* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download rec07
Genetic algorithm wikipedia , lookup
Mathematical optimization wikipedia , lookup
Corecursion wikipedia , lookup
Knapsack problem wikipedia , lookup
Multiple-criteria decision analysis wikipedia , lookup
Computational complexity theory wikipedia , lookup
Factorization of polynomials over finite fields wikipedia , lookup
Graph coloring wikipedia , lookup
Gene expression programming wikipedia , lookup
Comp. Genomics
Recitation 7
Clustering and analysis of microarrays
Exercise 1
• A microarray that contains probes for all
the N metabolic enzymes of the bacterium
D.Angerous was used for the following
time-series microarray experiment: The
bacteria population were exposed to a
drug, and gene expression was measured
every hour for M hours.
• The expression values are discretized to
{-1,0,1}
Exercise 1
• Find the longest expression pattern that is
common to at least k enzymes. Each
enzyme may start the pattern at a
different time.
K=3
T1
T2
T3
T4
T5
T6
T7
E1
-1
0
1
-1
0
1
-1
E2
-1
-1
0
1
-1
0
1
E3
-1
-1
-1
0
1
-1
0
E4
0
0
0
0
0
0
0
E5
1
1
1
1
0
1
1
E6
1
-1
-1
-1
-1
0
-1
Solution
• Treat each expression vector as a string
• Create a generalized suffix tree O(MN)
• Find longest k-common substring
Exercise 2
• Expression of N genes was measured
under a certain condition using a
microarray.
• No discretization was performed.
• Give a polynomial time algorithm for
clustering these genes into exactly k
clusters.
k
• The objective function is max{ d (i, j ) | i, j Cl }
l 1
Pictorially
G1
G2
G3
G4
G5
G6
If {G3,G4,G5}is a cluster, its contribution to the
objective function is d(G3,G5)
Expression level
Solution
• Create a weighted directed graph, every
gene is a node and the edge from i to j
has weight d(i,j-1) if i’s expression is
lower than j’s (otherwise ∞)
G1
G2
G3
G4
G5
G6
The path in the graph that corresponds to this clustering
is G1G3G6. The value of the objective function is
d(G1,G2)+d(G3,G5)+0
Solution
• Next:
• Find the shortest path that visits exactly k
nodes
• Dynamic programming:
Pj (k ) min Pl (k 1) d (l , j )
lk ,.., j 1
Start from k because if l<k Pl(k-1)=∞
Exercise 3
• A microarray experiment with N genes
and M conditions was conducted
• Describe a polynomial algorithm that
determines whether the genes can be
clustered into 2 clusters such that the
maximum distance d(Gi,Gj) in each cluster
<W
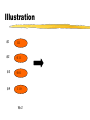
Illustration
G1
111
G2
011
G3
001
G4
110
W=2
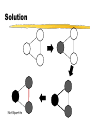
Solution
• Create a graph with a node for every gene
• Add an edge (i,j) if d(i,j)> W
• Check if the resulting graph is bipartite:
Run BFS, if you discover an edge (u,v) to
a gray node and the depths of u and v are
both even or both odd, answer: “no”.
Solution
Not Bipartite
Exercise 4
• We are given a microarray with N genes
and M experiments
• We want to cluster the genes into k
clusters such that the distance between
genes that belong to the same cluster will
be < W
• Can you give a polynomial algorithm that
solves this problem?
Solution
• Probably not
• More specifically, if we could solve this
problem in polynomial time, we could
solve a large class of problem that are
widely believed to be unsolvable in
polynomial time
Solution
• How can we show that we can probably
not find a solution in polynomial time?
• We will take a problem for which this has
already been shown
• We will construct a polynomial time
reduction to our problem
• So, if our problem could be solved
efficiently the “hard” problem could also
be solved efficiently
Graph description
The following graph can describe our
problem:
G2
G1
G3
G6
G5
G4
There’s an edge (Gi,Gj) if
the distance between Gi and
Gj is less than W
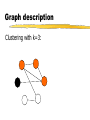
Graph description
Clustering with k=3:
3COL
3-Colorability: Given a graph G, can we dye
its vertices with 3 different colors such that
no two adjacent nodes have the same
color?
Comparing the problems
•
•
•
•
What is common to both these problems?
In both we “cluster” the nodes
What are the differences?
First, in 3COL there are only 3 clusters
instead of k
• Second, the elements that belong to the
same group in 3COL must not have edges
between them
Reduction
• Now that we understand the differences,
we can take a graph G that is an input to
3COL, and transform it to a graph G’ and
a constant k that are the input to the kclustering problem
• We assume that we have a polynomial kclustering algorithm, and we apply it to
(G’,k) and translate the solution to 3COL
Reduction
• Given the first difference that we noted,
what should be the value of k?
• We set k to 3, i.e. the algorithm should
find exactly 3 clusters
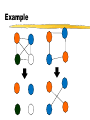
• How do we change G to get G’?
• G’ has the complement edges of G
Example
Proof
Suppose that G is 3 colorable. Let
V1,V2,V3 be the groups of nodes that can
be colored by distinct colors. There are
no edges between any pair of nodes in V1
,and therefore it forms a legal cluster in
G’. Similarly, the nodes of V2 and V3 form
clusters. Since V1UV2UV3 contains all the
nodes all the genes are clustered in the 3
corresponding clusters.
Proof, second direction
Suppose that G’ contains a clustering to 3
legal clusters. These clusters correspond to
3 nodes sets in G such that within each set
there are no edges between pairs of nodes.
Therefore, assigning a different color to
every set is a 3-coloring.