* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Document
Gene therapy of the human retina wikipedia , lookup
Genomic imprinting wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Neocentromere wikipedia , lookup
Gene therapy wikipedia , lookup
Polymorphism (biology) wikipedia , lookup
Pharmacogenomics wikipedia , lookup
Skewed X-inactivation wikipedia , lookup
Genetic engineering wikipedia , lookup
Gene expression programming wikipedia , lookup
Nutriepigenomics wikipedia , lookup
X-inactivation wikipedia , lookup
Fetal origins hypothesis wikipedia , lookup
Tay–Sachs disease wikipedia , lookup
Human genetic variation wikipedia , lookup
Hardy–Weinberg principle wikipedia , lookup
Medical genetics wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Designer baby wikipedia , lookup
Neuronal ceroid lipofuscinosis wikipedia , lookup
Genome (book) wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Genome-wide association study wikipedia , lookup
Genetic drift wikipedia , lookup
Population genetics wikipedia , lookup
Public health genomics wikipedia , lookup
Dominance (genetics) wikipedia , lookup
HLA A1-B8-DR3-DQ2 wikipedia , lookup
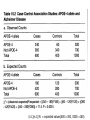
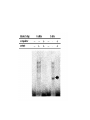
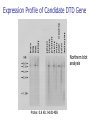
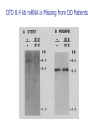
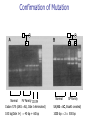
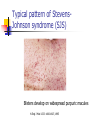
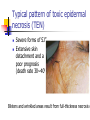
Allelic Association and Transmission Disequilibrium Test 成大醫學院 病理科分子病理實驗室 分子診斷實驗室 呂政展 (x 2641) Linkage Disequilibrium (allelic association) • Def: nonrandom association of alleles at linked loci • f(A, B)= f(A) x f(B) •: Haplotype • Haplotype: a series of alleles found at linked loci on a single chromosome. e.g, – A33-Cw10-B58-DRB1*0301-DQB1*02(6.3%); A2-B46-Cw11DRB1*09-DQB1*0303 (orientals); – A30-Cw5-B18(Bw6)-DRB1*0301-DQB1*02 (most frequent in Mediterranean) – A1-Cw7-B8(Bw6)-DRB1*0301- DQB1*02 (commonest in European Caucasoids) The Relationship of Disease Locus and Marker Defined by Linkage Disequilibrium The stronger the linkage disequilibrium, the closer the marker is to the disease locus Mapping and/ or gene identification using LD is particularly useful in genetically unique or isolated populations (Amish and Finnish populations) LD rarely extends more than 1 cM from the susceptibility locus, its detection indicates a significant narrow down of candidate region Allelic Association Studies How it happens? Direct biological action of the genetic polymorphism Linkage disequilibrium with a adjacent susceptibility gene How useful in the analysis of genetic complex disease? Evaluation of candidate gene loci Fine mapping of region that is indicated by linkage analysis for follow-up studies Association vs Linkage Association and linkage: distinct from each other Linkage: a specific genetic relationship between loci Association: Statistical statement about the cooccurrence of alleles or phenotypes “Allele A is associated with disease D” : if individuals who have D also have A more often than would be predicted from the individual frequencies of D and A in the population Allelic Association Def: A significantly increased or decreased frequency of a marker allele with a disease trait and represents deviations from the random occurrence of the alleles regarding the disease phenotypes Linkage disequilibrium means allelic association caused or maintained by tight linkage Linkage Disequilibrium in Genetic Analysis Defining the ancestral haplotype of a disease gene in relation to several marker loci Fine-mapping of the disease gene even when complete linkage ( = 0.0) is established in the family studies The slowness of linkage disequilibrium decay make LD a useful mapping tool Conditions for LD Mapping Founder population Geographically isolated population, traceable ancestry, most individuals of current