* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download NMR Spectroscopy
Survey
Document related concepts
Transcript
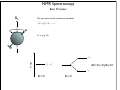
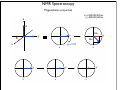
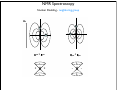
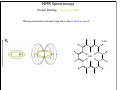
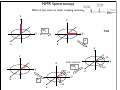
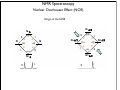
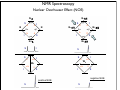
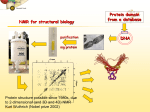
NMR Spectroscopy Applications Drug design MRI Food quality Metabonomics Structural biology NMR Spectroscopy Basic Principles N.M.R. = Nuclear Magnetic Resonance Spectroscopic technique, thus relies on the interaction between material and electromagnetic radiation The nuclei of all atoms possess a nuclear quantum number, I. (I0, always multiples of .) Only nuclei with spin number (I) >0 can absorb/emit electromagnetic radiation. Even atomic mass & number: I = 0 (12C, 16O) Even atomic mass & odd number: I = whole integer (14N, 2H, 10B) Odd atomic mass: I = half integer (1H, 13C, 15N, 31P) The spinning nuclei possess angular momentum, P, and charge, and so an associated magnetic moment, . = x P Where is the gyromagnetic ratio NMR Spectroscopy Basic Principles B0 The spin states of the nucleus are quantified: I, (I - 1), (I - 2), … , -I I= (e.g. 1H) Energy – – E=h=hB0/2 B0=0 B0>0 NMR Spectroscopy Basic Principles In the ground state all nuclear spins are disordered, and there is no energy difference between them. They are degenerate. Bo Since they have a magnetic moment, when we apply a strong external magnetic field (Bo), they orient either against or with it: There is always a small excess of nuclei (population excess) aligned with the field than pointing against it. NMR Spectroscopy Basic Principles Bo>0 Bo=0 E=h0=hB0/2 0 is the Larmor Frequency 0=B0, angular velocity z B0 y x NMR Spectroscopy Basic Principles Each level has a different population (N), and the difference between the two is related to the energy difference by the Boltzmman distribution: N/N = eE/kT E for 1H at 400 MHz (B0 = 9.5 T) is 3.8 x 10-5 Kcal/mol N/N =1.000064 The surplus population is small (especially when compared to UV or IR). That renders NMR a rather insensitive technique! NMR Spectroscopy The electromagnetic spectrum -rays 600 1022 1020 Mossbauer 500 400 X-rays visible 1014 infrared electronic 100 rotational 108 NMR 106 /Hz 4 200 aldehydic aromatic 6 31P vibrational 1010 radiofrequency 8 2 0 1012 microwave 19F 300 1018 ultraviolet 1016 10 1H 13C /MHz olefinic acetylenic aliphatic /ppm NMR Spectroscopy The Vector Model M0 y x y x y y x x NMR Spectroscopy NMR excitation z B1 = C * cos (ot) Mo y B1 x Bo i Transmitter coil (x) B1 is an oscillating magnetic field y y -0 +0 x x NMR Spectroscopy Laboratory vs. Rotating frame z z’ +0 M0 M0 y +0 x y’ B1 -0 Laboratory frame x’ -20 Rotating frame NMR Spectroscopy Effect on an rf pulse z z M0 =1tp (degrees) =90° B1 x y y B1 x =180° NMR Spectroscopy Magnetization properties 1H=400,000,000 Hz A=400,000,005 Hz z A y y B1 MAy MAx =5 Hz x x y x x y x y x y NMR Spectroscopy Magnetization properties 1H=400,000,000 Hz A=400,000,005 Hz z A B1 MAy y y y MAx =5 Hz x x x MAy=MA cost detector MAx=MA sint IM IM MA MA t t t NMR Spectroscopy The Fourier Transform FT time domain IM frequency domain =1/t t FT t time frequency NMR Spectroscopy The Fourier Transform IM Signal Induction Decay (FID) time NMR Spectroscopy The Fourier Transform FT NMR Spectroscopy Continuous wave vs. pulsed NMR o or Bo o or Bo time NMR Spectroscopy Continuous wave vs. pulsed NMR • For