* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Slide
Genetic code wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Protein structure prediction wikipedia , lookup
Homology modeling wikipedia , lookup
Community fingerprinting wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Point mutation wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
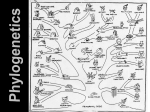
Molecular Phylogeny Biology 224 Instructor: Tom Peavy Nov 3, 8 & 10 <Images adapted from Bioinformatics and Functional Genomics by Jonathan Pevsner> Introduction Charles Darwin’s theory of evolution. --struggle for existence induces a natural selection. --Offspring are dissimilar from their parents (that is, variability exists), and individuals that are more fit for a given environment are selected for. --over long periods of time, species evolve. --Groups of organisms change over time so that descendants differ structurally and functionally from their ancestors. The basic processes of evolution are [1] mutation, [2] genetic recombination [3] chromosomal organization (and its variation); [4] natural selection [5] reproductive isolation, which constrains the effects of selection on populations At the molecular level, evolution is a process of mutation with selection. Molecular evolution is the study of changes in genes and proteins throughout different branches of the tree of life. Phylogeny is the inference of evolutionary relationships. Traditionally, phylogeny relied on the comparison of morphological features between organisms. Today, molecular sequence data are also used for phylogenetic analyses. Goals of molecular phylogeny Phylogeny can answer questions such as: • How many genes are related to my favorite gene? (gene tree) • Are humans more closely related to chimps or gorillas? (species tree) • How related are whales, dolphins & porpoises to cows? • Where and when did HIV originate? • What is the history of life on earth? The Structure of Phylogenetic Trees Molecular phylogeny uses trees to depict evolutionary relationships among organisms. These trees are based upon DNA and protein sequence data. 2 A 1 I 2 1 1 G B H 2 1 6 1 2 C 2 D B C 2 1 E A 2 F D 6 one unit E time Tree nomenclature Branches are unscaled... 2 Branches are scaled... A 1 I 2 1 1 G B H 2 1 6 1 2 C 2 D B C 2 1 E A 2 F D 6 one unit E time …OTUs are neatly aligned, and nodes reflect time …branch lengths are proportional to number of amino acid changes Tree nomenclature operational taxonomic unit (OTU) such as a protein sequence taxon 2 A 1 I 2 1 1 G B H 2 1 6 1 2 C 2 D B C 2 1 E A 2 F D 6 one unit E time Tree nomenclature Node (intersection or terminating point of two or more branches) branch 2 A A 2 (edge) F 1 I 2 1 1 G B H 2 1 6 1 2 C 2 E C 2 1 D B D 6 one unit E time Tree nomenclature bifurcating internal node multifurcating internal node 2 A 1 I 2 1 1 G B H 2 1 6 A 2 F B 2 C 2 2 1 D E C D 6 one unit E time Tree nomenclature: clades Clade ABF (monophyletic group) 2 F 1 I 2 A 1 B G H 2 1 6 C D E time Tree nomenclature Clade ABF/CDH/G 2 A F 1 I 2 1 G B H 2 1 6 C D E time Tree roots The root of a phylogenetic tree represents the common ancestor of the sequences. Some trees are unrooted, and thus do not specify the common ancestor. A tree can be rooted using an outgroup (that is, a taxon known to be distantly related from all other OTUs). Tree nomenclature: roots past 9 1 7 5 8 6 2 present 1 7 3 4 2 5 Rooted tree (specifies evolutionary path) 8 6 3 Unrooted tree 4 Tree nomenclature: outgroup rooting past root 9 10 7 8 7 6 2 present 9 8 3 4 1 Rooted tree 2 5 1 3 4 5 6 Outgroup (used to place the root) Numbers of trees Number of OTUs 2 3 4 5 10 Number of rooted trees 1 3 15 105 34,459,425 Number of unrooted trees 1 1 3 15 105 Species trees versus gene/protein trees Molecular evolutionary studies can be complicated by the fact that both species and genes evolve. speciation usually occurs when a species becomes reproductively isolated. In a species tree, each internal node represents a speciation event. Genes (and proteins) may duplicate or otherwise evolve before or after any given speciation event. The topology of a gene (or protein) based tree may differ from the topology of a species tree. Species trees versus gene/protein trees past speciation event present species 1 species 2 Species trees versus gene/protein trees Gene duplication events speciation event OTUs species 1 species 2 Molecular Evolution Historical background: insulin By the 1950s, it became clear that amino acid substitutions occur nonrandomly e.g. most amino acid changes in the insulin A chain are restricted to a disulfide loop region. Such differences are called “neutral” changes rate of nucleotide (and of amino acid) substitution is about sixto ten-fold higher in the C peptide, relative to the A and B chains. Mature insulin consists of an A chain and B chain heterodimer connected by disulphide bridges The signal peptide and C peptide are cleaved, and their sequences display fewer functional constraints. 