population are descendants of a few individuals back 250 years ( 20 generations) The prevalence of a genetic disease is derived from an allele for a disease gene from a common ancestor Both parents in many of the marriages within a founder populations are heterozygous for a recessive disease gene, will have a 25% chance of having an affected offspring Rate of Decay of Linkage Disequilibrium: time and distance-dependent Dt = Do(1-)t t : current generation number Dt : current amount of disequilibrium Do : disequilibrium at generation 0 : recombination fraction between loci Decay in linkage disequilibrium Significance of Allelic Association Allelic associations reflect sharing of ancestral chromosomes, only alleles at loci tightly linked to the disease susceptibility locus will still be shared For a locus showing recombination fraction (θ) with the susceptibility locus, a proportion (θ ) of ancestral chromosome will lose the association each generation, and a proportion (1- θ) will retain it. (1-0.01)44 = 0.64, loci 1cM apart (1-0.03)44 = 0.26, loci 3 cM apart Linkage Disequilibrium as a Mapping Tool-I Cystic fibrosis mapped to 7q32, the candidate region was still very large XV2.c and KM19 marker within the candidate gene show strong association between (XV2.c*1, KM19*2) haplotype and CF Linkage Disequilibrium as a Mapping Tool-II Nijmegen breakage syndrome (NBS) Rare autosomal recessive disease characterized by chromosomal breakage, growth retardation, microcephaly, immunodeficiency and a predisposition to cancer Genetic linkage in families mapped NBS to an 8Mb regions between D8S271 and D8S270 Linkage Disequilibrium as a Mapping Tool-II Nijmegen breakage syndrome (NBS) 74 haplotypes related to a common ancestral haplotype. Some do not have this haplotype, possibly carry independent NBS mutations. Others share only part of haplotype, showing the effect of recombination in distant ancestors Ancestral haplotype in patients with Nijmegen breakage syndrome A: allele attributed to AH 51 unrelated patients typed for microsatellites Linkage Disequilibrium Mapping I1 G1 E1 C1 A1 N B1 D1 F1 H1 J1 K1 Mutation N→M I1 G1 E1 C1 A1 10 M B1 generations D1 F1 H1 J1 K1 I2 G3 E1 C1 A1 90 M B1 generations D1 F1 H1 J3 K4 I3 G2 E2 C3 A1 M B1 D1 F2 H3 J4 K2 Jennings 1917 first developed the concept of LD Richard Lewontin (1964) first used LD to measure D Linkage Disequilibrium Coefficient (D)(Lewontin 1964) D = P11- p1q1 (if D significantly differs from zero, LD is thought to exist) locus A(1,2) locus B(1,2) P11 :observed frequency of the 1/1 haplotype p1 : frequency of the allele “1” at the locus A q1 : frequency of the allele “1” at the locus B p1 = f(A1), p2 = 1- p1= f(A2) q1 = f(B1), q2 = 1-q1 = f(B2) Dmax = min(p1q2, p2q1) Dmin = max(-p1q1, -p2q2) D’ =D/Dmax Alternatively, D = h11- p1q1 , where h11 is the frequency of the haplotype with the rarer allele at each locus, and where p and q are frequencies of the rarer alleles at loci 1 and 2, respectively (Devlin and Risch, 1995). D’ is positive when the rarer alleles at each locus are associated and is negative when a common allele is associated with a rare allele Dmax = min[(pq, (1-p)(1-q)], for D <0 Dmax = min[(p(1-q), q(1-p)], for D >0 Determination of LD Coefficient D = d n bd cd x n n •a, b, c, d are the phenotype frequencies of the +/+, +/-, -/+ and -/- combinations of the alleles in each haplotype and n is the sum of the a, b, c, d. •HFij = Dij + GFi x GFj •GF = 1 - 1 AF Measure LD by D(2) (Hill and Robertson 1968) R or =√2/N = D/√(p1p2q1q2) 2 statistic= obtained from 2 x 2 table N: total number of haplotypes in the sample p11 P12 p21 p22 Genome research 10: 1435-1444 Measure of LD-(3) (Bengtsson and Thomson 1981) = D/(q1p22) q1: The population frequency of a disease allele, B1 P22: the frequency of chromosome that contain marker allele A2 and normal allele, B2 and D’ : give more reliable estimates of physical distances than do D and R, because D and R depends on allele frequency Quantitative Measurement of Linkage Disequilibrium Yule coefficient = (p1,1- p1,2)/(p1,1 + p1,2 - 2 p1,1 p1,2)(used in Huntington’s disease) A1, A2 locus A B1, B2 locus B p1,1 = the frequency of allele A1 on the chromosome p1,2 carrying allele B1 = the frequency of allele A1 on the chromosome carrying allele B2 Allelic association around the locus for Huntington Disease Acc I Total Distance= 2500 kb 250 kb Mbo I Taq I L19ps11 BS674 (n1.)