Problems in Replication and Protein Synthesis
... Eukaryotic Regulation • Enhancers – sites that call for specific activators to stimulate the production of certain proteins. ...
... Eukaryotic Regulation • Enhancers – sites that call for specific activators to stimulate the production of certain proteins. ...
differential gene expression
... 2. Physically there are more obstacles as eukaryotic chromatin is wrapped around histone proteins. ...
... 2. Physically there are more obstacles as eukaryotic chromatin is wrapped around histone proteins. ...
Chapter 16
... is absent. If operator is bound, promoter region is partially blocked-genes can not be transcribed. • This two switch control mechanism thus causes the cell to produce only what the cell needs, when it needs it. ...
... is absent. If operator is bound, promoter region is partially blocked-genes can not be transcribed. • This two switch control mechanism thus causes the cell to produce only what the cell needs, when it needs it. ...
problem set
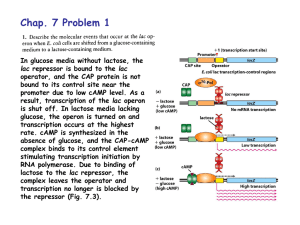
... bound to its control site near the promoter due to low cAMP level. As a result, transcription of the lac operon is shut off. In lactose media lacking glucose, the operon is turned on and transcription occurs at the highest rate. cAMP is synthesized in the absence of glucose, and the CAP-cAMP complex ...
... bound to its control site near the promoter due to low cAMP level. As a result, transcription of the lac operon is shut off. In lactose media lacking glucose, the operon is turned on and transcription occurs at the highest rate. cAMP is synthesized in the absence of glucose, and the CAP-cAMP complex ...
control of gene expression
... RNA polymerase binding) • -10 and –35 sequences and how conserved • Sigma factors • Whether or not a repressor protein is present • Enhancer/activator sequences • Once the transcript has been produced there is the opportunity for anti sense RNAs to get involved in control e.g. by binding to sense me ...
... RNA polymerase binding) • -10 and –35 sequences and how conserved • Sigma factors • Whether or not a repressor protein is present • Enhancer/activator sequences • Once the transcript has been produced there is the opportunity for anti sense RNAs to get involved in control e.g. by binding to sense me ...
Essential knowledge 3.B.1
... to DNA and blocking transcription (negative control). Regulatory proteins stimulate gene expression by binding to DNA and stimulating transcription (positive control) or binding to repressors to inactivate repressor function. Certain genes are continuously expressed; that is, they are always tur ...
... to DNA and blocking transcription (negative control). Regulatory proteins stimulate gene expression by binding to DNA and stimulating transcription (positive control) or binding to repressors to inactivate repressor function. Certain genes are continuously expressed; that is, they are always tur ...
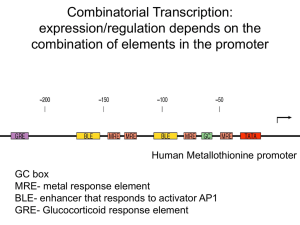
Combinatorial Transcription: expression/regulation depends on the
... domain is now free to form other domains with alternative boundary elements (in this case containing genes Z1and Z2). Enhancer 1(en1) is now unable to act on the promoter of gene Y because of the new location of the gypsy insulator. Nevertheless, this enhancer is still functional and competent to ac ...
... domain is now free to form other domains with alternative boundary elements (in this case containing genes Z1and Z2). Enhancer 1(en1) is now unable to act on the promoter of gene Y because of the new location of the gypsy insulator. Nevertheless, this enhancer is still functional and competent to ac ...
Slide 1 - AccessPharmacy
... Schematic diagram showing the transcription control regions in a hypothetical mRNA-producing, eukaryotic gene transcribed by RNA polymerase II. Such a gene can be divided into its coding and regulatory regions, as defined by the transcription start site (arrow; +1). The coding region contains the DN ...
... Schematic diagram showing the transcription control regions in a hypothetical mRNA-producing, eukaryotic gene transcribed by RNA polymerase II. Such a gene can be divided into its coding and regulatory regions, as defined by the transcription start site (arrow; +1). The coding region contains the DN ...
Control of Gene Expression
... • Each tissue in our body is very diff. despite having same DNA • Even identical twins have differences due to gene expression ...
... • Each tissue in our body is very diff. despite having same DNA • Even identical twins have differences due to gene expression ...
401Lecture8Sp2013post
... modular proteins composed of a DNA binding domain and an activation domain • Repressors inhibit transcription and are modular proteins composed of a DNA binding domain and a repressor domain • Both repressor and activators recruit other proteins to affect gene expression • A cell must produce the sp ...
... modular proteins composed of a DNA binding domain and an activation domain • Repressors inhibit transcription and are modular proteins composed of a DNA binding domain and a repressor domain • Both repressor and activators recruit other proteins to affect gene expression • A cell must produce the sp ...
Gene Expression
... • Introns and exons are transcribed to form PremRNA. Enzymes cut out the introns and join remaining exons together forming mRNA. This leaves the nucleus and travels through the nuclear pore to the cytoplasm where translation occurs. ...
... • Introns and exons are transcribed to form PremRNA. Enzymes cut out the introns and join remaining exons together forming mRNA. This leaves the nucleus and travels through the nuclear pore to the cytoplasm where translation occurs. ...
