* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Atomic models of complexes - Cryo
Survey
Document related concepts
Transcript
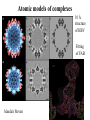
Data Mining Text mining, e.g.: • Structures at specific resolution • Structures by negative staining • Structures by vitrification Mining structural data, e.g.: • Segmentation • 3D shape recognition • Secondary structure recognition • Fold modules recognition Goals • Atomic scale model of a cell • Atomic models of interacting complexes • Conformational dynamics Difficulties • Range of size • Range of resolution • Range of complexity Towards the atomic model of a cell Wolfgang Baumeister Segmentation Actin mesh & aldolase Niels Volkmann Volkmann, 2002, JSB 138:123 Atomic models of complexes 15 Å 10 Å Helixhunter Situs 5Å b sheets Helix orientation Wah Chiu If no structure available: SCOP: 1.65 released December 2003. CATH: 2.5.1: Released January 2004 Atomic models of complexes 10 Å structure of HBV Fitting of FAB Alasdair Steven Helix orientation: the fold The directionality of the helices was determined by collecting the best-ftting orientations resulting from a search through the 3D experimental map for a large number of a-helical fragments. QuickTime™ and a Video decompressor are needed to see this picture. 3D cryo EM 4.5 Å resolution Mitsuoka et al. J. Struct. Biol. (1999) De Groot et al., J. Mol. Biol (2000) How to describe protein motion without sequence and atomic coordinates The 3D continuous object is decomposed into a set of Voronoi cells that approximate its shape and density distribution. The anisotropic elastic network theory allows the thermal fluctuations to be predicted. Ming et al. PNAS (2002) 99: 8620-25 Tama et al. J. Mol. Biol. (2002) 321: 297–305 Scheuring et al. E. Biophys J, 2001