* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
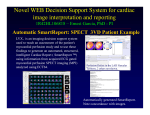
Download imaging three-dimensional cardiac function
Cardiac contractility modulation wikipedia , lookup
Management of acute coronary syndrome wikipedia , lookup
Electrocardiography wikipedia , lookup
Echocardiography wikipedia , lookup
Quantium Medical Cardiac Output wikipedia , lookup
Myocardial infarction wikipedia , lookup
Arrhythmogenic right ventricular dysplasia wikipedia , lookup
P1: FHY June 13, 2000 11:12 Annual Reviews AR106-16 Annu. Rev. Biomed. Eng. 2000. 02:431–56 c 2000 by Annual Reviews. All rights reserved Copyright IMAGING THREE-DIMENSIONAL CARDIAC FUNCTION ? Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. W. G. O’Dell and A. D. McCulloch Department of Bioengineering, University of California San Diego, La Jolla, California 92093-0412; e-mail: [email protected] Key Words review, myocardial strain, myocardial stress, 3D imaging ■ Abstract The three-dimensional (3-D) nature of myocardial deformations is dependent on ventricular geometry, muscle fiber architecture, wall stresses, and myocardial-material properties. The imaging modalities of X-ray angiography, echocardiography, computed tomography, and magnetic resonance (MR) imaging (MRI) are described in the context of visualizing and quantifying cardiac mechanical function. The quantification of ventricular anatomy and cavity volumes is then reviewed, and surface reconstructions in three dimensions are demonstrated. The imaging of myocardial wall motion is discussed, with an emphasis on current MRI and tissue Doppler imaging techniques and their potential clinical applications. Calculation of 3-D regional strains from motion maps is reviewed and illustrated with clinical MRI tagging results. We conclude by presenting a promising technique to assess myocardial-fiber architecture, and we outline its potential applications, in conjunction with quantification of anatomy and regional strains, for the determination of myocardial stress and work distributions. The quantification of multiple components of 3-D cardiac function has potential for both fundamental-science and clinical applications. CONTENTS INTRODUCTION . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . IMAGING MODALITIES FOR GEOMETRIC MEASUREMENT . . . . . . . . . . . . . Overview . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . X-ray Ventriculography . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . Radionuclide Ventriculography . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . Echocardiography . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . Computed Tomography . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . Magnetic Resonance Imaging . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . ANALYSES OF VENTRICULAR SIZE AND GEOMETRY . . . . . . . . . . . . . . . . . Wall Thickness Measurements . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . Ventricular-Cavity Volume Estimates . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . Surface Modeling with Ellipsoids . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . Volumes from Three-Dimensional Surface Reconstructions . . . . . . . . . . . . . . . . . 1523-9829/00/0825-0431$14.00 432 433 433 433 434 434 435 436 437 437 438 439 440 431 P1: FHY June 13, 2000 432 11:12 Annual Reviews O’DELL ¥ AR106-16 MCCULLOCH Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. DEFORMATION ANALYSIS . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . Implanted Markers . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . Magnetic Resonance Imaging Tagging . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . Strain Rate Analysis . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . Slice Motion Correction . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . Model-Based Deformation Analysis . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . Limitations of the Reconstructions . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . REGIONAL STRAINS IN DISEASE . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . MEASUREMENT OF THREE-DIMENSIONAL TISSUE ARCHITECTURE . . . . . FUTURE DIRECTIONS . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . ? 441 443 443 444 445 446 446 447 448 449 INTRODUCTION The three-dimensional (3-D) nature of heart deformation is well appreciated and a subject of great interest both to cardiovascular scientists and clinical cardiologists. The application of clinical imaging to cardiac function has traditionally been limited to global geometric measurements, including left ventricular (LV) wall mass, ventricular volume, stroke volume, ejection fraction, and wall thickness (1). These measures of heart function provide invaluable diagnostic information to the clinician regarding the severity of disease and the long-term prognosis (2). The ability to measure regional myocardial deformation provides additional information that is valuable for identifying the location and extent of affected areas, quantifying the degree of mechanical dysfunction, and differentiating between functionally distinct disease states. Among the investigational tools available, visualization and quantification of regional cardiac mechanical function are perhaps the most direct and reliable indicators of cardiac health (3). For example, although quantification of coronary artery patency is important in directing treatment of regional myocardial ischemia, the existence of severe vessel occlusion is not necessarily associated with insufficient mechanical function, owing to myocardial revascularization and ventricular wall remodeling. The realm of 3-D cardiac mechanical analysis encompasses global and regional myocardial functions, 3-D fiber tissue architecture, electrical propagation of activation, the generation of internal wall stresses by the contracting myocytes, and the distribution of those stresses by the surrounding tissue matrix. This article describes and contrasts the prevailing modalities for monitoring 3-D heart function, describes the current techniques for manipulating image data to model cardiac anatomy and deformation, and outlines what lies ahead for the field of functional cardiac imaging. Length constraints preclude coverage of other areas of cardiac imaging research and applications, such as metabolic imaging [positron emission tomography (PET) and magnetic resonance spectroscopy (MRS)], electric- and magnetic-field imaging, electrical propagation and mechanoelectrical feedback, and perfusion imaging. P1: FHY June 13, 2000 11:12 Annual Reviews AR106-16 IMAGING 3-D CARDIAC FUNCTION 433 IMAGING MODALITIES FOR GEOMETRIC MEASUREMENT Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. Overview Various imaging modalities exist to measure heart wall geometry. These can be compared objectively by the following criteria: signal quality [indicated by the signal to noise ratio (SNR)], discrimination between myocardial and neighboring tissue [indicated by the contrast to noise ratio (CNR)], temporal and spatial resolution, susceptibility to image blurring and artifact, acquisition and analysis time, ease of use, relative cost, and availability. For accurate quantification of heart wall geometry and ventricular volumes, CNR is often the more informative measure because it governs the ability to discern tissue boundaries. Spatial resolution is typically given by the pixel dimensions in the two-dimensional (2-D) image (voxel dimensions for 3-D); however, for some modalities, such as MR imaging (MRI) and computed tomography (CT), the image slice thickness can be comparatively large, leading to partial-volume artifacts that diminish the effective in-plane resolution. Temporal resolution relates to both the time interval between successive images (temporal sampling rate) and the time over which the data for each image are acquired (acquisition window). In X-ray imaging, motion while the shutter is open leads to blurring and loss of image quality. In MRI, motion during acquisition causes a phase dispersion across the image, creating a characteristic artifact. Generally one needs to acquire data on a time scale that is smaller than the fastest motions of interest. For a typical heart rate of 1 Hz, the duration of isovolumic relaxation or isovolumic contraction is ∼100 ms; hence a temporal sampling rate of <50 ms is required to capture these events. In many instances, increasing either spatial or temporal resolution competes with the desire to minimize acquisition time. Reduction of spurious cardiac motion is critical for all imaging modalities. Image acquisition during periods of patient breath holding (preferably at end-expiration) and electro-cardiography gating, either prospective (triggering the acquisition to occur at a specific point in the cardiac cycle) or retrospective (assigning time stamps to the data after acquisition), can substantially reduce image blurring and motion artifact. With retrospective gating, image data can be acquired continuously so that overall imaging time may be reduced, possibly improving patient compliance. However, a longer period of post-processing is then required. ? X-ray Ventriculography By 1964, X-ray ventriculography with contrast enhancement was considered the gold standard for ventricular volume and mass measurement (4). In this method, conventional X-ray images are generated by projecting a wide beam of X-ray energy onto a film. Radiopaque substances in the tissue absorb and scatter the incoming energy; hence an X-ray image is an inverse mapping of tissue opacity. P1: FHY June 13, 2000 Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. 