* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Molecular modeling of HIV-1 reverse
Silencer (genetics) wikipedia , lookup
Clinical neurochemistry wikipedia , lookup
Proteolysis wikipedia , lookup
RNA polymerase II holoenzyme wikipedia , lookup
Drug design wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Eukaryotic transcription wikipedia , lookup
Catalytic triad wikipedia , lookup
Genetic code wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Biochemistry wikipedia , lookup
Specialized pro-resolving mediators wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
Protein structure prediction wikipedia , lookup
Biosynthesis wikipedia , lookup
Metalloprotein wikipedia , lookup
Enzyme inhibitor wikipedia , lookup
Point mutation wikipedia , lookup
Discovery and development of neuraminidase inhibitors wikipedia , lookup
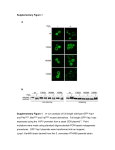
Protein Engineering vol.10 no.12 pp.1379–1383, 1997 Molecular modeling of HIV-1 reverse transcriptase drug-resistant mutant strains: implications for the mechanism of polymerase action Marilyn B.Kroeger Smith1,*, Christopher J.Michejda1, Stephen H.Hughes1, Paul L.Boyer1, Paul A.J.Janssen2, Koen Andries2, Robert W.Buckheit, Jr3 and Richard H.Smith, Jr1,4 A computer model of human immunodeficiency virus type 1 (HIV-1) reverse transcriptase (RT) either alone, or complexed with a non-nucleoside inhibitor (NNI), was constructed using crystal coordinate data from a subset of the protein surrounding the binding pocket region. Molecular mechanics calculations were carried out on solvated wildtype RT and RT that contained modifications corresponding to resistance-engendering mutations. Results from the calculations revealed that the r.m.s. difference between 12 modified proteins and that of wild-type RT could be qualitatively correlated with the measured polymerase activity of the enzyme in the presence of these mutations. In addition, the level of activity was related to the measured distance between the primer grip and dNTP binding regions of the protein. These data suggest a direct correlation between RT structure and function. Complexes of RT–8C1 TIBO and RT–α-APA were also minimized in models containing modifications corresponding to key drug-resistant mutants. The variant complexes all showed weaker binding than wild-type RT, while giving rise to similar, but critical changes in the protein. Therefore, the design of new inhibitors should center on obtaining stronger binding drugs to key drug-resistant RT variants. Keywords: AIDS/computer/inhibitor/resistance/reverse transcriptase nucleoside inhibitors (NNIRTs) results in the development of resistance, although the specific mutants that are responsible differ for the most part between the two classes of drugs (Boyer et al., 1994a,b). In order to combat this problem, combination therapy, which combines inhibitors from both classes, is now being used as a means to enhance virus suppression (Larder and Kemp, 1990; St Clair et al., 1991; Larder, 1992; Tisdale et al., 1993). One such combination involved administration of zidovudine (AZY) along with a second nucleoside analog such as ddI (Iversen et al., 1996). Alternative treatment strategies involve concomitant treatment with either one or more nucleoside and/or non-nucleoside RT inhibitors, two non-nucleoside drugs or RT inhibitors and HIV-1 protease inhibitors. Selection of compounds for these treatments should involve inhibitors that have different resistance spectra with the goal of obtaining escape mutants that replicate poorly. As an aid to understanding how mutations affect RT structure and function, in both the presence and absence of nonnucleoside inhibitors, molecular modeling studies were undertaken on RT modified at specific residues. Assays to determine polymerase activity of the mutant strains of RT showed that while some of these strains had levels of activity comparable to that of the wild-type enzyme, other mutant strains had much lower levels of enzymatic activity. Correlation of structure with enzymatic function in RT without inhibitor present may lead to an explanation for the experimental findings. It has been suggested that in the presence of NNIRTs, a deformation of the polymerase active site occurs, locking it into an inactive conformation (Ren et al., 1995). Presumably, mutation may either restore the active site conformation through structural realignment or could destabilize inhibitor binding. Therefore, assessment of the structural adjustment resulting from these local changes and the accompanying energetic consequences may provide insight into the design of more effective RT