2 group of the ‘next’ aminoacyl-tRNA, according to the genetically encoded peptide sequence –OH group in growing polyester molecule C S CH 2 H 2C N H pantothenate CoA phosphate phosphate ribose-3'-phosphate adenine O-Acetyl ester in polysaccharide Thioester Acetyl–CoA CH3 O C –OH group on (nascent?) polysaccharide chain S CoA continued 2 ENCYCLOPEDIA OF LIFE SCIENCES / & 2001 Nature Publishing Group / www.els.net Plant Cell Wall Biosynthesis Table 1 – continued Bond to be made Methyl ester or O-methyl ether in polysaccharide Bond broken in donor substrate Example of donor substratea Trialkylsulfonium S-Adenosyl-methionine Acceptor substrate (nucleophile) COOH HC H 2N NH 2 N CH 2 N CH 2 S H 3C CH 2 OH a N O (a) –OH group on (nascent?) polysaccharide chain (b) –COOH group on (nascent?) pectin chain N OH Pink, group transferred (the electrophilic carbon atom is shown in bold type); black, leaving group; ---, indicates that the bond is broken. Table 2 Important NDP sugars Abbreviation Full name UDP-Glc UDP-Gal UDP-GlcA UDP-GalA UDP-GlcNAc UDP-GalNAc UDP-Xyl UDP-Ara UDP-Rha UDP-Api GDP-Man GDP-Fuc GDP-Gal UDP-a-d-glucose UDP-a-d-galactose UDP-a-d-glucuronic acid UDP-a-d-galacturonic acid UDP-(N-acetyl-a-d-glucosamine) UDP-(N-acetyl-a-d-galactosamine) UDP-a-d-xylose UDP-b-l-arabinose UDP-b-l-rhamnose UDP-a-d-apiofuranose GDP-a-d-mannose GDP-b-l-fucose GDP-b-l-galactose UDP, uridine diphosphate; GDP, guanosine diphosphate. Sugar nucleotides NDP-sugars are primarily derived from a cytosolic pool of five rapidly interconverting hexose monophosphates (Figure 1; enzymes 1–4). Glc-1-P and Man-1-P are drawn off from the pool to form UDP-Glc and GDP-Man, respectively (5, 6). Also, some Glc-6-P may be drawn off to form UDP-GlcA via the inositol pathway (7–11). UDP-Glc and GDP-Man are precursors of all other major NDP sugars (12–20), including UDP-GlcA: enzyme 12 potentially short-circuits the inositol pathway. In addition to the primary pathways of NDP sugar formation (1–20), there are ‘scavenger’ pathways by which certain monosaccharides, released for example during polysaccharide turnover, can be recycled by kinases (10, 21–27) and pyrophosphorylases (5, 6, 11, 28–31). To biochemists, scavenger pathways are valuable because they enable exogenous 3H- or 14C-monosaccharides to be used to radiolabel selected sugar residues in newly synthesized polysaccharides in vivo. Cell wall phenolics Phenolic building blocks (except tyrosine) are formed from cinnamate, itself synthesized from l-phenylalanine by the action of phenylalanine ammonia-lyase (reaction [I]). phenylalanine!cinnamate 1 NH3 [I] Cinnamate is rapidly converted to a family of hydroxycinnamates through reactions [II], where [h] represents the action of hydroxylases, using O2; and [m] represents the action of methyltransferases, using S-adenosylmethionine. cinnamate }[h]! p-coumarate }[h]! caffeate }[m]! ferulate }[h]! 5-hydroxyferulate }[m]! sinapate [II] Hydroxycinnamates do not usually occur as free acids but as esters and amides. Particularly important, although present at low concentrations, are CoA thioesters, formed by (hydroxy)cinnamate:CoA ligase(s) (CCL), as shown in reaction [III] (PPi represents inorganic pyrophosphate). p-coumarate 1 CoA 1 ATP! p-coumaroyl–CoA 1 AMP 1 PPi [III] The CoA thioesters may be converted to the corresponding alcohols (monolignols, e.g. coniferyl alcohol; Figure 2) by (hydroxy)cinnamoyl–CoA reductase (CCR) and (hydroxy)cinnamyl alcohol dehydrogenase (CAD). Hydroxy fatty acids (HFAs) HFAs are assembled on an acyl carrier protein (ACP), using acetyl–CoA