434 11:12 Annual Reviews O’DELL ¥ AR106-16 MCCULLOCH The theoretical resolution of a film image is dependent on the film grain; however, the effective resolution depends on the relative opacity of neighboring objects, the dispersion of the X-ray beam, and the distances of objects from the X-ray tube focal spot and film. Projection images in orthogonal views depict major heart long- and short-axis dimensions. X-ray projection imaging with contrast is considered to be mildly invasive because of both the contrast injection and the ionizing radiation. For wall geometry measurements, there are the problems of limited CNR at the cavity boundaries, silhouette hiding of concavities in the ventricular surface, and various projection and registration artifacts. In biplanar cineradiographic imaging, there are also technical concerns about temporally and spatially registering the images from the two cameras. As an example of the potential accuracy of biplanar X-ray imaging, when implanted bead markers are imaged in canine subjects, singleplane views can distinguish and locate the centers of even partially overlapping 1-mm-diameter beads to an accuracy of 0.2 mm (5). However, a typical bead registration error between two cameras that are oriented perpendicularly to each other is 0.1 mm (6, 7). Movie film acquisition rates of 60–90 Hz are common. X-ray projection systems are relatively inexpensive, easy to use, and widely available, and they can provide real-time feedback, but the accumulated radiation dose is a concern for repeated measurements. ? Radionuclide Ventriculography Blood-pool or equilibrium radionuclide ventriculography is another common modality for estimating ventricular volumes. Erythrocytes are labeled in vivo with the radioactive tracer technetium-99m. A large-field gamma camera equipped with a low-energy collimator records the number of radio-decay counts, which is proportional to the number of erythrocytes and, hence, the blood volume. The resulting time-activity curves are often filtered in time, normalized by the end-diastolic volume count, and used to compute ejection fraction, peak filling/ejection rates, time to peak filling/ejection rates, and related indices (8). This technique is easy to use and measures relative volumes without the need for geometric assumptions, but it is susceptible to background signal sources, has relatively low spatial resolution, and is susceptible to variable attenuation during respiration. Long acquisition times of ∼5 min require a regular cardiac rhythm and/or post-processor elimination of data from premature beats and temporal and spatial smoothing. Tomographic radionuclide imaging or gated blood pool single photon emission CT (SPECT), was first demonstrated in 1980 (9) but so far has had only limited clinical application. The SPECT acquisition is accomplished within 30 min, with a rotating gamma camera and 60 projection angles (10). The 3-D images can then be reconstructed with conventional CT back-projection algorithms. Echocardiography Among the current cardiac imaging modalities, echocardiography (ECG) is perhaps the most prevalent because of its cost effectiveness, ease of use, real-time P1: FHY June 13, 2000 11:12 Annual Reviews AR106-16 Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. IMAGING 3-D CARDIAC FUNCTION 435 feedback, portability, and widespread availability. Echocardiographic imaging is based on the partial reflection of acoustic waves at tissue boundaries and echocardiographic image contrast is governed by the difference in acoustic properties between adjacent tissues. A pencil-shaped beam (M-mode) is swept though an arc of ∼80◦ to create a 2-D image at 16–38 frames/s. Transthoracic imaging (in which the echocardiographic transducer is manually placed on the skin) can be used to obtain multiple long-axis, short-axis, and oblique-image views, but it is limited to parasternal and apical windows. Transesophageal imaging (in which an echocardiographic transducer is inserted into the esophagus) overcomes the windowing constraints and places the transducer in close proximity to the heart. Many of the limitations of 2-D and M-mode ECG are overcome by 3-D ECG, including limited viewing angles and imaging obliquely through the LV wall, and this method is quickly approaching clinical use (11, 12). In 1990, von Ramm & Smith (13) introduced the first fully 3-D, phased-array cardiac ultrasound imager. Currently, 3-D ECG in patients is generally applied by one of two methods: transthoracic imaging in multiple views with transducer position registration [with either a mechanical arm or an acoustic spatial-location system (14)]; or transesophageal imaging with either incremental rotation of the transducer at various long-axis imaging planes or translation of the transducer at various short-axis imaging planes (11). A typical transthoracic 3-D ECG imaging set, as described by Gopal et al (14), is composed of 7–10 short-axis images and is acquired in 6–8 min. Analogous long-axis 3-D image sets acquired from rotated apical views are also feasible; a method introduced by Ghosh et al (15) in 1982. 3-D ECG can also be used to assess right ventricular (RV) geometry (16). ECG image quality is limited by relatively low contrast between myocardial and adjacent tissues, fading of the endocardial boundaries (typically, ∼25% of the total boundary in a time-elapsed M-mode recording is indiscernible) (17), noise, and related artifacts. With a contracting LV phantom, Lange et al (17) found that the error in volume estimation with multiple parallel ECG slices and Simpson’s rule was 3% at the larger volume (simulating end-diastole) and 4% at the smaller volume (end-systole). CNR differences account for a discrepancy of 9% for the end-diastolic volume and 11% for the end-systolic volume between the ECG-estimated LV volumes and those produced by the more reliable tissue Doppler imaging (TDI) method for contour detection (17). Without the addition of contrast agents, ECG CNRs generally tend to be lower than those for MRI and contrast-enhanced CT. ? Computed Tomography The conventional CT scanner consists of an array of detectors and a single X-ray source, which is rotated about the sample. The transmitted fan-shaped X-ray beam is recorded at several angles, and a 2-D image is reconstructed by using a backprojection algorithm. Radiographic contrast injection and rapid scanning are commonly used to improve image quality. CT with contrast offers very good CNR with P1: FHY June 13, 2000 Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. 