cos(t) FT absorptive lines • For sin(t) FT despersive lines NMR Spectroscopy Continuous wave vs. pulsed NMR A monochromatic radiofrequency pulse is a combination of a wave (cosine) of frequency 0 and a step function * = tp Since f=1/t, a pulse of 10 s duration excites a frequency bandwidth of 105 Hz! FT o NMR Spectroscopy Continuous wave vs. pulsed NMR E t ~ h or t ~1 NMR Spectroscopy Single-channel signal detection z y x FT + + 0 - NMR Spectroscopy Quadrature detection z y x My Mx My Mx + NMR Spectroscopy Quadrature detection cos y sin x + 0 + + - 0 Hz Hz NMR Spectroscopy The Chemical Shift The NMR frequency of a nucleus in a molecule is mainly determined by its gyromagnetic ratio and the strength of the magnetic field B The exact value of depends, however, on the position of the nucleus in the molecule or more precisely on the local electron distribution this effect is called the chemical shift NMR Spectroscopy The Chemical Shift E=h=hB/2 Nuclei, however, in molecules are never isolated from other particles that are charged and are in motion (electrons!). Thus, the field actually felt by a nucleus is slightly different from that of the applied external magnetic field!! NMR Spectroscopy The Chemical Shift E=h=hBe/2 Beff, is given by B0-B= B0-B0=B0(1-) = and is the chemical shift = (-ref) ref B0(1-) 2 106 106 (ref-) NMR Spectroscopy The Chemical Shift methyl protons amide protons protons methylene protons aromatic ring 750 MHz 1H spectrum of a small protein shielding frequency magnetic field NMR Spectroscopy The Chemical Shift NMR Spectroscopy The Chemical Shift NMR Spectroscopy Nuclear Shielding =dia + para + nb + rc + ef + solv diamagnetic contribution paramagnetic contribution neighbor anisotropy effect ring-current effect electric field effect solvent effect NMR Spectroscopy Nuclear Shielding - diamagnetic contribution The external field B0 causes the electrons to circulate within their orbitals B0 hB0 hB0(1-) B’ The higher is the electron density close to the nucleus, the larger the protection is! NMR Spectroscopy Nuclear Shielding - diamagnetic contribution Depends on the electronegativity CH3X NMR Spectroscopy Nuclear Shielding - paramagnetic contribution The external field B0 mixes the wavefunction of the ground state with that of the excited state The induced current generates a magnetic field that enhances the external field and deshields the nucleus LUMO B0 HOMO p = 1 1 R3 NMR Spectroscopy Chemical shift range 1H; ~10 ppm 13C; ~200 ppm 19F; ~300 ppm 31P; ~500 ppm Local diamagnetic and paramagnetic currents make only modest contributions to 1H shielding! NMR Spectroscopy Chemical Shift Anisotropy Nuclear shielding, , is a tensor. The distribution of the electrons about the nucleus is non-sperical- thus, the magnitude of the shielding depends on the relative orientation of the nucleus with respect to the static field. In isotropic cases: = (11 + 22 + 33) In static cases, e.g. solid state NMR Spectroscopy Nuclear Shielding - neighboring group B0 B A B μpar > μper A μpar < μper + + + - - + NMR Spectroscopy Nuclear Shielding - neighboring group + - C C - μpar > μper C + μpar < μper + + C - C 2H 4 7 6 C 2H 2 5 4 3 C 2H 6 2 1 0 ppm NMR Spectroscopy Nuclear Shielding - ring-current effect More pronounced in aromatic rings due to the electron clouds Bo 9.28 e- -2.99 NMR Spectroscopy Nuclear Shielding - hydrogen bonding Hydrogen bonding causes deshielding due to electron density decrease at the proton site [EtOH] in CCl4 CH3 CH2 OH 1M 0.1M 0.01M 0.001M 6 4 2 0 ppm NMR Spectroscopy Spin-spin (scalar) coupling