inhibitors. Introduction Reverse transcriptase (RT) of human virus type 1 (HIV-1) is an important enzyme for drug therapy because it plays a critical role in the viral replication cycle. RT, which is virally encoded, converts the single-stranded viral RNA genome into a double-stranded linear DNA intermediate that is subsequently integrated into the host cell DNA (Goff, 1990; Mitsuya et al., 1990). The success of treatment of HIV-1 with inhibitors that target RT has been limited by the development of resistance of the virus to the various drugs (Larder and Kemp, 1990; St Clair et al., 1991; Richman et al., 1994; Zhang et al., 1994). The reasons behind the eventual failure of the inhibitors include the large numbers of virions in the patient and their rapid turnover, in addition to continual mutation in the virus (Coffin, 1995). Treatment of patients with either nucleoside or non- Materials and methods Computational methods All molecular modeling of the enzyme complexes was carried out using Insight II software (MSI), with all calculations performed using the C force field (cff91) within the Discover module mounted on a Cray YMP-8 supercomputer at NCIFCRDC. The modeling calculations were based on the X-ray structure coordinates of the complexes of RT–DNA and RT–8-chloro-tetrahydro-imidazo(4,5,1-jk)(1,4)-benzodiazepin2(1H)-thione (8-C1 TIBO) (R86183) (Jacobo-Molina et al., 1993; Ding et al., 1995). For the model of the enzyme without inhibitor, only the protein coordinates were used. Calculations with α-anilino-2,6-dibromophenylacetamide (α-APA) were carried out by superimposing the inhibitor (in its non-minimized cognate complex) over 8-C1 TIBO (in its minimized complex) and reminimizing the resulting RT–α-APA site. At present, since there are no crystallographic data available for 1ABL-Basic Research Program, NCI-Frederick Cancer Research and Development Center, P.O. Box B, Frederick, MD 21702, USA 2Center for Molecular Design, Janssen Research Foundation, Antwerpsesteenweg 37, B2350 Vosselaar, Belgium, 3Virology Research Group, Southern Research Institute, Frederick, MD 21701 and 4Department of Chemistry, Western Maryland College, Westminster, MD 21157, USA *To whom correspondence should be addressed. © Oxford University Press 1379 M.B. Kroeger-Smith et al. mutant forms of RT either alone or in a complex with nonnucleoside inhibitors, computations involving RT variants were constructed by extrapolation from wild-type HIV-1 RT crystal data. The construction of a suitable computer model from the respective coordinate data has been documented previously (Kroeger Smith et al., 1995). The model site consisted of p51 subdomain residues 132–142 and residues 89–116, 156–211, 213–243, 265–271, 313–323, 346—351 and 380–384 from the p66 subdomain. The C-terminal ends of these strands were capped with a methylamino group and the N-terminal ends were capped with an acetyl group. The pH of the site was set to 7.5, resulting in a total charge of zero, with the charged residues including Lys1, Arg1, Asp– and Glu–. To simulate mutations in the protein, amino acid residues were changed using the Biopolymer module within Insight II. To account for solvation effects, all RT–inhibitor complexes were surrounded with a ~6 Å layer of water prior to minimization (the thickness of the water layer was adjusted in order to obtain 1283 water molecules surrounding each complex), using the higher cvff partial charges for water (O, 20.820; H, 0.410). Calculations were carried out in three stages, with each stage consisting of steepest descent minimization followed by a conjugate gradient minimization until the r.m.s. deviation was ø0.001 Å. The protein was divided into two layers: a primary layer (~10 Å from the inhibitor), which was constrained during initial solvent minimization and then progressively freed (side chain first, followed by backbone) during subsequent stages of the minimization, and a secondary layer (~10–20 Å from the inhibitor) that was held fully constrained throughout the calculations (for the specific residues in these two layers Kroeger Smith et al., 1995). The solvent molecules changed position between ~0.75 and 3.0 Å during the course of the minimization. The resultant binding energy for each RT–inhibitor complex obtained from the minimizations was taken to be the nonbonded interaction energy resulting from the formation of the enzyme–inhibitor complex from the initial inhibitor and enzyme reactant species. The magnitude of the drug–protein interaction energy, in addition to its van der Waals and electrostatic components, was measured using the Docking module within the Insight