436 11:12 Annual Reviews O’DELL ¥ AR106-16 MCCULLOCH high spatial resolution (typically 2- × 2- × 5-mm voxel dimensions), high temporal resolution (∼1 s), and the ability to acquire axial images at arbitrary locations through the chest. CT scanners, however, are more expensive and less available than ECG imagers. The simultaneous acquisition of multiple 2-D CT images for 3-D–image reconstruction was initially demonstrated in 1980, with the dynamic spatial reconstructor [DSR (18)]. The DSR uses multiple X-ray sources with coneshaped beams and a hemicylindrical fluorescent screen to acquire volumetric data with high temporal resolution. Whereas the DSR has remained primarily a research tool, electron-beam CT (EBCT) (19) has become increasingly popular for clinical use. In EBCT, X-rays are produced by scanning a single-source electron beam onto a tungsten target that is positioned in a semicircle below the patient (20). Mechanical motion in the gantry is eliminated (bypassing the typical 1-s interscan delay), and exposure times of 50–100 ms are possible. In typical multislice mode, four tungsten target rings and two detector rings are used to acquire up to eight image slices without movement of the patient table, reducing motion artifact. Spiral CT (also known as helical CT) is another emerging technique for very rapid acquisition of 3-D image data; with higher spatial but lower temporal resolution than EBCT. Spiral CT imaging uses a conventional fan-shaped beam source, which rotates continuously about the patient, and a fixed array of receivers. Contiguous 2-D axial images are acquired as the sample moves along the scanner bore. Multiple receivers are often used to acquire four slices simultaneously. Retrospective gating is commonly used in EBCT and spiral CT to reduce motion artifact (21). With both multislice EBCT and multislice spiral CT, 3-D image data sets can be acquired in times on the order of seconds—quickly enough that susceptibility to remaining motion artifact is significantly reduced. On-line reconstruction algorithms have been developed to aid in the graphical display of 3-D CT cardiac data sets and in simulating views from obliquely oriented cross sections (22). ? Magnetic Resonance Imaging Magnetic resonance (MR) imaging (MRI) both provides high CNRs between soft tissues and blood without the injection of contrast agents and enables imaging at arbitrary image angles. MRI scans are relatively expensive and time consuming, and they require a certain amount of patient cooperation. In-plane spatial resolution can be high (1 × 1 mm); however, the slice thickness is typically 5–10 mm; thus the images are highly susceptible to partial-volume artifacts. This is problematic especially towards the heart apex, where the wall slope is high. The quality and resolution of MR cardiac images have steadily improved with the inclusion of breath holding, black-blood saturation (which improves the contrast between the myocardium and blood in the cavity), rapid imaging techniques (i.e. gradient echo and echo-planar imaging for quicker acquisition), k-space segmentation (for improved temporal resolution), and dedicated cardiac radiofrequency receiver coils (for improved SNR) (23). With prospective gating and gradient echo sequences, P1: FHY June 13, 2000 11:12 Annual Reviews AR106-16 Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. IMAGING 3-D CARDIAC FUNCTION 437 the image data for 10–14 temporal images at a single slice location in the heart can be acquired within a 16- to 20-s breath hold. Breath holds of this duration are feasible in cardiac patients, even those with acute myocardial infarction (3). The acquisition of cine data for 8–10 short-axis slices thus requires ∼10–15 min, allowing for patient recovery after each breath hold. Newer sequences, using SMASH (simultaneous acquisition of spatial harmonics) for example (24), can cut this time by a factor of 2 or 3. MRI can also be used to perform retrospective respiratory gating with navigator echo signals from the diaphragm, although the implementation of this technique is difficult (25). A recent review of MRI applied to cardiac motion analysis by McVeigh (25) is highly recommended. ? ANALYSES OF VENTRICULAR SIZE AND GEOMETRY LV volume, ejection fraction, and wall thickness are perhaps the most frequently used indices of cardiac performance and patient prognosis (26, 27). However, the pertinent geometric information must first be extracted from the images before 3-D surface reconstruction. The challenge is to detect, within the image, heart contour features that are irregular in shape and position, changing with time, and varying in contrast and whose image intensity gradient profile can change in both magnitude and sign. Contour segmentation is the most laborious and error-prone part of most analysis schemes. Real-time clinical application of modern 3-D cardiac mechanics techniques is going to require faster and better automated segmentation. We refer the reader to the article in this volume of Annual Review of Biomedical Engineering that pertains to image segmentation, by Pham et al (27a). Wall Thickness Measurements LV wall thickness measurements have been linked to regional ischemia (27) and lead to estimates of mean wall stress (28) and mean wall stiffness. Wall thickness measurements were made as early as 1964, by X-ray angiography (4), but more recently Jakob et al (29) improved the technique by using injected-contrast and digital-subtraction angiography. In comparison with the more commonly used Mmode ECG, the Jakob method showed good agreement towards end-diastole but poorer agreement at end-systole. Although wall thickness changes are often the most apparent geometric indicator of altered mechanical function (30), the relationship between wall thickness and cardiac health is not as clear. Significant correlations were found by Lawson et al (31) between thallium uptake and systolic wall thickness in ischemic hearts, but not with diastolic wall thickness and thallium uptake (31, 32). Similarly, Curiel et al (33) found that the magnitude of inotropic reserve is not related to diastolic wall thickness or to basal systolic wall thickening. Heupler et al (34) showed that the LV end-diastolic cavity diameter, not wall thickness, was associated with thallium perfusion defects and therefore myocardial ischemia. Dong et al (35) concluded P1: FHY June 13, 2000 Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. 438 11:12 Annual Reviews O’DELL ¥ AR106-16 MCCULLOCH that regional and interindividual heterogeneity in wall thickness in diseases such as hypertrophic cardiomyopathy make meaningful comparisons difficult. Finally, acute changes in wall thickness do not necessarily indicate chronic dysfunction because long-term wall remodeling may act to compensate for mild injury by restoring wall thickness to normal values, as described by Pouleur et al (36) with LV dilation. The relationship between wall thickening and fiber shortening is also not clear. Wall thickening at various wall depths was shown by Hexeberg et al (30) to correlate not with fiber shortening or myocardial perfusion in those layers, but with thickening of the entire wall and local wall geometry. McCulloch et al (37) supported this conclusion for fiber strains but found that transmural location did have a significant effect on cross-fiber strains. The estimate of the contour location in an image is accurate to within 1–2 mm, typically, with MRI (38). For a 10-mm-thick wall, this error leads to a 10%–20% uncertainty in the thickness estimation. The apparent wall thickness is increased in planar images that intersect the heart wall at oblique angles to the local surface normal, creating a bias in the thickness measurement. Through-plane motion introduces additional error because the same piece of myocardial tissue is not imaged over time at a fixed cross-sectional location. Both the bias and the through-plane motion artifact can be corrected with 3-D ventricular surface reconstruction by using data from orthogonal views (see below). ? Ventricular-Cavity Volume Estimates Traditionally, the measurement of LV volume by thermodilution techniques (39), projection ventriculography (40), or radionuclide techniques has been subject to substantial error, that is, ∼20% (39). Despite many innovations in imaging techniques and computerized analyses, substantial errors of this order remain in clinical measurements of LV volume. Currently, the task of estimating the volume of the ventricular cavity often reduces to defining the cavity boundaries and integrating over the enclosed space. Significant, common sources of error are poorly defined basal and apical limits of the ventricle, oversimplified geometric models, and uncompensated imaging artifacts. A common convention is to model the base of the heart as a planar surface passing through the mitral valve ring. However, the entirety of the valve plane is not easily imaged with ECG or 2-D CT imaging because the mitral valve is obliquely oriented in the chest cavity. With MRI, it is possible to orient the imaging plane to encompass the entire mitral valve ring, but, because the slice thickness is typically 5–7 mm, there is a ±2.5- to 3.5-mm discrepancy in the plane’s exact location within the slice. Even if accurately imaged at one time, the mitral valve ring translates apically ∼10 mm (41) from end-diastole to end-systole; hence it does not remain within a single, fixed short-axis image volume. For a mitral valve ring that is 25 mm in diameter (area, 491 mm2), a jump from one slice location to the next represents an estimated volume change, using Simpson’s rule, P1: FHY June 13, 2000 11:12 Annual Reviews AR106-16 Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. IMAGING 3-D CARDIAC FUNCTION 439 of 2.5–3.4 ml or ∼6.2%–8.5% of the total end-systolic volume (for a representative end-systolic volume of 40 ml). Similarly, it is