HF (1H-19F) H F JHF JHF NMR Spectroscopy Spin-spin (scalar) coupling HF (1H-19F) H F H H F Bo 19F 1H H F H H F 19F 1H Nuclear moment Magnetic polarization of the electron E=h JAX mA mX where m is the magnetic quantum number JAX is the spin-spin coupling constant NMR Spectroscopy Spin-spin (scalar) coupling AX2 AMX AX3 NMR Spectroscopy Spin-spin (scalar) coupling Strong coupling – <10|J| NMR Spectroscopy Spin-spin (scalar) coupling The principal source of scalar coupling is an indirect interaction mediated by electrons involved in chemical bonding The magnitude of interaction is proportional to the probability of finding the electron at the nucleus (R=0) Magnitude in Hz- independent of the external magnetic field H3C – CH3 H2C – CH2 HC CH 125 Hz 160 Hz 250 Hz NMR Spectroscopy Spin-spin (scalar) coupling Three-bond coupling most useful since it carries information on dihedral angles Empirical relationship: the Karplus relation 3J = A + B cos + C cos2 NMR Spectroscopy Chemical shifts on the rotating frame 500 MHz z 3 y 2 0 x t z y x =500 Hz NMR Spectroscopy Spin couplings on the rotating frame J z y 0 x t z =-J/2 Hz y x =+J/2 Hz NMR Spectroscopy The basic spin-echo pulse sequence 90° applied along x axis 180° applied along y axis x delay y acquisition NMR Spectroscopy 90° applied along x axis Effect of spin echo on chemical shift evolution 180° applied along y axis x y acquisition delay z z y 90x y x x z z y z y x 180y A x y x A NMR Spectroscopy 90° applied along x axis Effect of spin echo on scalar coupling evolution 180° applied along y axis x y acquisition delay z z y 90x 1H-X y x x z z -J/2 y z (only 180y y y -J/2 x +J/2 x +J/2 x 1H) NMR Spectroscopy 90° applied along x axis Effect of spin echo on scalar coupling evolution 180° applied along y axis x y acquisition delay z z y 90x 1H-X y x x z z -J/2 z 180y y y +J/2 x +J/2 x -J/2 x y (both 1H and X) NMR Spectroscopy Water suppression by the Jump and Return method z z y 90x y x x z z y z y A 90-x x y x x NMR Spectroscopy Water suppression NMR Spectroscopy Spin decoupling decouple 1H 13C H C H H C H C H H C NMR Spectroscopy The J-modulated spin echo decouple 1H y x 13C NMR Spectroscopy The J-modulated spin echo decouple 1H y x 13C NMR Spectroscopy The J-modulated spin echo If =180J degrees C: CH: CH2: CH3: I=1 I cos I cos2 I cos3 decouple 1H y x 13C NMR Spectroscopy The J-modulated spin echo decouple 1H y x 13C =1/J 13C (ppm) NMR Spectroscopy Sensitivity enhancement NMR has poor sensitivity compared to other analytical techniques The intrinsic sensitivity depends upon the gyromagnetic ratio, A greater contributes to: a high resonant frequency- large transition energy difference- greater Boltzmann population difference high magnetic moment and hence a stronger signal high rate of precession which induces a greater signal in the detection coil So, the strength of NMR signal is proportional to 3 Noise increases a square-root of observed frequency } S/N 5/2 NMR Spectroscopy Sensitivity enhancement by polarization transfer Signal sensitivity enhancement by transferring the greater population differences of high spins onto their spin-coupled low- partners. C2 1H-13C H2 H1 C1 spin pair NMR Spectroscopy Sensitivity enhancement by polarization transfer Signal sensitivity enhancement by transferring the greater population differences of high spins onto their spin-coupled low- partners. C 2C C2 H2 +C 2 2 H1 C2 H2 +C 2 C C1 C 2+2C 2 H1 C1 (inverted) 2C C 2+2C +C +C C1 H1 H1 H2 C1 C2 H2 C2 -3:5 NMR Spectroscopy Sensitivity enhancement by polarization transfer Signal