II package. Biochemical procedures Construction of the expression vector for RT (Hughes et al., 1990; Jacobo-Molina et al., 1993) and the preparation of clones expressing individual drug-resistance encoding mutations in the p66 subunit and a wild-type p51 subunit (Boyer et al., 1994a,b) were described previously. RNA-dependent DNA polymerase activity was assayed in Escherichia coli extracts as described previously (Boyer et al., 1992). The cytoprotective activity of 8-C1 TIBO or α-APA against either wild-type virus or strains containing defined mutations in RT was evaluated in a screening assay described earlier (Weislow et al., 1989). Results and discussion Modeling studies on HIV-1 RT containing amino acid substitutions The structure of wild-type HIV-1 RT is represented by a model containing 155 amino acid residues surrounding the nonnucleoside binding pocket. Minimizations on a subset of the enzyme were undertaken owing to memory limitations. It should be recognized that calculations performed on only part 1380 Table I. Results from calculations on mutant variants of RT-DNA RT strain Polymerase activity (%)a Distance (primer grip to dNTP) (Å)b r.m.s., difference (Å)c Wild-type L100I K101E K103N V106A D110E V179D Y181C D185E D186E Y188H G190E E138K 100 40 140 100 80 ,5 100 115 ,5 ,5 75 20 150 11.7 10.7 11.3 11.6 11.5 10.8 11.7 11.1 10.2 10.5 11.0 10.9 11.2 – 0.6 0.4 0.4 0.5 0.5 0.5 0.4 0.6 0.7 0.5 0.6 0.4 aData for p66 homodimer reported as % of wild-type RT bThe measured distance is from G231:CA to D186:CA. activity. cThe difference is a comparison for the mutant and wild-type strains of RT, as measured by backbone superposition. of the enzyme are not necessarily representative of the response of the entire protein and, as such, are subject to errors which may permeate throughout the study in a differential manner. However, since the aim of the study was to see if computations would produce quantative trends that might shed light on the mechanistic effects of mutations on polymerase activity and resistance to NNIRTs, it was felt that the method was valid within these limits. This model was modified separately at each of 12 key amino acid residues (see Table I) and the resultant sites were subjected to molecular mechanics energy minimization. Nine of the amino acid residues that were modified in the model are representative of changes observed in drug-resistant strains that emerge either in vitro (cell assays) (Kleim et al., 1996) or in vivo (patients) (Moermans et al., 1995) following treatment with non-nucleoside inhibitors. The other three amino acid mutations that were analyzed, D110, D185 and D186, compose the triad of aspartyl residues in the polymerase active site and are crucial for polymerase function. These were included in the calculations since they show that removal of one of aspartic acid residues disrupts the site. The results from the calculations on these modified RT sites are shown in Table I, along with the measured polymerase activity of wild-type RT and the various drug-resistant strains. The r.m.s. differences (measured by backbone superposition of all 155 residues) of the modified RTs, as compared with the wild-type starting structure, are also shown in Table I. These overall measures of difference were found, in general, to correlate with the measured polymerase activities of the mutant enzymes. RT drug-resistant strains that showed substantially reduced polymerase activity had r.m.s. values .0.5 Å, while those variants that maintained near wild-type activity had r.m.s. values ,0.5 Å. Two notable exceptions are the V179D and D110E changes, which both showed r.m.s. values of 0.5 Å. Clearly, measurement of r.m.s. differences is only a crude measure of possible disruption of the geometry at the polymerase active site. A more detailed analysis is necessary to elucidate the mechanism that underlies these differences. The mutation that caused the greatest local structural change was the G190E substitution. In this case, structural differences were especially evident in the β12–β14 sheet which contains the amino acid Molecular modeling of HIV-1 reverse transcriptase Fig. 1. Chart of the distance in Å between an amino acid residue in the dNTP binding site region (D186) and one in the primer grip region (G231) vs the polymerase activity for various mutant RTs. residue W229. This residue appears to play an essential role in the polymerase process since no viable mutations at that position have been found to date (Jaques et al., 1994; P.L.Boyer, H.Q.Gao and S.H.Hughes, in press). The displacement observed for the CB atom of this residue (1.52 Å) was