(n.1)/N = expected values(600 x 300 /1000 =180) Advantages of Association Studies Association studies are easier to conduct than linkage analysis , no multicase families or special family structures LD is a short range phenomenon, if association exists, it defines a small candidate region in which to search for disease gene Association is more powerful than linkage for detecting weak susceptibility alleles Possible Causes of Positive Association-I Direct causation: having allele A make you susceptible to disease D; possession of A is neither necessary nor sufficient to develop disease, but it increases the likelihood Natural selection: People who have disease D might be more likely to survive and have children if they also have allele A Population stratification: population contains several genetically distinct subsets. Both the disease D and allele A happen to be particularly frequent in one subset (HLAA1 and chopsticks user in San Francisco bay area). Possible Causes of Positive Association-II Statistical artefact: association studies often tests a range of loci, each with several alleles, for association with a disease. The raw p values need correction for the numbers of genes tested Linkage disequilibrium: If LD exist between the disease D and the locus A, there should be a gene near to the A locus that has mutations in people with disease D. Advantages and Pitfalls of Association Study Advantages Not restricted to nonmendelian genetics Easy to perform (case and control) Pitfalls Selection of controls is very crucial ( representative?!) Probabilities calculated from Association studies must be corrected for the number of questions asked Threshold of significance is p = 0.05/n (Bonferroni correction) Relative Risk (RR)/Odds Ratio(OR) Estimate how many times more the carrier of a specific allele or haplotype was likely to have a specific disease associated with the particular genetic marker studied Calculation of RR RR= a x d/ b x c a: the number of patients positive for the allele or haplotype b: the number of patients negative for the allele or haplotype c: the number of controls positive for the allele or haplotype d: the number of controls negative for the allele or haplotype Calculation of RR (Relative Risk) Risk factor Present Absent Total Cases a c a+c Controls b d b+d Sample Total a+b c+d n Linkage disequilibrium mapping is carried out after genetic linkage between a polymorphic locus and the disease gene is determined The Genetic Distance() between a Marker Locus and the Disease Gene that are in Linkage Disequilibrium-I Pexcess= Paffected-Pnormal/(1- Pnormal)= (1- gq-1 )(1- )g = recombination fraction between marker and disease loci = mutation rate (1 x 10-6) g= the number of generations after since the common ancestor q= world wide frequency of the disease ( =0.001) The Genetic Distance() between a Marker Locus and the Disease Gene that are in LD-II Pnormal= the proportion of the marker allele in normal chromosomes Paffected = the proportion of the marker allele in diseased chromosomes Pexcess = measure of disequilibrium, the fraction of the excess occurrence of a chromosome with the disease gene and a marker allele compared with the chromosome with the nondisease gene and the marker allele Functional SNPs in the lymphotoxin-α gene that are associated with susceptibility to myocardial infarction (Nature Genetics 32: 650-654, 2002) Linkage disequilibrium mapping Haplotype analysis D’= D/Dmax 65,671/92,788 = 70.8% A cut-off P value of 0.01 for association in either recessive and dominant mode A SNP in intron 1 of LTA (252A->G) on chromosome 6p21was associated with myocardial infraction in the initial screening Construct a high-density SNP map for LD mapping by sequencing 16 MI patients and 16 normal controls. 