0.1 x 10-9 1 x 10-9 0.1 x 10-9 Number of nucleotide substitutions/site/year for insulin Historical background: insulin Surprisingly, insulin from the guinea pig (and from the related coypu) evolve seven times faster than insulin from other species. Why? The answer is that guinea pig and coypu insulin do not bind two zinc ions, while insulin molecules from most other species do. There was a relaxation on the structural constraints of these molecules, and so the genes diverged rapidly. Molecular clock hypothesis In the 1960s, sequence data were accumulated for small, abundant proteins such as globins, cytochromes c, and fibrinopeptides. Some proteins appeared to evolve slowly, while others evolved rapidly. Linus Pauling, Emanuel Margoliash and others proposed the hypothesis of a molecular clock: For every given protein, the rate of molecular evolution is approximately constant in all evolutionary lineages Molecular clock hypothesis Richard Dickerson (1971) plotted data from three protein families: cytochrome c, hemoglobin, and fibrinopeptides. The x-axis shows the divergence times of the species, estimated from paleontological data. The y-axis shows m, the corrected number of amino acid changes per 100 residues. n is the observed number of amino acid changes per 100 residues, and it is corrected to m to account for changes that occur but are not observed. N = 1 – e-(m/100) 100 Hidden mutation due to multiple substitutions corrected amino acid changes per 100 residues (m) Dickerson (1971) Millions of years since divergence • For each protein, the data lie on a straight line. Thus, the rate of amino acid substitution has remained constant for each protein. • The average rate of change differs for each protein. The time for a 1% change to occur between two lines of evolution is 20 MY (cytochrome c), 5.8 MY (hemoglobin), and 1.1 MY (fibrinopeptides). • The observed variations in rate of change reflect functional constraints imposed by natural selection. Molecular clock for proteins: rate of substitutions per aa site per 109 years Fibrinopeptides Kappa casein Lactalbumin Serum albumin Lysozyme Trypsin Insulin Cytochrome c Histone H2B Ubiquitin Histone H4 9.0 3.3 2.7 1.9 0.98 0.59 0.44 0.22 0.09 0.010 0.010 Molecular clock hypothesis: implications If protein sequences evolve at constant rates, they can be used to estimate the times that sequences diverged. This is analogous to dating geological specimens by radioactive decay. N = total number of substitutions L = number of nucleotide sites compared between two sequences K= N L = number of substitutions per nucleotide site See Graur and Li (2000), p. 140 Rate of nucleotide substitution r and time of divergence T r = rate of substitution = 0.56 x 10-9 per site per year for hemoglobin alpha K = 0.093 = number of substitutions per nucleotide site (rat versus human) r = K / 2T T = .093 / (2)(0.56 x 10-9) = 80 million years See Graur and Li (2000), p. 140 Neutral theory of evolution Kimura’s (1968) neutral theory of molecular evolution: --the vast majority of DNA changes are not selected for in a Darwinian sense. --The main cause of evolutionary change is random drift of mutant alleles that are selectively neutral (or nearly neutral). --Positive Darwinian selection does occur, but limited role. e.g. the divergent C peptide of insulin changes according to the neutral mutation rate. “fast-clock” organisms • These organisms with long branches are called “fastclock” • They really acumulate substitutions faster than the rest of organisms (their rate of substitution is higher) • Some authors have proposed various hypothesis to try to explain this phenomenon: – higher metabolic rate, short generation time, differences in the number of replications of DNA in the germinal line, deficiences in DNA repair, mutagens, Solutions? • Use methods less sensitive to this type of inconsistency (ML?) • If it is possible, eliminate long branches: – eliminate the “fast-clock” organism – substitute by another of the same group that is not “fast-clock” – increase the number of organisms of that group Solutions? • We first need to know if we really have a “fast-clock” organism • Relative Rate Test – Sarich and Wilson, 1973 for proteins – Wu and Li (1985) and Li and Tanimura (1987) extended it to nucleotides Relative Rate Test • Uses 3 species A, B and one “outgroup” C • Tests if A and B have the same rate of substitution since their split: O dAO = dBO dAC = dBC d = dAC - dBC = 0 A B C Relative Rate Test • This method is time independent • We have to be sure about the phylogeny O A B C Relative Rate Test O • Our null hypothesis is: d = dAC - dBC = 0 A B • It is assumed that the number of nucleotide substitutions follows a Poisson, • then we can use the standarized normal distribution to test if the number of substituions in the 2 lineages is the same C Relative Rate Test • d = dAC - dBC = 0 • d ± Var(d) • Var(d) = Var(dAC) + Var(dBC) - 2 Cov (dAC,dBC) – |d| > 1.96 Var(d) = significant at the 5% level – |d| > 2.96 Var(d) = significant at the 1% level x – 1.96 0 x + 1.96 How to Construct Phylogenetic Trees Four stages of phylogenetic analysis Molecular phylogenetic analysis may be described in four stages: [1] Selection of sequences for analysis [2] Multiple sequence alignment [3] Tree building [4] Tree evaluation Stage 1: Use of DNA, RNA, or protein -Protein alignments are more informative as to structure function relationships -Although DNA may be preferable for the phylogenetic analysis since the protein-coding portion of DNA has synonymous and nonsynonymous substitutions -RNA is useful for the other non-protein coding genes (e.g. tRNAs) if looking at structure-function relationships But often use the gene instead for phylogeny (e.g. genes For rRNA) Stage 1: Use of DNA, RNA, or protein For phylogeny, protein sequences are also often used. --Proteins have 20 states (amino acids) instead of only four for DNA, so there is a stronger phylogenetic signal. Nucleotides are unordered characters: any one nucleotide can change to any other in one step. An ordered character must pass through one or more intermediate states before reaching the final state. Amino acid sequences are partially ordered character states: there is a variable number of states between the starting value and the final value. Synonymous vs Nonsynonymous rates If the synonymous substitution rate (dS) is greater than the nonsynonymous substitution rate (dN), the DNA sequence is under negative (purifying) selection. This limits change in the sequence (e.g. insulin A chain). If dS < dN, positive selection occurs. For example, a duplicated gene may evolve rapidly to assume new functions. DNA can be more informative also due to: --Rates of transitions and transversions can be measured. --Noncoding regions (such as 5’ and 3’ untranslated regions) may be analyzed using molecular phylogeny. --Pseudogenes (nonfunctional genes) are studied by molecular phylogeny -- Additional mutational events can be inferred by analysis of ancestral sequences. These changes include parallel substitutions, convergent substitutions, and back substitutions. -- in order to predict ancestral sequence, other distantly related sequences are analyzed Stage 2: Multiple sequence alignment The fundamental basis of a phylogenetic tree is a multiple sequence alignment. (If there is a misalignment, or if a nonhomologous sequence is included in the alignment, it will still be possible to generate a tree.) Consider the following (see Fig. 3.2) Alignment of 13 orthologous retinol-binding proteins Some positions of the multiple sequence alignment are invariant (arrow 2). Some positions distinguish fish RBP from all other RBPs (arrow 3). Stage 2: Multiple sequence alignment [1] Confirm that all sequences are homologous [2] Adjust gap creation and extension penalties as needed to optimize the alignment [3] Restrict phylogenetic analysis to regions of the multiple sequence alignment for which data are available for all taxa (delete columns having incomplete data). [4] Many experts recommend that you delete any column of an alignment that contains gaps (even if the gap occurs in only one taxon) Stage 3: Tree-building methods Discuss two tree-building methods: distance-based versus character-based. Distance-based methods involve a distance metric, such as the number of amino acid changes between the sequences, or a distance score. Examples