difficult to detect the most apical intracavitary point by using only short-axis images. Both of these sources of error can be overcome by incorporating planar image data from the orthogonal, long-axis view. When referenced to 3-D ECG and equilibrium radionuclide angiography, limited-plane 2-D ECG methods were no better than subjective visual estimation for determining the LV ejection fraction (14). To compensate for the inherent limitations, some applications use correction factors that are calibrated to match the known volume errors as tested on postmortem or experimental heart models (40). However, these correction factors are computed for a “typical” heart geometry that may not necessarily be appropriate for abnormal or diseased hearts. Poor CNRs and partial-volume effects both contribute to errors in the segmentation of heart contour data, which are magnified in the LV volume estimate. For a sphere of radius 25 mm, a typical LV minor-axis radius dimension, an error of 1 mm (4%) in the estimated radius leads to a volume estimation error of ∼12%. All imaging modalities suffer misregistration of the heart contour data when the image data are acquired over many heartbeats or different times, as a result of beat-to-beat variations in the heart contraction and uncorrected respiration-derived motions. With projection images, the contrast silhouette obscures most of the indentations in the wall (i.e. at the papillary muscle sites), creating an overestimation of the true ventricular dimensions. A comparison between cross-sectional ECG and X-ray ventriculography found that projection imaging leads to an overestimation in ejection fraction of 22% (42). ? Surface Modeling with Ellipsoids When only a few ventricular surface measurements are available, it is convenient to model the LV surface geometry as an ellipsoid. This simplification allows for rapid calculation of volume but disregards the asymmetry of the actual ventricular surface, the complexity of the mitral valve plane, and the existence of the papillary muscles. Elliptical models were initially used with projection ventriculography (40), but are now also used commonly with 2-D ECG data (43). The volume of an ellipse is given by: V = 4 πab2 3 where a is the radius along the major axis and b is the radius of the minor axis. For single-plane projections, Dodge (40) used the formula b= Aproj πa where b is the minor axis dimension and Aproj is the area of the long-axis projection 4A2proj (making V = 3πa ). An analogous formula exists for biplanar data. Ellipsoidal P1: FHY June 13, 2000 Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. 440 11:12 Annual Reviews O’DELL ¥ AR106-16 MCCULLOCH models cannot as readily approximate RV volumes, because their geometry does not possess such symmetry (44). The Simpson’s-rule method computes the cavity volume by summing the areas of 2-D planar slices, scaled by the slice separation. The implied geometric assumption for this method is that the area of the measured cross-section remains constant over the distance between slices (or at least is representative of the average area), which becomes increasingly error-prone as the slice separation increases. There are also the previously mentioned errors associated with the identification of the mitral-valve plane, through-plane motion, and registration artifacts. A common modification to the volume equation involves reducing the volume contribution of the most apical and/or most basal slices by 50%. Many other variations are commonly used. ? Volumes from Three-Dimensional Surface Reconstructions Regardless of the modality used to acquire the anatomical data, once these are attained, 3-D surface reconstructions allow more exact descriptions of the heart geometry than simple elliptical models. Many 3-D surface reconstruction techniques are also able to incorporate anatomical data from multiple views/projections. Many recent advances in 3-D surface reconstruction have been driven by computer vision research (45, 46). Piecewise mathematical constructs have also been proposed, including 3-D splines and finite-element basis functions (47, 48). Deformable sheet or balloon models (3-D extensions of “snakes”) are able to interact directly with features of the images to satisfy both the need to match image data and to smooth out noisy image data. The smoothing is accomplished by incorporating energies associated with stretching and bending into the deformable sheet or balloon (49–51). Global polynomial representations are also applicable (52, 53). For example, we have used a surface model expressed as a polynomial series in prolate spheroidal coordinates. Here the base geometry is an ellipsoid, to which are superposed spatial modulations as a function of angular position, analogous to spherical harmonics. Mathematically, the radial coordinate λ is represented as a function of the circumferential and longitudinal angles (θ, φ): λ= l L X X a j Plm (cos(θ ))eimφ (1) l=0 m=−l Here Plm are the associated Legendre polynomials. The coefficients ai in the polynomial are fit, in a least-squares sense, simultaneously to contour data points from multiple views. The goodness/smoothness of the reconstruction can be adjusted to match the expected uncertainty in the contour data, by admitting or omitting higher-order terms in the series, which is achieved by adjusting the series limit “L.” Figure 1A (see color insert) shows the fitted endocardial surfaces for a zerothorder fit, that is, the best-fit prolate spheroid. From left to right in the figure, P1: FhN/FGI Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. July 31, 2000 P2: FPX/FOK QC: FHN/fgm 4:55 Annual Reviews T1: FhN AR106-01 ? Figure 1 Left ventricular endocardial surface-fitted data sets for canine heart. The light lines are the raw contour data, and the dark lines are generated from the fitting equation (Equation 1). Top: Top view; bottom: side view. The order of fitting goes from zero-th to third, sixth, and tenth from left to right. P1: FHY June 13, 2000 11:12 Annual Reviews AR106-16 Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. IMAGING 3-D CARDIAC FUNCTION 441 additional polynomial terms are added, giving the model increased spatial modulation (for deformable sheets and balloons, this is analogous to decreasing the bending energy). The error between the fitted surface and the original contour data can be computed as a function of the number of fitting parameters. The error will generally decrease as the number of fitting parameters increases, but there is a point at which the inclusion of additional high-order terms would fit the noise to the data rather than to the actual heart geometry. The combination of data from multiple views and the spatial and temporal smoothing will filter out noise and misregistration errors, creating a more accurate representation of the 3-D heart surfaces. Once the 3-D surface is mathematically reconstructed, the cavity volume and LV mass can be computed numerically. The time required to compute volume estimates from contour data from multiple views and multiple locations is negligible compared with the time required for the acquisition and segmentation steps. In the example shown in Figure 1D (see color insert), the 49 polynomial coefficients were fit to >1800 contour points in <1 min, using an SGI R5000 Indy workstation. The 3-D analytical surface representations are also useful for accurate measurement of regional wall thickness, where thickness compared with the true epicardial surface normal can be quickly and accurately computed. In review, LV volume and ejection fraction remain two of the most sought-after clinical indices of cardiac health. However, errors of 20% in LV volume estimation and ejection fraction are common clinically, although this uncertainty is often overlooked. Modern investigational and computational strategies afford much higher accuracy with only marginally more effort and time although the necessary imaging technologies and computation software are not yet widely available. ? DEFORMATION ANALYSIS Measures of ventricular cavity volume, volume changes over time, and myocardial mass have traditionally been used to gauge global myocardial mechanical dysfunction and overall patient prognosis. However, the ability to quantify the deformation of the myocardial tissue locally, for example to describe the amount of shortening in the direction of the myocardial fibers, is of paramount importance in cardiac mechanics research and promises to provide clinicians with much more powerful indices of regional tissue viability. A material point is a physical piece of myocardial tissue. The description of the deformation in the neighborhood of a material point begins