found to be respresentative of the alteration of the entire backbone chain. The sheet was also displaced when each of the other amino acid substitutions were made, although to a lesser extent. Other notably large residue adjustments were as follows: for the V179D mutation, the CB atom of F227 moved 1.8 Å compared with its position in wild-type; for the Y188H change, the backbone residues for K101 to V106 shifted ~1.0 Å; and for the K101E modification, the CB atom of Y183 moved 0.9 Å. A more direct measure of this geometry alteration is the distance between the D186 residue (CA atom) near the dNTP binding site and a reference residue in the primer-grip region (Jacobo-Molina et al., 1993) of RT (G231:CA). Reduction of this distance to ,11.0 Å (wild-type 11.7 Å) appears to lead to polymerase activities less than 50% of wild-type (Table I, Figure 1), although slightly less reduction in some cases leads inexplicably to an increase in polymerase activity. In order to test that the default placement of the mutant residues by the software program is not artificially influencing the results of the calculations, the position of the isoleucine side chain of amino acid 100, for example, was adjusted to an alternative initial orientation. The measured distance of 10.5 Å following minimization was comparable to that found from the first calculation (10.7 Å), ruling out system artifacts. The major effect of the mutations appears to be a local distortion in the position of the catalytic triad (as opposed to major changes in the backbone atoms of the β12–β14 sheet), with the CB atom of D186, for example, moving 0.3–1.2 Å. This adjustment is largest (1.2 Å) in the case of D186E substitution, followed by the G190E and K101E modifications (~1.0 Å). For mutations that retain a level of polymerase activity comparable to wildtype (with the exception of K101E noted above), the D186:CB displacement distance is smaller (,0.8 Å). Correlation of RT structure and function The locations of the residues in RT that were modified prior to minimization, together with the corresponding level of polymerase activity for the mutant RTs, are shown in Figure 2. From the data in Table I, it is apparent that the importance Fig. 2. Representation of the location and polymerase activity of key drugresistant variants in wild-type RT–DNA. Amino acid residues shown in red maintain 100% of the wild-type activity upon mutation, those in blue 50– 100% and those in green ,50%. of these residues with respect to polymerase activity falls into three classes. Of utmost importance are the aspartic acid triad residues, D110, D185 and D186 (shown in green), which play a direct role in the polymerase activity of the enzyme. Mutation in any of these residues results in a drop in polymerase activity to ,5% of wild-type. In addition, the D110E, D185E and D186E mutations reduce the dNTP to primer grip distance by 0.9–1.3 Å, more than that for any other modification we have tested. Among the key ramifications of these aspartic acid mutations is that lengthening the carboxylate-bearing side chain will result in substantial movement of the Mg21 ion to which it is complexed (Steitz and Steitz, 1993; Patel et al., 1995). Hence the location of the dNTP that in turn complexes with this Mg21 ion through its α-phosphate would be altered from that found in the wild-type enzyme. The end result is that the E186 variant would be less able to process the incoming dNTP. Mutation of amino acid residues L100 and G190 (also shown in green) produces enzymes that retain only 20–40% of wild-type polymerase activity. In these cases, the dNTPprimer grip distance is significantly shortened (see Table I). This change could interfere with the ability of the incoming dNTP to be appropriately positioned to permit sugar–phosphate bond formation. Although no key function has yet been postulated for L100 in either unliganded RT or RT complexed with DNA, examination of the structures shows that in its location on the β5–β6 connecting loop, L100 is situated between Y181 and Y188, making important van der Waals contacts with both residues. Hence, in the absence of inhibitor binding, which causes the reorientation of the side chains of Y181 and Y188 as well as F227 and W229 during the formation of the inhibitor binding pocket (Hsiou et al., 1966), L100 is clearly crucial to the integrity of the folded protein. Replacement of leucine with isoleucine could account for substantial structural rearrangement which is reflected in the 1381 M.B. Kroeger-Smith et al. distortion of the polymerase active site geometry and results in a marked reduction in polymerase