and malonyl–CoA as C2 donors, with ENCYCLOPEDIA OF LIFE SCIENCES / & 2001 Nature Publishing Group / www.els.net 3 Plant Cell Wall Biosynthesis the reduced form of nicotinamide–adenine dinucleotide phosphate (NADPH) as reductant. Major intermediates are palmityl-ACP and oleyl-ACP, which are hydrolysed to release palmitic and oleic acids (saturated C16 and 9,10unsaturated C18 chains, respectively). Palmitic acid is hydroxylated (using O2 1 NADPH) to form the C16 HFAs. Oleic acid may be hydroxylated at C18 and/or epoxidated at the C=C group; the epoxide thus formed may be hydrolysed to yield a 9,10-diol. HFA–CoA thioesters are then presumably formed by ligases (see above). Gal ATP 21 23 ADP ADP GalA-1-P Gal-1-P UTP Ara-1-P UTP 28 UTP 29 30 PPi NADP+ + H2O 19 ATP 22 ADP UDP-Rha Ara GalA ATP PPi PPi UDP-D-Gal UDP-GalA UDP-Ara * 14 * 15 * 16 CO2 Glc-1-P 5 12 UDP-Glc UDP-GlcA 2NAD+ 2NADH +2H+ UTP PPi 25 7 Glc-6-P Ins-1-P * CO2 20 GlcA-1-P Pi ADP 26 Fru-6-P UDP-Api ADP 8 * 2 10 ATP O2 9 Inositol GlcA * 3 ATP Man ADP 27 Man-6-P GDP-L-Gal Fuc ATP * 4 * 18 24 ADP PPi GTP Man-1-P 6 GTP PPi 4 Wall proteins are synthesized by ribosomes on the outer surface of the rough endoplasmic reticulum (ER). At the N-terminus of the nascent polypeptide is a hydrophobic ‘signal sequence’ that leads this end of the chain through the membrane into the ER lumen. The signal sequence is later removed enzymatically. The new protein is carried through the endomembrane system: ER!cis-Golgi! medial-Golgi!trans-Golgi!plasma membrane. During this journey, the protein may be posttranslationally modified. In extensin, for example, most of the proline residues get oxidized to 4-hydroxyproline (Hyp) about 2.5 min after their incorporation into the polypeptide chain. This was shown by pulse-labelling carrot roots with [14C]proline (Sadava and Chrispeels, 1971). Later, most of the Hyp residues are O-arabinosylated as in reaction [IV] (where n 5 1–4) and some Ser residues are O-galactosylated. [IV] Ara n Many wall enzymes are N-glycosylated: oligosaccharides (rich in Man and GlcNAc; sometimes also Fuc, Gal, Xyl, etc.) are attached to the –CONH2 groups of specific Asn residues. In some enzymes, e.g. xyloglucan endotransglycosylase (XET), this N-glycosylation is essential for full enzymic activity. ADP H2O Fru UDP-Xyl PPi UTP ATP 13 11 * 1 ATP Proteins and glycoproteins … −(Hyp)− … + nUDP-Ara → … −(Hyp)− … + nUDP ?NADPH + H+ Glc Polymer Assembly in Endoplasmic Reticulum and Golgi Cisternae GDP-Man 17a–c GDP-Fuc NADPH NADP+ + H+ + H2O 31 Fuc-1-P Figure 1 Major pathways for the synthesis and interconversion of the NDP sugars used in the biosynthesis of plant cell wall polysaccharides. Some indication of the relative flux through the various pathways is given by arrow thickness. In addition, solid arrows indicate primary pathways for de novo synthesis of NDP sugars and inositol. Pecked arrows (————) imply great variation between tissues. Dot-dashed arrows (–––) indicate scavenger pathways involved in recycling monosaccharides, e.g. released by polysaccharide turnover. The ‘box’ is the pool of hexose monophosphates mentioned in the text. Reactions marked * are isomerizations, with no other reactants. Numbered enzymes are: 1, phosphoglucomutase; 2, glucose 6-phosphate isomerase; 3, mannose 6phosphate isomerase; 4, phosphomannomutase; 5, UDP-glucose pyrophosphorylase; 6, GDP-mannose pyrophosphorylase; 7, myo-inositol 1-phosphate synthase; 8, myo-inositol 1-phosphatase; 9, myo-inositol oxygenase; 10, glucuronokinase; 11, UDP-glucuronate pyrophosphorylase; 12, UDP-glucose