of distance-based algorithms are UPGMA and neighbor-joining. Character-based methods include maximum parsimony and maximum likelihood. Parsimony analysis involves the search for the tree with the fewest amino acid (or nucleotide) changes that account for the observed differences between taxa. common carp zebrafish Fish RBP orthologs rainbow trout teleost African clawed frog chicken human mouse rat horse pig cow rabbit 10 changes Other vertebrate RBP orthologs Distance-based tree Calculate the pairwise alignments; if two sequences are related, put them next to each other on the tree Character-based tree: identify positions that best describe how characters (amino acids) are derived from common ancestors Stage 3: Tree-building methods Regardless of whether you use distance- or character-based methods for building a tree, the starting point is a multiple sequence alignment. ReadSeq is a convenient web-based program that translates multiple sequence alignments into formats compatible with most commonly used phylogeny programs such as PAUP and PHYLIP. Mega has its own text converter. Stage 3: Tree-building methods: distance The simplest approach to measuring distances between sequences is to align pairs of sequences, and then to count the number of differences. The degree of divergence is called the Hamming distance. For an alignment of length N with n sites at which there are differences, the degree of divergence D is: D=n/N But observed differences do not equal genetic distance! Genetic distance involves mutations that are not observed directly Stage 3: Tree-building methods: distance Jukes and Cantor (1969) proposed a corrective formula: D = (- 3 ) ln (1 – 4 p) 4 3 This model describes the probability that one nucleotide will change into another. It assumes that each residue is equally likely to change into any other (i.e. the rate of transversions equals the rate of transitions). In practice, the transition is typically greater than the transversion rate. Models of nucleotide substitution A transition G transversion transversion C T transition Jukes and Cantor one-parameter model of nucleotide substitution a A G b b b b T a C Stage 3: Tree-building methods: distance Jukes and Cantor (1969) proposed a corrective formula: D = (- 3 ) ln (1 – 4 p) 4 3 Consider an alignment where 3/60 aligned residues differ. The normalized Hamming distance is 3/60 = 0.05. The Jukes-Cantor correction is D = (- 3 ) ln (1 – 4 0.05) = 0.052 4 3 When 30/60 aligned residues differ, the Jukes-Cantor correction is more substantial: D = (- 3 ) ln (1 – 4 0.5) = 0.82 4 3 Many software packages are available for making phylogenetic trees. http://evolution.genetics.washington.edu/phylip/software.html This site lists 200 phylogeny packages. Perhaps the bestknown programs are PAUP (David Swofford et al.), PHYLIP (Joe Felsenstein) and MEGA (Kumar et al.) UPGMA (distance-based tree) Tree-building methods: UPGMA UPGMA is unweighted pair group method using arithmetic mean 1 2 3 4 5 Tree-building methods: UPGMA Cluster the smallest pairwise alignments And repeat until all clusters are drawn 1 2 3 4 5 Step 1 1 2 6 3 1 4 2 5 1 2 6 Step 2 1 3 4 5 7 2 4 5 Step 3 1 8 2 7 6 3 4 1 5 2 4 3 5 9 1 2 8 Step 4 7 3 6 4 5 1 2 4 5 3 Distance-based methods: UPGMA trees UPGMA is a simple approach for making trees. • An UPGMA tree is always rooted. • An assumption of the algorithm is that the molecular clock is constant for sequences in the tree. If there are unequal substitution rates, the tree may be wrong. • While UPGMA is simple, it is less accurate than the neighbor-joining approach (described next). Making trees using neighbor-joining The neighbor-joining method of Saitou and Nei (1987) Is especially useful for making a tree having a large number of taxa. Begin by placing all the taxa in a star-like structure. Tree-building methods: Neighbor joining Next, identify neighbors (e.g. 1 and 2) that are most closely related. Connect these neighbors to other OTUs via an internal branch, XY. At each successive stage, minimize the sum of the branch lengths. dXY = 1/2(d1Y + d2Y – d12) Example of a neighbor-joining tree: phylogenetic analysis of 13 RBPs Tree-building methods: character based Rather than pairwise distances between proteins, evaluate the aligned columns of amino acid residues (characters). Tree-building methods based on characters include maximum parsimony and maximum likelihood. Making trees using character-based methods The main idea of character-based