187 SNPs, 130 kb within 6p21 Select 120 SNPs (>10%) and genotyping 94 MI patients and 94 subjects from general population 26 SNPs with minor allele frequency > 0.25 +ive 1 2 3 4 Located in one extended block of intense LD, with D’ dropping off near p5-1 and AIF1 They concluded that the gene associated with susceptibility to myocardial infraction was probably located between these two loci The Disatrophic Dysplasia Gene Encodes a Novel Suflate Transporter: Positional Cloning by Fine-Structure Linkage Disequilibrium Mapping Cell 78: 1073-1087, 1994 Diastrophic dysplasia (DTD) (彎曲發育不全) One form of Osteochondrodysplasia Constitutional disorders of skeletal system result in disturbed growth and/or density of bone First described by Lamy and Maroteaux in 1960 Autosomal recessive Diastrophic dysplasia)(DD) (彎曲發育不全) Short-limbed short stature, kyphoscoliosis(脊柱 後側彎), generalized dysplasia of the joints, peculiar flexion limitation of the finger joints, hitchhiker thumbs, metatarsus adductus (蹠內收) deformity of the feet, and deformation of the ear lobes and cleft palate Joint changes are progressive in nature Osteoar’throses(非炎性骨關節病) and contractures develop at an early age Diastrophic dysplasia)(DD) (彎曲發育不全) Patients are severely physically handicapped and need repeated corrective orthopedic surgery Increased mortality in infancy due to respiratory difficulties and spine anomalies Intelligency and sexual development are normal Life expectancy is not clearly shortened Abnormal cartilage, number and morphology of chondrocytets, the organization of collagen fibrils, and the deficiency of sulfated proteoglycans Diastrophic dysplasia and D5S72 is Genetically Closely Linked No recombination between DD and D5S72 (Z()= 7.37) Principle of Linkage Disequilibrium (LD) Mapping Among a collection of chromosomes carrying the same ancestral mutation, genetic markers nearest the disease gene will have recombined least often and thus should show the highest degree of allelic association with the disease on such chromosomes How to Do Linkage Disequilibrium Mapping Determine the haplotypes of disease-bearing chromosomes for an extremely dense collection of genetic markers Use these haplotypes to identify a subset of chromosomes likely to carry a common ancestral mutation Find the genetic markers that show the strongest allelic association with the disease on these chromosomes Genetic Linkage studies of DTD 18 multiplex and 59 singleton families (a total of 144 DTD-baring chromosomes) DTD was initially localized to the interval between GRL and SPARC on chromosome 5q CSF1R gene: 1-1 haplotype at these two marker loci was present on 95% of DTD-baring chromosomes, but only 4% of normal chromosomes (RFLP analysis) Physical map shows locations and direction of transcription AC3: STS from the 3’ end of CSF1R BT1: CA-repeat (SSR) STS content mapping of the clones Known genomic organization of PDGFRB-CSF1R Genetic map ordered by recombinational mapping DTD was estimated to lie about 0.064 cM from CSF1R Genetic map order of each pair of adjacent loci supported by obligate crossovers with exchange of flanking markers either in CEPH pedigrees or DTD families GRL-ADRB2-D5S413-D5S372-BT1-CSF1RPDGRRB-RPS14-SPARC-D5S72 Physical map shows locations and direction of transcription AC3: STS from the 3’ end of CSF1R BT1: CA-repeat (SSR) STS content mapping of the clones Known genomic organization of