* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download The Epigenetic Memory Different timing of bud burst between epitypes
Survey
Document related concepts
Transcript
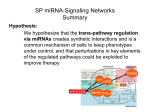
31st May 2016 IUFRO Genomics and Forest Tree Genetics Igor Yakovlev The Epigenetic Memory Induced by the Temperature During Embryo Development Impact on Bud Phenology Different timing of bud burst between epitypes • Related to development • Involved in adaptation and decease resistance • Encoded in chromatin and non-coding part of genome What we know about epigenetic memory so far The epigenetic memory is defined as a heritable change in gene expression or behavior that is induced by a previous stimulus (D’Urso & Brickner 2014) • Temperature sum during embryogenesis induce epigenetic memory • The effect lasts for life (at least 25 years……) • The effect may be adaptive and may decide life or death (i.e. frost damage) • Epigenetic memory is associated with visible transcriptome changes both during establishment in embryos and further maintenance in buds and needles (Yakovlev et al., 2014, 2016) Multiple molecular mechanisms are involved • Epigenetic regulation mechanisms Non-coding RNAs Long ncRNAs DNA Methylation Cytosine methylation Adenine methylation small small RNAs RNAs Gene Expression Histone Modifications methylation, acetylation, phosphorylation, ubiquitination, sumoylation, etc… Chromatin remodeling repositioning of nucleosomes incorporation of histone variants All these mechanisms realized by the specific pathways and demands specific enzymes miRNA pathway Plant microRNAs (miRNAs), a class of small non-coding regulatory RNAs, are canonically 20–24 nucleotides in length and bind to complementary target RNA sequences, guiding target attenuation via mRNA degradation or translation inhibition. They should be traceable back to precursor with a hairpin structure Yang & Li, 2012 Biology Experimental approaches The main task of current study is to identify and characterize miRNAs as important regulatory elements involved in the initiation and maintenance of epigenetic memory Somatic embryo tissues were induced from one genotype B10W of the full-sib family of Picea abies greenhouse and tested in the field conditions. Obtained epitypes show clear phenotypic differences related to memory of temperature during embryogenesis, confirming as obsereved differences in timing of bud burst We constructed 9 small RNA and 9 mRNA libraries Temperature conditions cold (18oC), normal (23oC) warm (28oC) + + + Intermediate – E2 + + + Mature embryos – E3 + + + Stages of embryo development Early – E1 miRNA profiling in somatic embryos of Norway spruce under different growth temperatures We defined about 2300 miRNA candidates with prevailing length 21 nt (41%) and 22 nt (34%). Among them we selected 1115 highly expressed miRNAs (>100 reads). Allowing up to 2 mismatches, we defined 522 conserved miRNAs belonged to 58 miRNA families and 593 novel miRNAs. Differential expression analysis revealed 676 differential expressed miRNAs 676 DE miRNAs could be grouped to 12 main clusters 202 DE miRNAs 23 DE miRNAs 423 DE miRNAs Differences in abundant transcript families between 18°C compared to the epitype inducing temperatures of 23°C and 28°C were largely qualitative, while the differences between 23°C and 28°C conditions were mostly quantitative Prediction of targets for highly expressed miRNAs We defined 470 miRNAs targeting to 1139 gene models, incl. 930 annotated gene models from around 212 gene families with diverse biological functions and 209 gene models without match at the Databases We found high redundancy in miRNA – mRNA target pairs We found that 317 DEGs could be regulated by several miRNAs and 290 miRNAs could regulate more than one gene a. MA_10433003g0010 Disease resistance protein (TIR-NBSLRR class) family; PF00931 - NB-ARC domain; PF01582; PF13676 TIR domain; PF01637 - Archaeal ATPase; PF01656 - CobQ/CobB/MinD/ParA nucleotide binding domain; PF03205-Molybdopterin guanine dinucleotide synthesis protein B; PF03308-ArgK protein; PF04665-Poxvirus A32 protein; PF05729NACHT domain; PF06745-KaiC; PF08477-Mirolike protein; PF13191-AAA ATPase domain; PF13173; PF13207; PF13238; PF13401-AAA domain Comparison of transcription profiles of selected differentially expressed miRNA–gene functional pairs, involved in temperature-dependent regulatory processes MA_97097g0010 MA_10433968g0020 MA_8090186g0010 