sensitivity enhancement by transferring the greater population differences of high spins onto their spin-coupled low- partners. coupled decoupled INEPT refocused INEPT refocused, decoupled INEPT NMR Spectroscopy Relaxation When perturbed, the nuclear spins need to relax to return to their equilibrium distribution E.g. when the sample is put into a magnet, the Boltzmann distribution of spins among the energy levels changes due to a change in the energy of the various levels E.g. after applying electromagnetic radiation, which induces transitions between energy levels, the system returns to its equilibrium This process is called relaxation NMR Spectroscopy Longitudinal Relaxation: Establishing Equilibrium z y x z z y y x x z z x x NMR Spectroscopy Longitudinal Relaxation: Establishing Equilibrium Recovery of the z-magnetization follows exponential behavior dMz dt = (M0-Mz) T1 Mz=M0 (1-2e-t/T1) where T1 is the longitudinal relaxation time NMR Spectroscopy Longitudinal Relaxation: Measurement x x z z z 180x z y x 90x y x y y x x z z 90x y x y x NMR Spectroscopy Longitudinal Relaxation: Measurement x x NMR Spectroscopy Longitudinal Relaxation: Exponential growth Mz=M0 (1-2e-t/T1) By the end of 5T1 sec, the magnetization has recovered by 99.33% NMR Spectroscopy Longitudinal Relaxation: optimizing sensitivity NMR Spectroscopy Longitudinal Relaxation: optimizing sensitivity NMR Spectroscopy Longitudinal Relaxation: optimizing sensitivity optimum pulse repetition time when using 90º Quantitative measurements and integration NMR Spectroscopy Transverse Relaxation: magnetization loss in the x-y plane y x - - - y y y + + + x x time x NMR Spectroscopy Transverse Relaxation: magnetization loss in the x-y plane = 1 T2* NMR Spectroscopy Transverse Relaxation: Measurement x z z z 90x - y y y x + x z 180y + x y z y y x x NMR Spectroscopy Transverse Relaxation: Measurement NMR Spectroscopy T1 vs T2 Relaxation T1 T2 For small molecules, T1 T2 For large molecules, T1 >> T2 Longitudinal relaxation causes loss of energy from the spins (enthalpic) Transverse relaxation occurs by mutual swapping of energy between spins (entropic) NMR Spectroscopy Relaxation mechanisms Nuclear spin relaxation is not a spontaneous process; it requires stimulation by suitable fluctuating fields to induce the necessary spin transitions Two main mechanisms Dipole-dipole Chemical shift anisotropy NMR Spectroscopy Relaxation mechanisms Longitudinal relaxation requires a time-dependent magnetic field fluctuating at the Larmor frequency The time-dependence originates in the motions of the molecule (vibration, rotation, diffusion etc) Molecules in solution “tumble”. This “tumbling” can be characterized by a rotational correlation time c c is the time needed for the rms deflection of the molecules to be ~ 1 radian (60°) NMR Spectroscopy Spectral density function Rotational diffusion in solution occurs at a range of frequencies 1/c ~ rms rotational frequency (radians s-1) The probability function of finding motions at a given angular frequency can be described by the spectral density function J() NMR Spectroscopy Spectral density function Frequency distribution of the fluctuating magnetic fields NMR Spectroscopy Spectral density function: Longitudinal relaxation Spins are relaxed by local fields fluctuating at the Larmor frequency 0 So, the relaxation rate (R1) will be proportional to the J(0) 1/T1= R1 = 2 <B2> J(0) Knowing the form of J() we can predict the dependence of the spin-lattice relaxation time (T1=1/ R1) on the correlation time c for a given NMR