with the task of identifying and tracking these points and their neighbors over time and space. The relative motion of neighboring points away from or toward each other reflects the amount of stretching or compression of the tissue, and is described by the onedimensional (1-D) displacement gradient, ∇U. If the 1-D displacement (along the x-axis, for instance) of one material point differs from that of a second, neighboring point located initially a distance 1x away and the displacement difference is given P1: FHY June 13, 2000 442 11:12 Annual Reviews O’DELL ¥ AR106-16 MCCULLOCH Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. by 1u x , then the displacement gradient is 1u x /1x. The infinitesimal displacement gradient is δu x /δx. If a second degree of motion is added, along a y-axis, then there is a potential for stretch or compression along this second axis and for rotation and/or shearing in the plane of motion. The complete 2-D motion is described by a nonsymmetric 2-D displacement gradient tensor: ∂u x ∂u x ∂x ∂y ∇U = ∂u y ∂u y ? ∂x ∂y At least three points are required to fully describe all of these potential motions in the neighborhood of a material point. Rigid-body rotations alter the displacement gradient component values, and therefore a rotation-invariant measure of deformation is generally preferred. One such measure is the Lagrangian strain tensor, E, computed as 1 [(∇U)T × (∇U) {+} (∇U)T {+} (∇U)] 2 Chen et al (52) and Young & Hunter (48) incorporated both surface geometry data and coronary tree bifurcation points to model epicardial motion and strains. McEachen & Duncan (54) used points of curvature extrema similarly. These methods used the tracking of these identifiable surface points along with smoothing constraints to estimate surface point trajectories. For investigational use, Arts et al (55) implanted a total of 14 radiopaque surface markers around the heart and computed coefficients in a 14-parameter kinematic model of heart motion. Clinically, similar studies with transplanted hearts embedded with tantalum helices have been performed by Moon et al (56) and Hansen et al (57), yielding similar results. The sparseness of the material markers in these studies severely limited the accuracy and application of these methods for strain calculation (6), yet they provided novel shape and shape-change information (50, 58). In canine hearts, Hashima et al (59) achieved much greater material point density by sewing 25 lead beads into the LV free wall. The resulting finite element-derived, nonhomogeneous epicardial strains showed substantial alteration during occlusion of the left anterior descending (LAD) coronary artery, compared with the nonoccluded state. Nevertheless, the absence of material points within the myocardium limits the conclusions that can be drawn from such surface analyses. To compute a local 3-D displacement gradient tensor requires a minimum of four noncoplanar material points (60). The magnitude and rate of change of these deformations vary regionally around the heart and throughout the cardiac cycle. To accurately assess 3-D cardiac function, one must first be able to define and track a sufficient number of material points with adequate spatial and temporal resolution. Spatially, a complete description of the local strain requires a sampling interval that is sufficiently small to recover the smallest fluctuations in the motion field. Douglas et al (61) estimated, using spherical models and wall motion data from E= P1: FHY June 13, 2000 11:12 Annual Reviews AR106-16 Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. IMAGING 3-D CARDIAC FUNCTION 443 canine hearts, that a 3-mm sampling interval through the wall is needed to model transmural wall motion in a healthy subject. Less stringent sampling intervals of 5–6 mm are needed in the circumferential and longitudinal directions. However, in diseased hearts, in which strong gradients in motion may occur in the border zone between healthy and ischemic regions, a higher density of points may be needed. Temporally, some purposes require strain patterns at only a single time point, that is, from end-diastole to end-systole, to evaluate the strains in the fully contracted state. However, the evolution of strain may also be critical for clinical diagnosis (62). Strain evolution may give information on the pattern of contraction and reveal electrical conduction pathway abnormalities. This requires the acquisition of data on a time scale that is smaller than the fastest motions of interest. As suggested earlier, a temporal resolution limit of 50 ms seems reasonable. ? Implanted Markers One investigational method that has provided valuable research data is the embedding of arrays of radiopaque markers into the heart wall, which are then imaged by using biplanar cineradiography. Combining the projection data from two orthogonal views, one is able to reconstruct the 3-D location of each bead in the array at each time frame (7). The motion of three or more planar beads that are attached to the surface of the heart give an estimate of the local 2-D surface strain (63, 64). Additional markers at various depths in the heart wall provide the additional information necessary for a complete 3-D motion analysis (65–67). Beads are commonly arranged in stacks of planar triangles, ∼5 mm on a side, with 4 mm of separation between stacks. Imaging a density of beads becomes difficult, but arrays with 2- to 3-mm separations have been successfully reconstructed. Implanted arrays of sonomicrometers have been used similarly, although their use is generally limited to 2-D surface studies. The disadvantage of these approaches is the invasiveness of the procedure, which precludes its routine use on human subjects. The implantation surgery may damage the tissue, the chest wall and pericardium are often resected, and the mere presence of several 0.5- to 1-mm-diameter beads through a wall 10–15 mm thick may alter the normal mechanical deformations in the subject. Magnetic Resonance Imaging Tagging An MRI tag is a region of the tissue where the net magnetization has been altered with carefully designed radio frequency pulses (3, 68). Upon imaging, these altered regions show good signal contrast compared with neighboring untagged regions. Each tag is created as a 3-D plane that extends through the entire subject, and it is seen as a tag line when imaged in an orthogonal view. Various tag generation schemes have been invented for creating numerous tags in a short amount of time. The more popular tagging schemes include stacks of parallel lines (69, 70), grids (71), and radial stripes (72). Because the tags result from alterations of the magnetization of the tissue itself, the motion of the tags matches the motion of the underlying tissue (73, 74). P1: FHY June 13, 2000 Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. 444 11:12 Annual Reviews O’DELL ¥ AR106-16 MCCULLOCH Typically, tags are created at end-diastole when the cavity volumes are largest, and the image data are acquired at 30- to 50-ms intervals as the heart contracts. The altered magnetization persists for ∼0.5 s, based on the T1 magnetization relaxation constant of cardiac tissue. Therefore, 10–12 images can be acquired during the ejection or filling phase of the cardiac cycle. The segmentation of tags is far more easily automated than is the segmentation of the heart contours, because the tags are human-made features of the image, for which the location and image intensity profile can be well predicted (75). The optimal image intensity profile and in-plane separation of tag planes can be computed analytically and have been validated experimentally. For parallel tag data, a tag width of 1–2 pixels with a tag center-to-center separation of 6 pixels gives an optimal tag centerline detection accuracy (76), which, for a typical image SNR of 15, is <0.1 pixels (0.1 mm for a typical clinical cardiac image). The detection of the location of grid tag intersections is slightly less accurate [∼0.3 mm mean error (77)], owing to the absence of the tag profiles at the intersections. Parallel tagging schemes enable the acquisition of images with lower resolution in the direction that is parallel to the tag lines. This enables the acquisition time to be reduced, leading to images that exhibit less motion artifact. However, two sets of images are then needed to sample the in-plane 2-D motion. Tag displacement data in three orthogonal directions are required to reconstruct the 3-D trajectory of material points. A typical 3-D tagging data set consists of two sets of short-axis images (or one set with a tag grid), one with the tag lines oriented 90◦ from the other set, and a set of