* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download bio98a_l10
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Metabolic network modelling wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Deoxyribozyme wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Biochemical cascade wikipedia , lookup
Nicotinamide adenine dinucleotide wikipedia , lookup
Epitranscriptome wikipedia , lookup
Magnesium in biology wikipedia , lookup
Metalloprotein wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
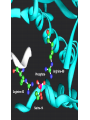
Catalytic triad wikipedia , lookup
Restriction enzyme wikipedia , lookup
Mitogen-activated protein kinase wikipedia , lookup
Lipid signaling wikipedia , lookup
Citric acid cycle wikipedia , lookup
Proteolysis wikipedia , lookup
Biochemistry wikipedia , lookup
Ultrasensitivity wikipedia , lookup
Adenosine triphosphate wikipedia , lookup
Biosynthesis wikipedia , lookup
Phosphorylation wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
Enzyme inhibitor wikipedia , lookup
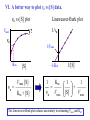
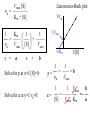
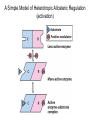
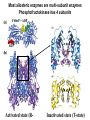
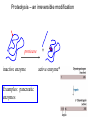
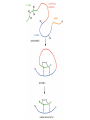
Bio 98 - Lecture 10 Enzymes III: Competitive Inhibition & Enzyme Regulation E+S k1 k-1 ES k2 E+P Note: [E], concentration of free enzyme, is not the same as [E]t VI. A better way to plot vo vs [S] data. vo vs [S] plot Vmax Lineweaver-Burk plot ? vo 1/vo 1/Vmax Km [S] Vmax [S] vo = ––––––––– Km + [S] -1/Km 1/[S] 1 Km 1 1 –– = –––– ––– + ––––– vo Vmax [S] Vmax The Lineweaver-Burk plot reduces uncertainty in estimating Vmax and Km. Vmax [S] vo = ––––––––– Km + [S] Take reciprocal of both sides of equation Km + [S] 1 = –––––––– vo Vmax [S] Expand [S] K 1 m + = –––––––– vo Vmax [S] Vmax[S] Thus 1 Km 1 1 –– = –––– ––– + ––––– vo Vmax [S] Vmax Lineweaver-Burk y = ax + b Lineweaver-Burk plot Vmax [S] vo = ––––––––– Km + [S] 1/vo 1 Km 1 1 –– = –––– ––– + ––––– vo Vmax [S] Vmax y = a x + 1/Vmax -1/Km 1/[S] b Solve for y at x=1/[S]=0: 1 1 y = –– = –––– = b vo Vmax Solve for x at y=1/v0=0: 1 1 Vmax b x = –– = - –––– –– = - –– [S] Vmax Km a How do you measure competitive inhibition? K1 [I] = 2KI K-1 [I] = KI [I] = 0 K-I -1 Km Vmax [S] vo = ––––––––– aKm + [S] where -1 aKm [I] a = 1 + ––– KI [E][I] KI = –––––– [EI] Vmax remains unchanged, but apparent Km increases with increasing [I] Modes of Enzyme Regulation 1. Allosteric* control/regulation • homotropic allostery (O2 for hemoglobin) • heterotropic allostery (H+, CO2, BPG for Hb) 2. Covalent modification • group addition - often reversible, ie phosphorylation allosteric* = allo (other); steric (shape, object) Regulation of Enzyme Activity It would be wasteful to continue to turn substrate into product if enough is available for proper cellular function. Therefore, enzymes often are highly regulated by binding small molecule regulators that can either decrease or increase activity. A classic example is in amino acid metabolism. Several enzymes are required to convert simple substrates into more complex amino acids: Thr E1 threonine dehydratase B E2 E3 E4 E5 Ile In this example, five enzyme-catalyzed steps are required to convert Thr to Ile. When there is sufficient Ile available, Ile will “feedback” inhibit enzyme 1 (E1) that converts Thr to intermediate B. Such inhibition effectively shuts down the entire pathway. There must be a careful balance between the concentration of Ile required for normal function and the concentration of Ile required to inhibit enzyme E1. FEEDBACK INHIBITION! A Simple Model of Heterotropic Allosteric Regulation (activation) Heteroallosteric Regulation 2 subunit enzyme model T-state R-state inhibitor activator substrate Heteroallostery: Effect of inhibitor or activator on vo vs [S] plot Vmax R-state vo Vmax T-state 2 Kmapp Km Kmapp [S] Phosphofructokinase-1 is regulated by both activators and inhibitors activates (ADP) phosphoenolpyruvate PEP pyruvate + ATP F 6-bisF deactivates (phosphoenolpyruvate) + ADP + PPP F 6-P (mM) When the concentration of phosphenolpyruvate (PPP) reaches a certain level, then P6-F kinase activity is lowered by feedback inhibition from phosphenolpyruvate. On the other hand, ADP is the allosteric activator; this may seem strange but the net objective of glycolysis is generation of ATP. If the ATP levels drop (ie [ADP] goes up), glycolysis needs to be stimulated. Thus ADP binds to and activates phosphofructokinase-1 to increase ATP production. Most allosteric enzymes are multi-subunit enzymes Phosphofructokinase has 4 subunits F6bisP + ADP ADP Activated state (R- Inactivated state (T-state) Covalent modifications Phosphorylation - a reversible modification ATP ADP protein kinase activity state 1 Enz Enz protein phosphatase = HOPO3 H20 = -OPO3 activity state 2 Phosphorylation is reversible and is used in many pathways to control activity. Enzymes that add a phosphate to a hydroxyl side chain are commonly called kinases. Enzymes that remove a phosphate from a phosphorylated side chain are called phosphatases. Proteolysis – an irreversible modification protease inactive enzyme Examples: pancreatic enzymes + active enzyme*