methods is to find the tree with the shortest branch lengths possible. Thus we seek the most parsimonious (“simple”) tree. • Identify informative sites. For example, constant characters are not parsimony-informative. • Construct trees, counting the number of changes required to create each tree. For about 12 taxa or fewer, evaluate all possible trees exhaustively; for >12 taxa perform a heuristic search. • Select the shortest tree (or trees). As an example of tree-building using maximum parsimony, consider these four taxa: AAG AAA GGA AGA How might they have evolved from a common ancestor such as AAA? Tree-building methods: Maximum parsimony AAA 1 AAA AAG AAA 1 1 AGA GGA AGA Cost = 3 AAA 1 AAA 1 AAG AGA AAA AAA 2 AAA GGA Cost = 4 1 AAA 2 AAG GGA AAA 1 AAA AGA Cost = 4 In maximum parsimony, choose the tree(s) with the lowest cost (shortest branch lengths). In PAUP’s implementation of maximum parsimony, many arrangements are tried and the best trees (lowest branch lengths) are saved Phylogram (values are proportional to branch lengths) Rectangular phylogram (values are proportional to branch lengths) Cladogram (values are not proportional to branch lengths) Rectangular cladogram (values are not proportional to branch lengths) These four trees display the same data in different formats. 37 A1AG HUMA 40 A1AG RABI 25 21 24 A1AH MOUS A1AG RAT 40 23 44 APHR CRIC 36 OBP RAT 2 61 22 PBAS RAT 15 MUP1 MOUS 33 27 MUPM MOUS 25 MUP RAT 3 66 HUMA 40 CO8G AMBP HUMA 22 40 34 FAB1 MANS 24 FAB2 MANS 25 FABL CHIC 32 33 24 38 16 21 ILBP PIG ILBP18 RAT 14 19FABA HUMA 64 17 15 36MYP2 BOVI 27 FABE HUMA 15 20 20 FABH BOVI 34 21 26 FABL GINC 43 FABP ECHG 25 37 18 FABP SCHM 33 21 17 32 RET3 BOVI 27 23 25 RET1 HUMA 28 31 RET2 MOUS 54 FABI HUMA 38 FABL HUMA 52 26 71 AMBP PLEP RANP 38 58 OLFA LALP MACE 31 30 31 29 VEG1 RAT 24 VEGP HUMA 53 24 ERBP RAT 48 ESP4 LACV 64 57 QSP 46 CHICK 25 41 LIPO BUFM 35 PGHD HUMA 28 49 26 32 NGAL HUMA NGAL MOUS 21 LACA CANF 11 21 21 LACB BOVI 28 16 LACB PIG 34 37 30 2115 LACA EQUA LACB EQUA 43 40 PAEP HUMA 56 LACB 54 MACG APD HUMAN 28 53 BBP PIEBR 29 42 38 33 ICYA MANS 35 PURP 59 24 RET1CHIC ONCM 23 25 46 18 33 RETB BOVI RETB XENL 39 45 HOMG 49 CRA2 CRC1 HOMG 41 OBP BOVIN 50 changes odorant-binding protein (rat) lactoglobulin retinol-binding protein odorant-binding protein (bovine) Tree artifacts: long branch attraction For some phylogenetic trees, particularly those based on maximum parsimony, the artifact of long-branch attraction may occur. Branch lengths often depict the number of substitutions that occur between two taxa. Parsimony assumes all taxa evolve at the same rate, and all characters contribute the same amount of information. Rapidly evolving taxa may be placed on the same branch, not because they are related, but because they both have many substitutions. Long branch attraction (LBA) • When the length of the branches or the substitution rates are extremely unequal, there is a violation of the assumptions made by inference methods – termed the Felsenstein Zone Long branch chain attraction can confound phylogenetic analyses Making trees using maximum likelihood Maximum likelihood is an alternative to maximum parsimony. It is computationally intensive. A likelihood is calculated for the probability of each residue in An alignment, based upon some model of the substitution process. Stage 4: Evaluating trees The main criteria by which the accuracy of a phylogentic tree is assessed are consistency, efficiency, and robustness. Evaluation of accuracy can refer to an approach (e.g. UPGMA) or to a particular tree. Stage 4: Evaluating trees: bootstrapping Bootstrapping is a commonly used approach to measuring the robustness of a tree topology. Given a branching order, how consistently does an algorithm find that branching order in a randomly permuted version of the original data set? Stage 4: Evaluating trees: bootstrapping To bootstrap, make an artificial dataset obtained by randomly sampling columns from your multiple sequence alignment. Make the dataset the same size as the original. Do 100 (to 1,000) bootstrap replicates. Observe the percent of cases in which the assignment of clades in the original tree is supported by the bootstrap replicates. >70% is considered significant. In 61% of the bootstrap resamplings, ssrbp and btrbp (pig and cow RBP) formed a distinct clade. In 39% of the cases, another protein joined the clade (e.g. ecrbp), or one of these two sequences joined another clade.