long-axis images with tag planes that are parallel to the short-axis imaging planes. ? Strain Rate Analysis It is also possible with certain imaging modalities to measure directly the rate of change of strain. As described above, the tracking of material points leads to the computation of the local Lagrangian strain tensor, which is the description of motion around a given point in the tissue as it traverses through space and time. An alternative way to describe the deformation is to consider the relative velocity of motion at a particular location in space as a function of time. The local gradient of the velocity field at a given spatial location leads to the computation of the Eulerian strain rate tensor, which is analogous to the above computation of the strain tensor from the displacement gradient (78). In 1-D analysis, the velocity information can also be integrated over time to obtain the trajectory of material points and the local strain tensor. Magnetic Resonance Phase-Contrast Imaging MR phase-contrast imaging can be used to directly measure the velocity of the myocardial tissue on a pixel-by-pixel basis (79). In a phase-contrast image, the change in phase of the net magnetization inside each pixel, compared with a reference image, correlates to the velocity of the underlying tissue in the direction that is parallel to the applied magnetic field P1: FHY June 13, 2000 11:12 Annual Reviews AR106-16 IMAGING 3-D CARDIAC FUNCTION 445 Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. gradient. This method has the advantage over tagging in that each pixel provides a unique motion measurement, and MR phase-contrast imaging has great potential for reconstructing 3-D motion (80). However, current implementation of phasecontrast imaging for 3-D motion measurement is hampered by low SNRs, high susceptibility to motion artifacts, and, hence, limited accuracy of the reconstructed velocity profiles. ? Tissue Doppler Imaging The Doppler shift of the reflected echocardiographic wave also contains velocity information that is analogous to that obtained with MR phase-contrast imaging. Doppler ultrasound provides real-time velocity (81) and velocity gradient (82) information about blood and tissue. The frequency shift spectrum vs time information, presented as a sonogram, reveals the velocity range and distribution. The additional motion information from TDI has been shown to improve, over transthoracic ECG, detection of anomalous conduction pathways in patients with Wolff-Parkinson-White syndrome (83). However, velocity information is obtained only in the direction parallel to the echo beam and CNR noise (Doppler speckle), and the number of available viewing angles limit the image quality. Tessler et al (84), using a calibrated 1-D flow phantom, found that the Doppler-measured velocity had a baseline error of 6.8 cm/s (9%–17%) for peak flows that range from 40.5 to 75.3 cm/s. This error was independent of inter- or intraobserver variability or variability that is attributable to different flow probes or the flow phantom. Fleming et al (82) also used a test phantom and showed that 3 mm by 3 mm was the smallest region of tissue from which a measurable change in velocity could be detected with Doppler imaging. Velocity vector mapping in two and three dimensions is theoretically possible by using additional ECG receivers or by electronically separating a linear array receiver into spatially separate subapertures (85, 86). The unidirectional velocity magnitudes and the angle difference between the subapertures and the source echo are used to derive the direction and true magnitude of the velocity. Slice Motion Correction Accounting for the full 3-D motion of the heart, particularly the motion through the fixed short-axis image plane locations, is crucial for 3-D deformation analysis. The approaches mentioned above can accommodate this need by acquiring 3-D tagging or velocity data sets. However, alternatives, such as the slice isolation presented by Rogers et al (41), can also be used. A related technique involving subtraction of positive and negative tagged images has been shown by Fischer et al (87). In another approach, WeDeen et al (88) implemented a motionless movie scheme, in which the same slice was imaged at the same time in the cardiac cycle (to be certain that the same chunk of tissue was always imaged), but the tagging grid (or phase-encoding step) was imparted to the tissue at a sequence of preceding time increments. From this they were able to compute the 2-D Eulerian strain (or strain rate) at that slice location. Hybrid tagging/ P1: FHY June 13, 2000 446 11:12 Annual Reviews O’DELL ¥ AR106-16 MCCULLOCH velocity-encoding methods are inherently difficult because the tagged regions of tissue contain no velocity information (are a signal void), and the velocity encoding step requires a reference and a phase-encoding image; thus it does not offer an overall time advantage compared with 3-D tagging or 3-D phase-contrast imaging. Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. Model-Based Deformation Analysis ? Reconstruction of the 3-D strain field from discrete samples of displacement requires an interpolation scheme to describe the motion between samples in space, to correct for noise in the sampling, and to compute the local displacement gradients that are used for strain calculation (unless explicitly stated, considerations for velocity field and strain rate reconstruction can be assumed to be analogous to those described here for displacement field and strain reconstruction). The interpolation method usually includes a model of the 3-D displacement field. A priori assumptions of the displacement field can be used to generate a model that contains the expected modes of motion, and the displacement or velocity samples can be used to optimize the magnitude and direction of those modes (55). A more general formulation is to use 3-D basis functions to describe the local motion within each element of a finite element model of the heart (89), increasing the number of degrees of freedom in the reconstruction to the number of local basis functions per element times the number of elements. Taking this globally, a model containing hundreds of candidate modes can be formulated, and the sampled motion can be used to solve for the magnitude of those modes over the entire heart. This removes the requirement for a priori assumptions about the modes of motion, useful for reconstructing motions in regions of altered mechanical state, such as sites of ischemia. O’Dell et al (53) use a displacement field fitted to a polynomial series in prolate spheroidal coordinates (a 3-D analogy to the 2-D surface reconstruction presented earlier) to reconstruct 3-D material point trajectories from parallel-tag data sets. As shown in Figure 2 (see color insert), the contribution of each mode to the overall motion of the heart can be quantified and interpreted independently by using computer models of the heart (90). Another interesting approach, presented by Denney et al (91), uses samples of motion along with energy minimization equations that penalize large gradients in deformation to estimate the motion at each point in a rectangular grid that encompasses the entire heart. This approach is related to optical-flow techniques and is a powerful model-free method of computational analysis. Limitations of the Reconstructions Ultimately, the resolution and accuracy of the results from a reconstruction scheme are determined by the deformations of the heart tissue and the density and quality of original motion sampling. First, if the motion is simple, for example, the translation of a solid body, only a few simple parameters and a few samples of motions P1: FhN/FGI Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. July 31, 2000 P2: FPX/FOK QC: FHN/fgm 4:55 Annual Reviews T1: FhN AR106-01 ? Figure 2 Top six deformation modes from high-density healthy human left ventricular data set, demonstrated on a computer-generated model. λ is the radial coordinate, φ is the longitudinalangle–coordinate, and θ is the circumferential angle. (1) ∂φ = constant; longitudinal contraction of the base of the heart toward apex. (2) ∂λ = constant; concentric contraction without wall thickening. (3) ∂θ = cos(φ); clockwise twist of the apex with respect to the base. (4) ∂θ = sin(θ)sin(φ); circumferential contraction toward lateral wall. (5) ∂φ = cos(φ); longitudinal contraction toward the equator. (6) ∂λ = sin(φ)cos(θ ); bulging of the lateral free-wall. P1: FHY June 13, 2000 11:12 Annual Reviews AR106-16 Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. IMAGING 3-D CARDIAC FUNCTION 447 are needed. However, for the complex, 3-D