dehydrogenase; 13, UDPglucuronate decarboxylase; 14, UDP-glucose 4-epimerase; 15, UDPglucuronate 4-epimerase; 16, UDP-xylose 4-epimerase; 17, ‘GDP-fucose synthase’ (three individual activities: (a) GDP-D-mannose 4,6-dehydratase, (b) GDP-4-keto-6-deoxy-D-mannose 3,5-epimerase, and (c) GDP-4-keto-Lfucose 4-reductase); 18, GDP-mannose 3,5-epimerase; 19, ‘UDPrhamnose synthase’ (probably three activities, cf. 17); 20, UDP-apiose synthase; 21, D-galactokinase; 22, galacturonokinase; 23, arabinokinase; 24, fucokinase; 25, hexokinase or glucokinase; 26, fructokinase; 27, mannokinase; 28, UDP-D-galactose pyrophosphorylase; 29, UDPgalacturonate pyrophosphorylase; 30, UDP-arabinose pyrophosphorylase; 31, GDP-fucose pyrophosphorylase. ENCYCLOPEDIA OF LIFE SCIENCES / & 2001 Nature Publishing Group / www.els.net Plant Cell Wall Biosynthesis OH CH 3O HC CH CH 2OH Coniferyl alcohol [H] O• Free radicals: O CH 3O CH 3O O C • O CH 3O CH 3O C• HC HC CH CH 2OH HC CH CH 2OH A HC CH CH 2OH B CH 2OH C D A+B coupling D+D coupling O O O CH 3O • CH CH 3O CH 3O O CH 3O Initially formed dimers HC HC CH CH 2OH HC HC HC CH 2OH CH CH CH 2OH CH 2OH OH OH CH 3O CH 3O O Stable rearrangement products CH 3O O HC HC CH CH 2OH HC HC CH H 2C CH 2 CH CH O CH 2OH OCH 3 OH Figure 2 Early steps in lignin synthesis. Coniferyl alcohol (one of three monolignols) is oxidized enzymically, losing one hydrogen atom to form a free radical, which rapidly interconverts between four tautomers (A, B, C, D). These pair off nonenzymically to form dimers (two of the several possible dimers are illustrated), some of which (e.g. the D 1 D dimer) undergo intramolecular substitution reactions to form more stable products. ENCYCLOPEDIA OF LIFE SCIENCES / & 2001 Nature Publishing Group / www.els.net 5 Plant Cell Wall Biosynthesis Matrix polysaccharides Pectins and hemicelluloses (except callose) are synthesized in Golgi bodies. Polysaccharide assembly begins in ciscisternae, the finishing touches being applied in the medialand/or trans-cisternae. The basic reaction [V] is catalysed by transferases called NDP sugar : polysaccharide transglycosylases or polysaccharide synthases. NDP-sugar 1 (sugar)n!NDP 1 (sugar)n+1 [V] Specific permeases carry NDP-sugars into the Golgi lumen, where polysaccharide synthesis occurs. All NDP-d-sugars are a-anomers, whereas the d-sugar residues in polysaccharides may be a- and/or b-anomers. Simplistically, transglycosylation (as an SN2 reaction) should reverse the anomerism: thus UDP-a-Xyl could directly add a b-Xyl residue to a xylan chain. Retention of anomerism, on the other hand, e.g. when UDP-a-Xyl adds an a-Xyl residue to xyloglucan, suggests two consecutive transglycosylations (a!b!a). In such cases, the enzyme itself may accept the sugar residue before adding it to the nascent polysaccharide, reversal of anomerism occurring at each step (reactions [VI] and [VII]). UDP-a-Xyl 1 [enzyme]!b-Xyl–[enzyme] 1 UDP [VI] b-Xyl–[enzyme] 1 (sugar)n!a-Xyl–(sugar)n 1 [enzyme] The activity of the transglycosylases (rather than supply of NDP-sugars) is the main variable regulating polysaccharide biosynthesis. For example, pectin and xylan synthases decrease and increase, respectively, during the transition to secondary wall production during xylem differentiation (Bolwell and Northcote, 1981). Nonsugar groups (methyl, acetyl and feruloyl esters; methyl ethers) may also be added to polysaccharide chains in Golgi cisternae, as shown in reactions [VIII] and [IX] (Me, methyl ester or ether; Ac, acetyl ester). The feruloyl donor may be feruloyl–CoA. S-adenosyl-methionine 1 (sugar)n!