MA_2193g0020 PF00097 - Zinc finger, C3HC4 type (RING finger); PF03031 - NLI interacting factor-like phosphatase; PF11793 - FANCL C-terminal domain; PF12678 - RING-H2 zinc finger; PF12861 - Anaphase-promoting complex subunit 11 RING-H2 finger PF03031 - NLI interacting factor-like phosphatase; PF07527 - Hairy Orange domain PF00847 - AP2 domain PF00847 - AP2 domain Comparison of transcription profiles of selected differentially expressed miRNA–gene functional pairs, involved in temperature-dependent regulatory processes MA_10436505g0010 MA_17313g0010 MA_88843g0010 PF08744 - Plant transcription factor NOZZLE PF02045 - CCAAT-binding transcription factor (CBFB/NF-YA) subunit B PF00145 - C-5 cytosine-specific DNA methylase; PF01426 - BAH domain; PF12047 - Cytosine specific DNA methyltransferase replication foci domain Characterization of numbers of differentially expressed miRNAs and their putative differentially expressed targets* preliminary annotated based on Pfam domains ## Pfam ID 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 PF00560 PF00069 PF00931 PF01535 PF01582 PF00004 PF00515 PF00637 PF00394 PF02536 PF00646 PF00249 PF00847 PF01397 PF00418 PF06345 PF00566 PF01715 PF00046 PF00201 PF08263 PF11721 PF04937 PF08744 PF03110 PF00106 … Pfam description Leucine Rich Repeat (LRR) Protein kinase domain NB-ARC (nucleotide-binding adaptor R-gene shared) domain Pentatricopeptide (PPR) repeat Toll-Interleukin receptor (TIR) domain ATPase family associated with various cellular activities (AAA) Tetratricopeptide (TPR) repeat Clathrin heavy chain/ VPS (vacuolar protein sorting-associated) Multicopper oxidase mTERF (Mitochondrial transcription termination factor) F-box domain Myb-like DNA-binding domain AP2 domain Terpene synthase, N-terminal domain Microtubule-associated protein (MAP) Tau, tubulin-binding repeat DRF (Diaphanous-related formins) autoregulatory domain Rab-GTPase-TBC domain tRNA Delta(2)-isopentenylpyrophosphate transferase (IPP transferase) Homeobox domain UDP-glucoronosyl and UDP-glucosyl transferase Leucine rich repeat N-terminal domain Di-glucose binding within endoplasmic reticulum Protein of unknown function (DUF 659) Plant transcription factor NOZZLE SBP (SQUAMOSA promoter binding protein-like) domain short chain dehydrogenase Number of Number of miRNAs gene models 274 139 169 105 138 85 57 40 44 15 11 12 8 8 7 11 5 15 11 5 4 4 3 3 40 7 163 97 90 84 52 52 44 30 15 11 11 10 8 8 5 4 4 3 3 3 3 3 3 3 2 2 * - evaluation based on variants with negative correlation (-0.6 … -1.0) between DE miRNAs target gene expression patterns. Targets were predicted by psRNATarget Differentially expressed miRNAs targeting epigenetic regulators Number of Number of DEGs miRNAs DNA methylation 4 4 Histone methylation 41 38 Histone acetylation 7 7 136 70 Histone ubiquitination 9 9 Chromatin remodeling 7 7 sRNA pathways 4 4 Thermosensing 1 1 Histone phosphorylation Total 212 differentially expressed miRNA – target pairs, composed by 125 miRNAs and 204 gene models Conclusions • We showed that Norway spruce poses a variety of miRNAs and their isoforms with distinct temperature dependent expression patterns • We found around 500 differentially expressed miRNAs targeting to 1139 gene models • Target genes are mostly represented by transcripts of multiple repeats proteins like TIR, NBS-LRR protein genes, PPR and TPR repeat, Clathrin heavy chains/VPS proteins, etc. , including also genes involved in epigenetic regulation • We consider that TIR, NBS and LRR domain containing proteins could fulfill more general functions for signals transduction from external environment and conversion them into molecular response of any type Conclusions • More than 200 DEGs of epigenetic regulators (from about 700) could be post-transcriptionally regulated by 125 miRNAs • Except the protein kinases, miRNAs mostly involved in regulation of genes related to methylation modifications, both in histones and DNA • Unknown mechanisms provide fine-tuning of miRNAs pool content participating in developmental regulation and epigenetic memory formation. • Taking together, our data confirm significant involvement and importance of miRNAs into post-transcriptional gene regulation and epigenetic memory formation People involved: NIBIO: Igor Yakovlev, Vivian-Smith A. , Carneros E., and Carl G. Fossdal NMBU (Norwegian University of life Sciences) : Jorunn E. Olsen, Lee Y.K., Marcos Viejo Funding: FRIPRO (NFR), EU FP7, TOPPFORSK (NFR)