PDGFRB-CSF1R Number of Normal and DTD chromosomes carrying Alleles from Finnish Ancestral DTD Haplotype Most DTD-bearing chromosomes carrying a single predominant allele at each of the 11 markers studied 95% of DTD-bearing chromosomes with haplotype “1-1-1-1-1-1-1-1-1-1-1” Linkage Disequilibrium Mapping P = (P affected – P normal)/(1-P normal) ≒α(1-θ)g P directly related to proximity to the disease locus α = proportion of disease-bearing chromosomes g = the number of generations since the introduction of ancestral chromosome excess Linkage Disequilibrium in DTD Region Clues Regarding the Location of DTD: Proximal to CSF1R P excess falls rapidly a the PDGFRb-CSF1R, but remains high at the marker BT1 proximal to CSF1R. Haplotype pattern of 144 DTD chromosomes Single DTD chromosome lack ancestral allele at the genetic markers within the PDGFRB and CSF1R and present at the more proximal genetic markers BT1 and D5S372. Gene Identification of Region 100 Kb Proximal to CSF1R Exon trapping of P1-1013, P1-1014 Direct cDNA selection Analysis of genomic fragments from this region Clones (JH10140B, 0.8 kb fragment from centromeric end of P1-1014, show strong aa similarity to the 5’ end of rat gene (sat-1, sulfate transporter) from BLAST program Expression Profile of Candidate DTD Gene Northern blot analysis Probe: 0.8 Kb JH10140B DTD 8.4 kb mRNA is Missing from DD Patients DTDST gene contains two exons separated by 1.8 kb intron DRA gene (down-regulated adenoma) from subtractive hybridization between colon and colon carcinoma DTDST Shows 12 Transmembrane Helices (hydropathy analysis) Two potential N-glycosylation sites at the N-terminal extracellular loop Confirmation of Mutation Normal Normal SP Family FV Family JD IM Codon 575 (AAG→AG, Dde I eliminated) SA(AG→AC, BsaAI created) 100 bp(Dde I+) → 40 bp + 60 bp 1000 bp→ 2 x 500 bp Summary of Mutations Transmission disequilibrium test (TDT) Association studies with internal controls Couples who have one or more affected offspring, it is irrelevant wheterh either parents is affected or not To test whether marker alleles M1 is associated with the disease Those parents who are heterozygous for M1 Let a be the number of times a heterozygous parent transmits M1 to the affected offspring, and b be the number of times the other allele is transmitted. TDT test statistic is (a-b)2/(a+b) Stevens-Johnson Syndrome (SJS) or Toxic Epidermal Necrosis (TEN) In 1922, Stevens and Johnson described two children with fever and stomatitis, severe disseminated conjunctivitis and cutaneous eruptions The reaction develops within 1-4weeks from the beginning of drug therapy Mucosal lesion includes lesions of the mouth, eyes, GI, respiratory tract, anus and vagina Drugs associated with SJS and TEN Sulfonamides, anticonvulsant agents Allopurinol, are the most consistently associated with SJS and TEN Nonsteroidal antiinflammatory drugs (NSAIDs) Analgestic agents Nonsulfonamides antibiotics, controversial? Typical pattern of StevensJohnson syndrome (SJS) Blisters develop on widespread purpuric macules N Engl J Med 1333: 1600-1607, 1995 Typical pattern of toxic epidermal necrosis (TEN) Severe forms of SJS Extensive skin detachment and a poor prognosis (death rate 30~40%) Blisters and wrinkled areas result from full-thickness necrosis o A marker for Stevens-Johnson Syndrome carbamazepine, treatment for seizures(癲癎) 44 CBZ-SJS patients 101 CBZ-tolerant (93 Normal controls) Diagnostic criterion of SJS/TEN were based on clinical morphology SJS: skin detachment of 10%, TEN: skin detachment of 30%, Overlapping SJS/TEN: skin detachment of 1030% Nature 428: 486, 2004 Genotyping methods HLA-A, -B, -C, DRB1(sequence-specific oligonucleotide probe, SSOP) cytochrome p450 (157 Single nucleotide polymorphism); What is single nucleotide polymorphism ? ATG(M)→ATA(I); TTC(F) →ATC(I)) Nature 428: 486, 2004 Nature 428: 486, 2004