(sugar)n –Me 1 S-adenosyl-homocysteine Ac-CoA 1 (sugar)n!(sugar)n –Ac 1 CoA [VIII] [IX] The completed polysaccharides in the trans-cisternae are finally carried in vesicles to the plasma membrane, and thence into the apoplast, to be integrated into the wall. [VII] During xyloglucan synthesis, the a-Xyl side-chains are added only to those b-Glc residues that have themselves very recently been incorporated into the nascent polymer’s backbone; equally, further elongation of the backbone appears to depend on successful a-xylosylation at or near the nonreducing end. The enzymes cannot complete a glucan backbone and then add a-Xyl side-chains. In contrast, a-Fuc side-chains can be added to a preformed xyloglucan core. Similar conclusions have been reached through studies of galactomannan biosynthesis (Reid et al., 1995). Specific polysaccharides synthesized in the Golgi bodies include homogalacturonan, xyloglucan, (glucurono)(arabino)xylans and (galacto)(gluco)mannans, as demonstrated by two main approaches: 1. Isolated Golgi bodies can synthesize these polysaccharides from the appropriate NDP sugars (Hobbs et al., 1991; Reid et al., 1995). The transferase activities responsible are integral to Golgi membranes; few such enzymes have been solubilized in active form. However, partial fractionation of the cis-, medial- and trans-cisternae has indicated that different steps in polysaccharide assembly occur in different cisternae. 6 2. Immunocytochemical studies by electron microscopy have demonstrated the presence of epitopes characteristic of these polysaccharides within the Golgi cisternae and associated vesicles. Simultaneous labelling with two distinguishable antibodies confirms that different epitopes first appear in different cisternae (Zhang and Staehelin, 1992). Polymer Assembly at the Plasma Membrane Cellulose and callose are not synthesized within the endomembrane system but at the plasma membrane. Callose Callose synthase is readily demonstrated in isolated plasma membranes in vitro (reaction [X]) and can be solubilized in an active form. UDP-Glc 1 (Glc)n!(Glc)n+1 1 UDP [X] Callose synthase activity is immediately promoted by various stimuli, e.g. Ca2 1 , perhaps explaining the dramatic induction of callose as a wound response. Cellulose The mechanism of plant cellulose synthesis is poorly understood. This is particularly galling because cellulose is the world’s most abundant organic chemical! Quantitative metabolic studies in vivo showed that UDP-Glc is a precursor of cellulose. However, cellulose synthesis has ENCYCLOPEDIA OF LIFE SCIENCES / & 2001 Nature Publishing Group / www.els.net Plant Cell Wall Biosynthesis never been demonstrated with plant enzymes in vitro at rates approaching those observed in vivo. In vivo, new cellulose chains are almost instantly corralled into microfibrils, which appear by electron microscopy as strands issuing from ‘terminal complexes’ (rosettes) in the plasma membrane. Each terminal complex presumably contains several cellulose synthase molecules (equal to the number of cellulose chains in a microfibril cross-section). Since microfibrils are firmly embedded in the wall, the terminal complexes must move about in the plasma membrane as the microfibrils elongate. This movement, and hence the orientation of new microfibrils, may be guided by microtubules in the cytosol. Since plants have unique cell walls, inhibitors of wall biosynthesis could be highly specific herbicides. Cellulose synthesis in vivo is specifically inihibited by the herbicides dichlobenil (2,6-dichlorobenzonitrile) and