heart wall motion, with high-order transmural gradients in the displacement field and spatial heterogeneity (i.e. from septum to free wall and base to apex), many global degrees of freedom are required to describe the motion around a given material point. To describe the trajectory and strain environment of a given material point (e.g. to within 0.3-mm 3-D tracking accuracy) requires many parameters in a local deformation model or perhaps >100 in a global model. However, the resolution and accuracy one actually needs for a given purpose may be considerably less stringent. Models based on a priori assumptions and a few simple expected modes may give sufficient information in an efficient manner. Upon successful image segmentation, a typical MRI parallel-tagging data set generates 3000–10,000 samples of 1-D displacement. Points along tag lines are obtained typically at 1- or 2-mm intervals. In the clinical setting, the separation between tags is typically 6–7 mm, the distance between contiguous image slices is often 7–10 mm, and hence the 3-mm sampling criterion estimated by Douglas et al (61) is not met in all directions. Experimental conditions, however, can afford greater image quality and spatial resolution and can overcome these limitations. ? REGIONAL STRAINS IN DISEASE The ultimate clinical goal of 3-D cardiac deformation analysis is to quantify and characterize regions of altered mechanical function. Figure 3 (see color insert) compares subendocardial and subepicardial circumferential strains at end-systole (referenced to end-diastole) in a patient with LV anterior-wall dysfunction that is secondary to ischemia. This region is typically supplied by the LAD coronary artery; therefore, we can postulate that there is a critical occlusion of that artery. In the central ischemic zone, the circumferential strain is positive, indicating that the region is undergoing stretch or bulging in response to the contraction of the neighboring healthy tissue and the imposed cavity pressure. For clinical application, the primary concern is whether the tissue is viable. If the tissue were necrotic, then an aggressive and expensive treatment paradigm, e.g. open-chest coronary graft surgery, would not be beneficial. However, if it can be shown that the tissue is yet viable, then restoration of normal circulation will presumably restore normal cardiac function. The secondary issue is to quantify the degree of dysfunction. A patient exhibiting only a 20% loss of contractile function can perhaps be treated with a less aggressive, less expensive pharmacologic regimen, whereas a patient with an 80% loss of regional function will perhaps need more immediate aggressive therapy, such as balloon angioplasty, to restore vascular patency. Dobutamine or adenosine stress testing has been suggested for noninvasive detection of viable but noncontracting (hibernating) myocardium in conjunction with MRI motion tracking. Croisille et al (92) found improved function in response to P1: FhN/FGI Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. July 31, 2000 P2: FPX/FOK QC: FHN/fgm 4:55 Annual Reviews T1: FhN AR106-01 ? Figure 3 Three-dimensional rendering of end-systolic circumferential strain derived from magnetic resonance imaging tagging in a patient with left anterior descending coronary arteryassociated ischemia. Sub-endocardial (left) and sub-epicardial (right) strains are shown. Positive strain (stretching) is indicated by dark blue, and negative strain is rendered in bright yellow. Range, 0.2 to −0.2. MRI data provided by E McVeigh, Johns Hopkins University School of Medicine. P1: FHY June 13, 2000 448 11:12 Annual Reviews O’DELL ¥ AR106-16 MCCULLOCH inotropic stimulation in at-risk regions of canine hearts subject to LAD occlusion and reperfusion. Regions of unchanged function after dobutamine infusion were shown to correlate with areas of infarction as evidenced by TTC staining. Hopefully, conclusive clinical test results will soon be forthcoming. Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. MEASUREMENT OF THREE-DIMENSIONAL TISSUE ARCHITECTURE ? The heart has a complex 3-D anatomy that plays a significant role in the generation of 3-D wall deformation (93). There exists a predominant fiber direction that is nearly completely confined to a circumferential-longitudinal plane, with an in-plane angle with respect to the heart long-axis that varies nearly linearly with wall depth. However, conflicting theories remain as to the tertiary structure of the fibers. The dissections and histological findings of LeGrice et al (94) suggest a laminar structure of the fiber bundles, with loose collagen connections between bundles of 4–5 fibrils. They found that the laminar or sheet arrangement is consistent among canine subjects and hypothesized that the sheets facilitate large wall shears and increased wall thickening during contraction. A dilemma for any anatomical study is that intervention distorts the underlying 3-D fiber structure, and, hence, conclusions about the intact structure are suspect. Yet, dissection and histological sectioning remain the gold standards for elucidating the 3-D fiber architecture. Considerable data on fiber structure in canines have been presented by Streeter & Hanna (93) and LeGrice et al (94), but these findings do not necessarily apply to the human heart. Owing to difficulties in obtaining suitable human hearts and in performing the histological sectioning, detailed 3-D fiber structure analysis for a human heart has yet to be performed. A recent application of MR phase-sensitive imaging is now finding use for measurement of myocardial fiber architecture. MR diffusion-sensitive imaging uses phase-contrast imaging techniques, but these are tailored to motions on the scale of free-water diffusion. The tissue sample is spatially marked by applying a magnetic field gradient for a short period of time, creating a linear dispersion of the phase of the tissue magnetization across the sample along the direction of the applied gradient field. Nuclei that subsequently move will carry their phase information to the new location. A refocusing gradient then resets the stationary spins to their original phase state. In a population of nuclei, random motion that is parallel to the applied gradient field will cause a dispersion of phase around the stationary-phase value and a corresponding decrease in pixel intensity that varies exponentially with the square of the magnitude of the applied gradient (assuming that restricted diffusion effects can be ignored). The coefficient of that exponential is the diffusion coefficient. The longer the average displacement of the population, the greater the spread in the phase, the greater the signal attenuation, and the larger the diffusion coefficient. P1: FHY June 13, 2000 11:12 Annual Reviews AR106-16 Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. IMAGING 3-D CARDIAC FUNCTION 449 For many biological tissues, the diffusion coefficient is different for different directions. In three dimensions, this is described fully by the local diffusion tensor. Phase dispersion mapping with measured diffusion coefficients in six nonparallel directions (the diffusion tensor is symmetric) can be used to estimate the local 3-D diffusion tensor (95). From a tissue standpoint, the free diffusion of water is hypothesized to occur more readily in a direction more parallel to the local fibers than perpendicular. Studies by Hsu et al (96) in dogs and Scollan et al (97) in rabbits have shown that the principal direction of maximal diffusion is related to the local myocardial fiber direction, with a 12◦ uncertainty, using histological sectioning as a guide. Scollan et al (97) go on to describe a qualitative relationship between the laminar fiber architecture and the remaining diffusion eigen components. Reese et al (98) have applied such techniques to in vivo canine and human subjects. With this technology, it should be possible to noninvasively measure the 3-D fiber architecture in the same hearts that are used for mechanical testing. This will be especially enlightening in certain disease states where interindividual variation in fiber architecture can be large and unpredictable. ? FUTURE DIRECTIONS The 3-D nature of heart deformation is dependent on the 3-D ventricular geometry, regional myocardial strain, 3-D fiber tissue architecture, and internal wall stresses. We have shown examples of promising techniques to independently assess heart anatomy, regional strains, and fiber architecture. The additional measurements of myocardial stresses would enable analysis of the myocardial work output, which relates to the metabolic requirements of the tissue, including oxygen