frequency 0 0c=1 ,J(0) = c= 1/0 and T1 is minimum (R1 maximum) 0c<<1 (small molecules), J(0) ~ 2c and T1 decreases (R1 increases) with increasing c 0c<<1 0c>>1 (e.g.by decreasing the temperature) 0c=1 0c>>1 (large molecules), J(0) ~ 2/02c and T1 increases (R1 decreases) with increasing c (e.g. by decreasing the temperature) NMR Spectroscopy Relaxation mechanisms: Dipole-dipole Nuclei with non-zero quantum numbers have magnetic dipoles They behave like small magnets and induce small magnetic fields that affect neighboring nuclei Magnetic field, B, generated by a magnetic dipole NMR Spectroscopy Relaxation mechanisms: Dipole-dipole Representation of the dipolar magnetic field B, generated by a magnetic dipole lines of force density plots Bμz Bμz is zero for =±54.7˚ (magic angle) Bμx NMR Spectroscopy Relaxation mechanisms: Dipole-dipole The z component of their dipole magnetic field will affect the field experienced by the other nucleus and cause splitting ± sign refers to the quantum number of A (±) X A Bμ Thus, the splitting in the spectrum of X is A KAX vary with the distance e.g. KCH is 9000 Hz at 1.5 Å and 30 Hz at 10 Å NMR Spectroscopy Relaxation mechanisms: Dipole-dipole Splitting of the AX spectrum depends on In a crystal with fixed distances and angles the dipolar splitting vary with the crystal orientation with respect to the external magnetic field NMR Spectroscopy Relaxation mechanisms: Dipole-dipole Molecules in liquids rotate, “tumble” rapidly with typical frequencies between 1012 to 108 Hz for small molecules and proteins, respectively. Those frequencies are much larger than typical dipolar couplings (105 Hz) The angular part of the dipolar splitting is averaged over all possible orientation to 0 Although they are not directly observed in solution, dipolar couplings play an important role in spin relaxation The local field experienced at one nucleus as a result of its neighbor will fluctuate as the molecule tumbles NMR Spectroscopy Relaxation mechanisms: Dipole-dipole R1 depend of the gyromagnetic ratio of the nuclei (e.g. H-H relaxation more efficient than C-H) NMR Spectroscopy Relaxation mechanisms: Chemical shift anisotropy The distribution of the electrons about the nucleus is non-sperical- thus, the magnitude of the shielding depends on the relative orientation of the nucleus with respect to the static field. As the molecule tumbles, it creates a fluctuating magnetic field NMR Spectroscopy Nuclear Overhauser Effect (NOE) NOE: change in intensity of one resonance when the spin transitions of another are perturbed from their equilibrium populations perturbation: saturation or inversion The two spins should “communicate” through dipole-dipole interaction NOE is observed for spin I when spin S is perturbed NMR Spectroscopy Nuclear Overhauser Effect (NOE) Origin of the NOE N-/2 N- S N I I S N 0 N/2 I I 0 S N+/2 N+ S S I S I N+/2 NMR Spectroscopy Nuclear Overhauser Effect (NOE) Six possible transitions in a two-spin system W1S W1I W2 W0 W1I W1S Only single transitions can by observed by NMR (W1) W0 and W2 are cross-relaxation pathways, responsible for the NOE NMR Spectroscopy Nuclear Overhauser Effect (NOE) N-/2 N- S N I N 0 N/2 I S S I I S S I S I S N+/2 S > > W2 I 0 W0 < 0 I S I < 0 S negative NOE positive NOE S N+/2 0 N+ 0 I I S I NMR Spectroscopy Nuclear Overhauser Effect (NOE) W1 tends to reduce the magnitude of the NOE Saturating for a period of time that is long relative to the relaxation times allows a new steady-state of populations to arise IS, cross-relaxation rate: dictates the sign of