isoxaben. The interaction of these compounds with cellulose synthase may assist identification of the enzyme or a closely associated protein. Recent research has identified genes that may encode plant cellulose synthases. In cotton seed hairs, a gene called celA1 is massively upregulated when synthesis of the cellulose-rich secondary wall begins, 17 days after anthesis. The celA1 gene resembles a bacterial cellulose synthase gene (celA) and celA1 protein binds UDP-Glc. These observations suggest that celA1 encodes a cotton cellulose synthase (Pear et al., 1996). An Arabidopsis mutant (rsw1) exhibiting ‘radial swelling’ may be defective in cellulose synthase. The mutant fails to synthesize crystalline cellulose and the terminal complexes disorganize, suggesting that the wild-type allele encodes a protein involved in microfibril assembly (Arioli et al., 1998). Thus, although there are still severe difficulties in studying plant cellulose synthases, rapid progress is being made in the related genetics. Polymer Assembly in the Cell Wall Polysaccharide and glycoprotein crosslinking The final integration of polysaccharides and proteins, synthesized by the protoplast, occurs in the wall itself. For example: . XET may attach new segments of xyloglucans to the ends of existing wall-bound xyloglucans (Thompson et al., 1997); . enzymic removal of methyl-esters allows pectins to become crosslinked by Ca2 1 bridges; . oxidative crosslinking of ferulate and tyrosine sidechains of some polysaccharides and proteins, respectively, may occur in the wall by peroxidase action (cf. lignin synthesis). Cutin and suberin Cutin and suberin are water-insoluble; they are synthesized at their final destination in the wall and/or cuticle. HFA– CoAs (see above) are probably the major precursors, as indicated by the ability of a particulate fraction from bean leaf epidermis to incorporate [14C]HFAs into insoluble [14C]polyesters in the presence of CoA and ATP (Croteau and Kolattukudy, 1974). It is unclear how HFA–CoAs reach the cutinizing or suberizing part of the wall or how CoA, released during polymerization, is recycled. Lignin Lignin is synthesized in the wall by polymerization of secreted monolignols. In contrast to other major biopolymers, lignin forms from free radical intermediates. Each monolignol (e.g. coniferyl alcohol, C10H12O3) loses 1H, thus becoming a free radical, (C10H11O3).. The major hydrogen acceptor is probably H2O2, as in reaction [XI] (catalyst: peroxidase). 2 C10H12O3 1 H2O2!2 (C10H11O3). 1 2 H2O [XI] However O2 may also contribute, as in reaction [XII] (catalyst:oxidase). 4 C10H12O3 1 O2!4 (C10H11O3). 1 2 H2O [XII] The free radicals formed undergo rapid, nonenzymic interconversions (tautomerization; Figure 2), before eventually coupling, as in reaction [XIII], to form a dimer (C20H22O6). 2 (C10H11O3).!(C10H11O3)–(C10H11O3) [XIII] The new bond formed can be C–C or C–O, depending on which tautomers participate (Figure 2). Coupling is widely assumed to be stochastic (semirandom); however, recent evidence suggests that ‘dirigent’ proteins can favour specific couplings, at least in related intraprotoplasmic reactions (Davin et al., 1997). Some of the dimers (quinone methides) undergo nonenzymic intramolecular rearrangements or bonding to external nucleophiles (e.g. polysaccharides, generating ‘lignin–carbohydrate complexes’). A dimer can itself lose an H to become a new free radical, (C20H21O6)., which can couple with another free radical. If the latter is also a dimer, as in reaction [XIV], then the product is a tetramer (C40H42O12). 