File - EUREKA! Science
... Bacterial Genes Bacteria have less DNA than other organisms Genes organized into operons Operon: region of DNA that includes a promoter, an operator, and the genes that code for the protein Found only in prokaryotes and round worms ...
... Bacterial Genes Bacteria have less DNA than other organisms Genes organized into operons Operon: region of DNA that includes a promoter, an operator, and the genes that code for the protein Found only in prokaryotes and round worms ...
A Tale of Three Inferences
... gene G, if T1 binds to c1 and T2 binds to c2 in an inductive way, then the expression of G will remain the same if the promoter were to have twice the number of c1 and c2 goes to 0. • Boolean AND: Under same conditions, there will be no expression ...
... gene G, if T1 binds to c1 and T2 binds to c2 in an inductive way, then the expression of G will remain the same if the promoter were to have twice the number of c1 and c2 goes to 0. • Boolean AND: Under same conditions, there will be no expression ...
Control of Gene Expression Control of Gene Expression Regulatory
... promoter region of the gene. • RNA polymerase II then binds to the promoter to begin transcription at the start site (+1). • Enhancers are DNA sequences to which specific transcription factors (activators) bind to increase the rate of transcription. ...
... promoter region of the gene. • RNA polymerase II then binds to the promoter to begin transcription at the start site (+1). • Enhancers are DNA sequences to which specific transcription factors (activators) bind to increase the rate of transcription. ...
Control of Gene Expression
... • RNA polymerase II then binds to the promoter to begin transcription at the start site (+1). • Enhancers are DNA sequences to which specific transcription factors (activators) bind to increase the rate of transcription. ...
... • RNA polymerase II then binds to the promoter to begin transcription at the start site (+1). • Enhancers are DNA sequences to which specific transcription factors (activators) bind to increase the rate of transcription. ...
DNA-binding motifs
... • RNA polymerase II then binds to the promoter to begin transcription at the start site (+1). • Enhancers are DNA sequences to which specific transcription factors (activators) bind to increase the rate of transcription. ...
... • RNA polymerase II then binds to the promoter to begin transcription at the start site (+1). • Enhancers are DNA sequences to which specific transcription factors (activators) bind to increase the rate of transcription. ...
Regulation of Gene Expression
... coactivators, corepressors, chromatin and histone modifiers Common structural classes of gene regulatory proteins ...
... coactivators, corepressors, chromatin and histone modifiers Common structural classes of gene regulatory proteins ...
17. Gene regulation
... transcriptional control control determines whether or not transcription is initiated requires promoter of gene transcription factors bind to promoter and recruit RNA polymerase to initiate transcription post-transcriptional regulation of gene expression control after pre-mRNA synthesis can ...
... transcriptional control control determines whether or not transcription is initiated requires promoter of gene transcription factors bind to promoter and recruit RNA polymerase to initiate transcription post-transcriptional regulation of gene expression control after pre-mRNA synthesis can ...
Controlling Gene Expression
... When lactose binds to LacI protein, it changes and the new complex cannot bind to the operator of the lac operon. This results in RNA polymerase being able to bind to the DNA and start protein synthesis. ...
... When lactose binds to LacI protein, it changes and the new complex cannot bind to the operator of the lac operon. This results in RNA polymerase being able to bind to the DNA and start protein synthesis. ...
Lecture 40_GeneRegulationI_transcriptional_control_RoadMap
... 8. Polyadenylation: the 3’ end of eukaryotic transcripts is polyadenylated. This does not happen in prokaryotes. ...
... 8. Polyadenylation: the 3’ end of eukaryotic transcripts is polyadenylated. This does not happen in prokaryotes. ...
Subject Description Form
... DNA; the replicon, initiation, DNA polymerases, primase, leading strand and lagging strand; Okazaki fragments. Transcription: prokaryotic promoters and terminators; RNA polymerase and sigma factors. Eukaryotic promoter elements, promoter proximal elements, enhancers, general transcription factors, a ...
... DNA; the replicon, initiation, DNA polymerases, primase, leading strand and lagging strand; Okazaki fragments. Transcription: prokaryotic promoters and terminators; RNA polymerase and sigma factors. Eukaryotic promoter elements, promoter proximal elements, enhancers, general transcription factors, a ...
401Lecture5sp2013post
... What result would you expect if DNA exists in loops? Would you expect loops to be present at all stages of cell cycle? ...
... What result would you expect if DNA exists in loops? Would you expect loops to be present at all stages of cell cycle? ...
Regulation of Gene Expression
... Sometimes the environment can change almost instantly Eukaryotes have to respond as well, although typically not as drastically With multicellular organisms, different types of cells express different sets of genes Structural genes encode proteins involved in metabolic or biosynthetic pathways ...
... Sometimes the environment can change almost instantly Eukaryotes have to respond as well, although typically not as drastically With multicellular organisms, different types of cells express different sets of genes Structural genes encode proteins involved in metabolic or biosynthetic pathways ...
Modification of Genes and Proteins - sharonap-cellrepro-p3
... reversal is also used to reverse methylation ...
... reversal is also used to reverse methylation ...