the NOE IS, dipolar longitudinal relaxation rate of spin I: it serves to reduce the magnitude Thus, NOE is related to molecular motion! NMR Spectroscopy Nuclear Overhauser Effect (NOE) 1H at 400 MHz W1 at 400 MHz W2 at 800 MHz (WI+WS)- stimulated by rapidly tumbling molecules W0 at Hz-kHZ (|WI-WS|)- stimulated by slowly tumbling molecules Small molecules exhibit positive NOEs Large molecules exhibit negative NOEs NMR Spectroscopy Nuclear Overhauser Effect (NOE) c=1.12 Variation in NOE as a function of molecular tumbling rates NMR Spectroscopy Field gradient Bg Variation of magnetic field strength along the z axis NMR Spectroscopy Field gradient -!Bg +!Bg 90º -Bg, !g Bg, !g +!B -!Bg g defocused (dephased) refocused (rephased) NMR Spectroscopy Field gradient x RF Gz stronger gradient NMR Spectroscopy Field gradient x RF Gz Variation of the second gradient pulse (90 to 110% of the first) NMR Spectroscopy Diffusion-ordered spectroscopy Bg !! x !! x "! #! !! "! #! G gradient strength D diffusion coefficient NMR Spectroscopy Diffusion-ordered spectroscopy mobility NMR Spectroscopy Multi-dimensional NMR One dimension Two dimensions NMR Spectroscopy Multi-dimensional NMR To generate a spectrum with two frequency domains, f1 and f2, it is necessary to sample data as a function of two separate time variables, t1 and t2. General scheme for 2D NMR experiment D P: Preparation E: Evolution M: Mixing D: Detection P E t1 M t2 NMR Spectroscopy Multi-dimensional NMR A hv B NMR Spectroscopy Multi-dimensional NMR NMR Spectroscopy COSY (COrrelated SpectroscopY) Correlation through bonds (J-coupling) NMR Spectroscopy TOCSY (Total COrrelated SpectroscopY) Correlation through bonds (J-coupling) x !m t1 t2 spin-lock JAB A !m A A A B B B B JBC C D E C D E D E C C JCD D JDE E NMR Spectroscopy COSY vs. TOCSY Correlation through bonds (J-coupling) COSY TOCSY NMR Spectroscopy COSY vs. TOCSY Correlation through bonds (J-coupling) NMR Spectroscopy General schemes for 2D NMR Relative sensitivity P a) H E H t1 1H-13C 1H-15N 1 1 1 2.5 4 10 (traditional) t2 H t1 X d) 1H-31P t2 X c) D t1 X b) M 4 8 30 10 32 300 t2 H X t1 t2 (inverse) modern NMR Spectroscopy Heteronuclear Single Quantum Coherence (HSQC) 13C 1H NMR Spectroscopy Protein NMR 2D NOESY NMR Spectroscopy Protein NMR 2D NOESY NMR Spectroscopy Protein NMR Isotopically labeled proteins NMR Spectroscopy Protein NMR 1H-15N HSQC (protein’s fingerprint) 15N 1H NMR Spectroscopy Protein NMR Signal overlap problem alleviated by 3D & 4D NMR 110 ppm 120 ppm 130 ppm 120 ppm 130 ppm 110 ppm 15N 15N 15N 1H 1H 1H 6.5 ppm 6.5 ppm 6.5 ppm NMR Spectroscopy Protein NMR Signal overlap problem alleviated by 3D & 4D NMR 2D 110 ppm 3D 120 ppm 130 ppm 120 ppm 130 ppm 110 ppm 15N 15N 15N 1H 1H 1H 6.5 ppm 6.5 ppm 6.5 ppm F2 (15N) F3 (NH) NMR Spectroscopy Protein NMR Signal overlap problem alleviated by 3D & 4D NMR NMR Spectroscopy Protein NMR Signal overlap problem alleviated by 3D & 4D NMR NMR Spectroscopy Protein NMR Assignment - Triple Resonance Experiments H C H N C C H H O 3D HNCA NMR Spectroscopy Protein NMR Assignment - Triple Resonance Experiments NMR Spectroscopy Protein NMR Assignment - Triple Resonance Experiments 15N ppm 115 118 122 125 116 130 128 40 45 13C 50 55 6.5 6.2 7.0 7.2 1H ppm 7.8 8.5 8.0 ppm NMR Spectroscopy Protein NMR Assignment - Triple Resonance Experiments NMR Spectroscopy Protein NMR Assignment - Triple Resonance Experiments HNCA 130 ppm 8.5 HN(CO)CA 130 ppm 8.5 i-1 HNCA 40 HN(CO)CA 125 ppm 125 ppm HNCA 40 125 ppm HN(CO)CA 125 ppm 40 45 45 45 50 50 50 55 55 55 7.5 7.5 i 7.5 7.5 i+1