* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
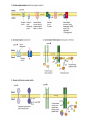
Download Figure 20-5. Common intracellular signaling proteins.
Survey
Document related concepts
Purinergic signalling wikipedia , lookup
Endomembrane system wikipedia , lookup
Phosphorylation wikipedia , lookup
Hedgehog signaling pathway wikipedia , lookup
Protein moonlighting wikipedia , lookup
NMDA receptor wikipedia , lookup
Intrinsically disordered proteins wikipedia , lookup
List of types of proteins wikipedia , lookup
Protein phosphorylation wikipedia , lookup
Mitogen-activated protein kinase wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Transcript
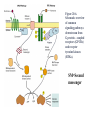
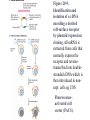
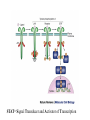
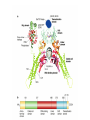
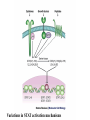
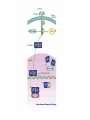
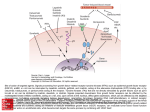
http://ncbi.nlm.nih.gov/entrez/query Books Molecular Cell Biology Lodish 20. Cell-to-Cell Signaling: Hormones and Receptors http://backserv.imbb.forth.gr CYTOKINES: Soluble factors required for hematopoietic Cell growth and differentiation IFN family gp130 gp 140 γc chain IFNα/β IL-6 IL-3 IL-2 IFNγ !L-11 IL-5 IL-4 IL-10 LIF GMCSF IL-7 CNTF IL-9 IL-15 Figure 20-5. Common intracellular signaling proteins. (a) GTP-binding proteins with GTPase activity function as molecular switches. When bound to GTP they are active; when bound to GDP, they are inactive. They fall into two categories, trimeric G proteins and Ras-like proteins(b) Protein kinases modulate the activity or the binding properties of substrate proteins by phosphorylating serine, threonine, or tyrosine residues. The phosphorylated form of some proteins is active, whereas the dephosphorylated form of other proteins is active. The combined action of kinases and phosphatases, which dephosphorylate specific substrates, can cycle proteins between active and inactive states. (c) Adapter proteins contain various proteinbinding motifs that promote the formation of multiprotein signaling complexes. Figure 20-6. Schematic overview of common signaling pathways downstream from G protein – coupled receptors (GPCRs) and receptor tyrosine kinases (RTKs). SM=Second messenger Figure 20-9. Identification and isolation of a cDNA encoding a desired cell-surface receptor by plasmid expression cloning. All mRNA is extracted from cells that normally express the receptor and reversetranscribed into doublestranded cDNA which is then introduced in nonexpr. cells eg COS Fluorescenceactivated cell sorter (FACS). Figure 15-47. Six subfamilies of receptor tyrosine kinases. Only one or two members of each subfamily are indicated. Note that the tyrosine kinase domain is interrupted by a "kinase insert region" in some of the subfamilies. The functional significance of the cysteine-rich and immunoglobulinlike domains is unknown. Figure 20-21. General structure and activation of receptor tyrosine kinases (RTKs). The ligands for some RTKs, such as the receptor for EGF depicted here, are monomeric; ligand binding induces a conformational change in receptor monomers that promotes their dimerization. The ligands for other RTKs are dimeric; their binding brings two receptor monomers together directly (see Figure 20-4d). In either case, the kinase activity of each subunit of the dimeric receptor initially phosphorylates tyrosine residues near the catalytic site in the other subunit. Subsequently, tyrosine residues in other parts of the cytosolic domain are autophosphorylated. See text for discussion. [See G. Panayotou and W. D. Waterfield, 1993, Bioessays 15:171; M. Mohammadi et al., 1996, Cell 86:577 Figure 15-57. Some of the protein kinases discussed in this chapter. The tyrosine kinases shown are bound to the plasma membrane, whereas most of the serine/threonine kinases are in the cytosol. Cytokines in the Immune system STATs and SMADs: Linking receptors to transcription. Pawson T and Nash P (2000). Genes Dev.,1027 STAT= Signal Transducer and Activator of Transcription Three examples of signaling in the JAK-STAT pathway. Specific ligandreceptor interactions generate active transcription complexes composed of distinct STAT proteins. GAS= gamma activated sequence Variations in STAT activation mechanisms Structural features of Src-kinases, JAKs and SOCS/CIS/JAB protein family proteins.