2 (C20H21O6).!(C20H21O6)–(C20H21O6) [XIV] Continuation of such reactions ( 5 oxidative polymerization) generates the lignin polymer. The source of H2O2 for lignin synthesis remains uncertain. It may arise from O2 by the action of wall-bound oxidases on polyamines (reaction [XV]; R– is ENCYCLOPEDIA OF LIFE SCIENCES / & 2001 Nature Publishing Group / www.els.net 7 Plant Cell Wall Biosynthesis NH2 –(CH2)4 –NH–(CH2)2 – in the case of spermidine) or some other reducing agent. R–CH2 –NH2 1 O2 1 H2O!R–CHO 1 H2O2 1 NH3 [XV] References Arioli T, Peng L, Betzner AS et al. (1998) Molecular analysis of cellulose biosynthesis in Arabidopsis. Science 279: 717–720. Bolwell GP and Northcote DH (1981) Control of hemicellulose and pectin synthesis during differentiation of vascular tissue in bean (Phaseolus vulgaris) callus and in bean hypocotyl. Planta 152: 225– 233. Croteau R and Kolattukudy PE (1974) Biosynthesis of hydroxy fatty acid polymers. Enzymatic synthesis of cutin from monomer acids by cell-free preparations from the epidermis of Vicia faba leaves. Biochemistry 13: 3193–3202. Davin LB, Wang H-B, Crowell AL et al. (1997) Stereoselective bimolecular phenoxy radical coupling by an auxiliary (dirigent) protein without an active center. Science 275: 362–366. Hobbs MC, Delarge MHP, Baydoun EA-H and Brett CT (1991) Differential distribution of a glucuronyltransferase, involved in glucuronoxylan synthesis, within the Golgi apparatus of pea (Pisum sativum var. Alaska). Biochemical Journal 277: 653–658. Pear JR, Kawagoe Y, Schreckengost WE, Delmer DP and Stalker DM (1996) Higher plants contain homologs of the bacterial celA genes encoding the catalytic subunit of cellulose synthase. Proceedings of the National Academy of Sciences of the USA 93: 12637–12642. Reid JSG, Edwards M, Gidley MJ and Clark AH (1995) Enzyme specificity in galactomannan biosynthesis. Planta 195: 489–495. 8 Sadava D and Chrispeels MJ (1971) Intracellular site of proline hydroxylation in plant cells. Biochemistry 10: 4290–4294. Thompson JE, Smith RC and Fry SC (1997) Xyloglucan undergoes inter-polymeric transglycosylation during binding to the plant cell wall in vivo: evidence from 13C/3H dual labelling and isopycnic centrifugation in caesium trifluoroacetate. Biochemical Journal 327: 699–708. Zhang GF and Staehelin LA (1992) Functional compartmentation of the Golgi-apparatus of plant-cells – immunocytochemical analysis of high-pressure frozen-substituted and freeze-substituted sycamore maple suspension-culture cells. Plant Physiology 99: 1070–1083. Further Reading Bolwell GP (1988) Synthesis of cell wall components: aspects of control. Phytochemistry 27: 1235–1253. Boudet AM, Lapierre C and Grima-Pettanati J (1995) Biochemistry and molecular biology of lignification. New Phytologist 129: 203–236. Delmer DP and Amor Y (1995) Cellulose biosynthesis. Plant Cell 7: 987– 1000. Delmer DP and Stone BA (1988) Biosynthesis of plant cell walls. In: Preiss J (ed.) Biochemistry of Plants: a Comprehensive Treatise, vol. 14, pp. 373–419. New York: Academic Press. Feingold DS and Barber GA (1990) Nucleotide sugars. In: Dey PM (ed.) Methods in Plant Biochemistry, vol. 2, pp. 39–78. London: Academic Press. Fry SC (1988) The Growing Plant Cell Wall: Chemical and Metabolic Analysis. Harlow, Essex: Longman. Kolattukudy PE (1996) Biosynthetic pathways of cutin and waxes, and their sensitivity to environmental stresses. In: Kerstiens G (ed.) Plant Cuticles – An Integrated and Functional Approach, pp. 83–108. Oxford: Bios. ENCYCLOPEDIA OF LIFE SCIENCES / & 2001 Nature Publishing Group / www.els.net