activity. Interestingly, we have also found that the polymerase activity of an RT carrying both the L100I and the K103N mutations is 85% of that observed for wild-type enzyme. Modeling of this variant showed that only a small decrease in the distance between the primer grip and dNTP binding site occurred compared with wild-type (0.1 Å). Turning to the G190 residue, it can be seen from the model that the substitution of glutamic acid for glycine leads to significant steric hinderance with Y188, with the resultant structural reorganization again compromising the critical geometry of the polymerase site. This explanation is in keeping with the suggestion by Chao et al. (1995) that the G190E mutation impairs protein folding of the recombinant enzyme, as indicated by a reduction in solubility. In our hands, however, a recombinant HIV-1 RT containing the G190E mutation was readily soluble. The V106 and Y188 mutations, shown in blue, retain at least 75% of the original activity upon mutation. These residues show slightly reduced dNTP–primer grip separation distances, in keeping with their relatively high levels of polymerase activity. V106 makes van der Waals contact with Y188, suggesting that changes in these residues affect the integrity of the folded protein, but the distortion of the geometry of the dNTP binding site is less severe than with other mutations. Some mutations (shown in red) do not lead to a measurable reduction in polymerase activity in our assay. K103N and V179D have activities identical with that of wild-type RT. Consistent with these data, alterations to those two residues do not markedly change the geometry of the polymerase active site, as measured by overall r.m.s. deviations and the dNTP– primer grip distance. Other changes such as K101E, Y181C and E138K actually lead to an increase in polymerase activity, although this should not be taken to mean that they are more efficient enzymes than wild-type. Other processes, such as fidelity, processivity, and/or strand transfer, may have defects that are not readily measurable by a straightforward polymerase assay. For example, the E138K mutant enzyme shows a significantly lower Vmax and a lower Km for poly (rC)·oligo (dG) or dGTP (P.L.Boyer, H.Q.Gao and S.H.Hughes, in press), enabling it to incorporate more dNTP than the wild-type enzyme. The effect of these latter mutations is a slight compression of the dNTP–primer grip distance, 11.2 6 0.1 Å, possibly suggesting that polymerization efficiency under the specific conditions used in the in vitro assay may be enhanced by this type of compression. The polymerase activity data shown in Figure 2 thus provide an informative picture of the correlation between structure and function in RT. When mutations arise in important residues in the enzyme, a displacement of catalytic site residues occurs which reduces the ability of the enzyme to incorporate the incoming nucleotide triphosphate. Modeling of RT–inhibitor complexes containing amino acid modifications Computer modeling studies were also undertaken of RT– inhibitor complexes that contained one of the amino acid substitutions (Y181C, K101E, or K103N) seen to emerge following treatment of patients with 8-C1 TIBO (Moermans et al., 1995). Following minimization of the complexes containing variant amino acids, the backbone r.m.s. values of these structures as compared with the wild-type were ~0.4 Å, with the majority of the deviations being local changes directly surrounding the site of mutation. 1382 Table II. Interaction energy and EC50 values for NNIs against drug-resistant strains of RT RT strain 8-CI TIBO–RTa (EC50b) α-APA–RTa (EC50b) Wild type Y181C K101E L100I K103N –54.0 –53.2 –50.8 –49.5 –49.5 263.5 (0.4) –52.3 (1.0) –58.3 (0.2) –61.4 (0.08) –55.7 (0.7) (0.3) (4.2) (17.4) (17.4) (17.4) aEnergies bValues in kcal mole–1. for EC50 in µM (in parentheses). The drug–protein interaction energy for 8-C1 TIBO or αAPA in the 8-C1 TIBO site (wild-type and variant) was measured following the minimizations. This value is useful in the evaluation of changes in the strength of inhibitor–protein binding among different complexes. The use of the same starting RT–inhibitor complex for all the minimizations eliminated any inherent error associated with structural anomalies between different crystal structures. In the present study, this energy value was seen to be more positive (hence less favorable) for all the variant RTs as compared with wild-type, whether RT was complexed with α-APA or 8-C1 TIBO (Table II). In addition, the order of the energy values corresponds not only to the measured EC50 values for 8-C1 TIBO and α-APA against wild-type RT, but also against the four drug-resistant strains (see Table II). Measurement of the dNTP–primer grip distance following modification of the complexed protein showed only small changes from those determined for inhibitor complexes with wild-type enzyme (15.93 Å), with distances generally 0.12– 0.22 Å shorter than in wild-type. L100I was the only mutant for which the changes seen upon drug binding were greater than in wild-type, with increases of 0.16 Å for 8-C1 TIBO and 0.57 Å for α-APA. Implications for the design of non-nucleoside inhibitors with a broader spectrum of activity No crystallographic data were available for complexes between non-nucleoside inhibitors with mutant forms of RT. In the absence of such structural data, calculations were based on wild-type RT–inhibitor complexes. In our previous studies where these complexes were used as models (Kroeger Smith et al., 1995; Smith et al., 1998), indirect evidence was obtained that binding of various non-nucleoside inhibitors interferes with polymerase activity in a straightforward manner. Specifically, the dNTP binding site to primer grip distance (as represented by residues D110, D185, D186 and G231) was correlated with the potency of the inhibitor against wild-type RT. It was found that the greater the separation of the two sites, the better was the EC50 value of the drug, with all of the distances for the complexed enzyme being substantially greater than that for RT alone. The resulting spatial rearrangements in the protein could reasonably be expected to impede the formation of the sugar phosphate bond. Hence we proposed a more detailed explanation of the allosteric mechanism of inhibition by the non-nucleoside compounds (Kroeger Smith et al., 1995). From data in the current study, complexes of inhibitors with variant forms of RT show distortions in the dNTP–primer grip separation similar to wild-type (data not shown). In previous work (Smith et al., 1998), where α-APA, 8-C1 TIBO and Molecular modeling of HIV-1 reverse transcriptase nevirapine were modeled in their cognate sites containing the Y181C mutation, this same distance was slightly shortened (0.1–0.5 Å as compared with the wild-type enzyme). In concert, it has recently been shown experimentally that none of the inhibitors examined here inhibit variant RT nearly as well as they arrest the wild-type enzyme (data not shown). In support of this idea, recent kinetic studies have shown (Spence et al., 1996) that the Kd of the non-nucleoside inhibitor nevirapine for RT carrying the Y181C mutation increased 500-fold compared with the wild-type RT complex. This study suggests that a similar mechanism (i.e. reduced binding to the various mutant forms of RT) might also be operative in the RT–TIBO complex aside from active site distortion. Contributing to the decreased binding affinity could be loss of aromatic stacking interactions, van der Waals non-bonded interactions, and/or coulombic (electrostatic) attraction. Depending on the specific amino acid change, the subsequent interaction of these modified residues with inhibitors of varying structure would result in a differing balance in the reduction of these types of interactions. Hence, in order for a substance to be an effective inhibitor of the polymerase activity of either wild-type or mutant HIV-1 RT, it must both bind tightly to the enzyme and, upon binding, give rise to structural changes in the protein that result in low polymerase activity. Two clinically observed mutational changes that appear to have important implications for RT inhibition are G190E and L100I. While the exact role of these two residues is unknown, they are clearly important for RT function (as seen from polymerase activity data) and in addition, play an important role in complexation with non-nucleoside inhibitors. Aside from inhibitor binding, it appears from their location that mutation of these residues might also interfere sterically (and electrostatically in the case of G190E) with key contacts along an entrance route that the inhibitors are believed to travel during the process of binding. Additional support for a key role of position of residue 190 comes from in vitro studies with the non-nucleoside inhibitor HBY 079, where five out of six resistant strains displayed a G190E mutation following treatment with the drug (Kleim et al., 1996). In the same paper the authors noted that the double mutant L100I/K103N may result in the same phenotype as that seen with the G190E mutant. Resistance, which develops following treatment with nonnucleoside inhibitors of RT, is the key factor undermining the ability of these drugs to lower viral load. While a drug that effectively inhibits all forms of RT (wild-type and variant) would be an ideal goal, a highly satisfactory alternative would be one that selects for weak mutant variants that replicate less well than the wild-type. With respect to either objective, an understanding of the structural consequences of mutations in RT in its unliganded and complexed states is vitally important. With this information at hand, one strategy to increase the activity of known inhibitors against variant forms of the virus would be to modify current inhibitors chemically with the aim of enhancing the hydrophobic character of the π-electronrich components of the molecule. The addition of a bulky, hydrophobic group to the inhibitor would fill the void left by the loss of an aromatic residue, such as Y181. A second tactic would be to design a chemically modified inhibitor that would covalently bind to an essential amino acid residue, such as W229. Hopefully, modifications such as these will lead to inhibitors that have a broad-based spectrum of action. Acknowledgements The authors thank Dr Edward Arnold for critical reading of the manuscript and Valerie Fliakas-Boltz for performing the anti-HIV-1 assays. In addition, they acknowledge the Frederick Biomedical Supercomputing Center of the Frederick Cancer Research and Development Center for access to the CrayYMP computer. Research sponsored input by the National Cancer Institute, DHHS, under contract with ABL. The contents of this publication do not necessarily reflect the views or policies of the Department of Health and Human Services, nor does mention of trade names, commercial products, or organizations imply endorsement by the US Government. References Boyer,P.L., Ferris, A.L. and Hughes, S.H. (1992) J. Virol., 66, 1031–1039. Boyer,P.L., Ding,J., Arnold,E. and Hughes, S.H. (1994a) Antimicrob. Agents Chemother., 38, 1909–1914. Boyer,P.L., Tantillo, C., Jacobo-Molina, A., Nanni, R.G., Ding, J., Arnold, E., and Hughes,S.H. (1994b) Proc. Natl Acad. Sci USA, 91, 4882–4886, Chao,S.-F., Loong Chan,V., Juranka,P., Kaplan,A.H., Swanstrom,R and Hutchinson, C.A. (1995) Nucleic Acids Res., 23, 803–810. Coffin,J. (1995) Science, 267, 483–489, Ding,J., Das,K., Moereels,H., Koymans,L., Andries,K., Janssen,P.A.J., Hughes,S.H. and Arnold,E. (1995) Nature Struct. Biol., 2, 407–415. Goff,S.P. (1990) J. Acquired Immune Defic. Syndr., 3, 817–831, Hsiou,Y., Ding,J., Das,K., Clark,A.D., Jr, Hughes,S.H. and Arnold,E. (1996) Structure, 4, 853–860. Hughes,S.H., Ferris,A. and Hizi,A. (1990) In Laver,W.G. and Air,G.M. (eds) Use of X-Ray Crystallography in the Design of Antiviral Agents. Academic Press, New York, pp. 297–307. Iversen,A.K.N., Shafer,R.W., Wehrly,K., Winters,M.A., Mullins,J.L., Chesebro, B. and Merigan,T.C. (1996) J. Virol., 70, 1086–1990. Jacobo-Molina, A. et al., (1993) Proc. Natl Acad. Sci. USA, 90, 6320–6324. Jacques,P.S., Wohrl,B.M., Ottmann,M., Darlix,J.-L. and Le Grice,S.F. (1994) J. Biol. Chem., 269, 26472-26478. Kleim,J.-P., Rosner,M., Winkler,I., Paessens,A., Kirsch,R., Hsiou,Y., Arnold,E. and Riess,G. (1996) Proc. Natl Acad. Sci. USA, 93, 34–38. Kroeger Smith,M.B. et al. (1995) Protein Sci., 4, 2203-2222. Larder,B.A. (1992) Antimicrob Agents Chemother., 36, 2664–2669. Larder,B.A. and Kemp,S.D. (1990) Science, 246, 1155–1158. Mitsuya,H., Yarchoan,R, and Broder,S. (1990) Science, 249, 1533–1544. Moermans,M. et al., (1995) Abstracts of the Fourth International Workshop on HIV-1 Drug Resistance, Sardinia, Italy, July 6–9. Patel,P.A., Jacobo-Molina,A., Ding,J., Tantillo,C., Clark,A.D., Jr, Raag, R., Nanni,R.G., Hughes,S.H. and Arnold,E. (1995) Biochemistry, 34, 5351– 5356, Ren,J., Esnouf,R., Hopkins,A., Ross,C., Jones,Y., Stammers,D. and Stuart,D. (1995) Structure, 3, 915–926. Richman,D.D. et al., (1994) J. Virol., 68, 1660–1666. Smith,R.H., Jr, Michejda,C.J., Hughes,S.H., Arnold,E., Janssen,P.A.J. and Kroeger Smith,M.B. (1998) J. Struct.Biol., (Theochem), in press. Spence,R.A., Anderson,K.S. and Johnson,K.A. (1996) Biochemistry, 35, 1054–1063. St Clair,M.H. et al. (1991) Science, 253, 1557–1559. Steitz,T.A. and Steitz,J.A. (1993) Proc. Natl Acad. Sci. USA, 90, 6498–6502. Tisdale,M., Kemp,S.D., Parry,N.R. and Larder,B.A. (1993) Proc. Natl Acad. Sci. USA, 90, 5653–5656. Weislow,O.S., Kiser,R., Fine,D.L., Bader,J., Shoemaker,R.H. and Boyd,M.R. (1989) J. Natl Cancer Inst., 81, 577–586. Zhang,D., Caliendo,A.M., Eron,J.J., DeVore,K.M., Kaplan,J.C., Hirsch,M.S. and D’Auuila,R.T. (1994) Antimicrob. Agents Chemother., 38, 282–287. Received April 29, 1997; revised July 25, 1997; accepted September 5, 1997 1383