consumption (99). It is not yet possible to directly measure myocardial stresses in vivo without destroying the tissue or significantly altering the measurement (100). However, it has been shown that pulsed Doppler ultrasound (101) can provide an estimate of tissue stiffness because the speed of propagation (and hence wavelength) of the ultrasonic beam varies with stiffness. This approach, however, suffers from the typical limitations of TDI. The promising method of MR elastography, introduced by Muthupillai et al (102), combines ultrasonic oscillating mechanical surface stress waves (at 10–1100 Hz) with MR phase-sensitive imaging to monitor the propagation of the ultrasonic beam through tissue, with increased sensitivity and clarity compared with Doppler ultrasound. These techniques have yet to be successfully applied to the in vivo heart. The approach that has seen some success is to estimate the stress-strain behavior of heart tissue by computational modeling (103). It is conceivable to use an inverse approach to fit constitutive law parameters from measured strains. Moulton et al (104) have shown the potential of such an approach by using a simplified constitutive law and finite element modeling of a 2-D cross section of tissue. The finite element model was loaded with approximated 2-D surface tractions and allowed P1: FHY June 13, 2000 Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. 450 11:12 Annual Reviews O’DELL ¥ AR106-16 MCCULLOCH to deform by the laws of equilibrium. The error was computed between the finite element-predicted strain field and that measured with an MRI tagging grid over that slice, and the constitutive law parameters were varied to achieve a minimal strain error. This approach is analogous to that used by Guccione et al (105), who used an axisymmetric 3-D finite-element model and transversely isotropic constitutive law to match 3-D strains measured at a single site, using radiopaque markers and cineradiography. When fully 3-D geometric representations are considered, including fiber anatomy and material anisotropy, compressibility, and heterogeneity, this approach still involves substantial computational time. It is also conceivable to design an inverse approach based on matching measured surface tractions and stress equilibrium constraints (106), but this is highly susceptible to uncertainties in the strain field and all of the 3-D properties. Just as new modalities for imaging cardiac structure and function are aiding the computational estimation of myocardial stress and work, a priori knowledge of myocardial mechanics can be used to improve image segmentation and analysis. Duncan et al (107) described the general theory in a recent review, and it could be applied directly to the finite element-based strain reconstruction methods for MRI tagging data by Kraitchman & Young (77) and Young & Axel (89). Apart from improvement in imaging individual components of 3-D cardiac deformation, the assessment of multiple components during a single imaging session (and within a reasonable time frame) is likely to become of increasing interest. Very recently, WeDeen et al (108) have shown preliminary data for measuring both myocardial strains and fiber direction in hearts from human volunteers and patients using MRI phase-sensitive imaging and MR diffusion tensor imaging. Despite the capabilities available currently, the use of 3-D cardiac functional analysis in the clinical setting remains limited. As outlined by the National Institutes of Health Cardiac MRI Work Group (109), the principal reasons for this include long imaging times (compared with single-plane/projection imaging), timeconsuming and labor-intensive post-processing requirements (primarily in image segmentation), and lack of understanding of benefits by clinicians. To this list can be added the difficulty in reducing and presenting the resulting voluminous and complex 3-D data sets into simple, informative parameters that can be readily used by the clinician. The challenges are for commercial, academic, and government agencies to improve the efficiency of 3-D image acquisition, to bring near real-time 3-D display of the processed image data into the examination room, and to perform the necessary clinical studies to demonstrate the capabilities of 3-D cardiac functional imaging. ? ACKNOWLEDGMENTS The authors acknowledge the many informative discussions and contributions from Paul Friedman, Sabrina Au, and Elliot McVeigh, and funding support from the AHA and NIH grant HL41603. W. G. O. is supported by an American Heart Association Post-Doctoral Fellowship. P1: FHY June 13, 2000 11:12 Annual Reviews AR106-16 IMAGING 3-D CARDIAC FUNCTION 451 Visit the Annual Reviews home page at www.AnnualReviews.org Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. LITERATURE CITED 1. Lamb HJ, Singleton RR, Vandergeest RJ, Pohost GM, Deroos A. 1995. MR imaging of regional cardiac function—low-pass filtering of wall thickness curves. Magn. Reson. Med. 34:498–502 2. Levy D, Murabito J, Anderson K, Christiansen J, Castelli W. 1992. Echocardiographic left ventricular hypertrophy— clinical characteristics—the Framingham Heart Study. Clin. Exp. Hypertens. A 14: 85–97 3. Reichek N. 1999. MRI myocardial tagging. J. Magn. Reson. Imaging 10(5):734–40 4. Rackley C, Dodge H, Coble Y, Hay R. 1964. A method for determining left ventricular mass in man. Circulation 29:666– 71 5. Waldman LK, McCulloch AD. 1993. Nonhomogeneous ventricular wall strain— analysis of errors and accuracy. J. Biomechan. Eng. 115:497–502 6. Hunter W, Zerhouni E. 1989. Imaging distinct points in the left ventricular myocardium to study regional wall deformation. In Innovations in Diagnostic Radiology, ed. JH Anderson, pp. 169–90. New York: Springer-Verlag 7. Garrison J, Ebert W, Jenkins R, Yionoulis S, Malcom H, et al. 1982. Measurement of three-dimensional positions and motions of large numbers of spherical radiopaque markers from biplane cineradiograms. Comput. Biomed. Res. 15:76–96 8. Clements I, Sinak L, Gibbons R, Brown M, O’Connor M. 1990. Determination of diastolic function by radionuclide ventriculography. Mayo Clin. Proc. 65:1007–19 9. Moore M, Murphy P, Burdine J. 1980. ECG-gated emission computed tomography of the cardiac blood pool. Radiology 134:233–35 10. Fischman A, Moore R, Gill J, Strauss H. 1989. Gated blood pool tomography: a 11. technology whose time has come. Semin. Nucl. Med. 19:13–21 Ofili E, Nanda N. 1994. Three-dimensional and four-dimensional echocardiography. Ultrasound Med. Biol. 20:669–75 DeCastro S, Yao J, Pandian N. 1998. Threedimensional echocardiography: clinical relevance and application. Am. J. Cardiol. 81:G96–102 von Ramm O, Smith S. 1990. Real-time volumetric ultrasound imaging system. J. Digit. Imaging 3:261–66 Gopal A, Shen Z, Sapin P, Keller A, Schnellbaecher M, et al. 1995. Assessment of cardiac function by three-dimensional echocardiography compared with conventional noninvasive methods. Circulation 92:842–53 Ghosh A, Nanda N, Maurer G. 1982. Three-dimensional reconstruction of echocardiographic images using the rotation method. Ultrasound Med. Biol. 8:655–61 Shiota T, Jones M, Chikada M, Fleishman C, Castellucci J, et al. 1998. Realtime three-dimensional echocardiography for determining right ventricular stroke volume in an animal model of chronic right ventricular volume overload. Circulation 97:1897–900 Lange A, Palka P, Caso P, Fenn L, Olszewski R, et al. 1997. Doppler myocardial imaging vs. B-mode grey-scale imaging: a comparative in vitro and in vivo study into their relative efficacy in endocardial boundary detection. Ultrasound Med. Biol. 23:69–75 Ritman E, Kinsey J, Robb R, Gilbert B, Harris L, Wood E. 1980. Threedimensional imaging of heart, lungs, and circulation. Science 210:273–80 Hsu T. 1981. Ultra-high-speed tomography: scanning electron beam CT systems. Appl. Radiol. 10:86–88 ? 12. 13. 14. 15. 16. 17. 18. 19. P1: FHY June 13, 2000 Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. 452 11:12 Annual Reviews O’DELL ¥ AR106-16 MCCULLOCH 20. McCollough C, Kanal K, Lannutti N, Ryan K. 1999. Experimental determination of section sensitivity profiles and image noise in electron beam computed tomography. Med. Phys. 26:287–95 21. Woodhouse C, Tanowitz W, Viamonte M. 1997. Retrospective cardiac gating technique to reduce cardiac motion artifact at spiral CT. Radiology 204:566–69 22. Sinak L, Hoffman E, Schwartz R, Smith H, Holmes DJ, et al. 1985. Three-dimensional cardiac anatomy and function in heart disease in adults: initial results with the dynamic spatial reconstructor. Mayo Clin. Proc. 60:383–92 23. McVeigh E, Atalar E. 1993. Balancing Contrast, Resolution and Signal-to-Noise Ratio. The Physics of MRI, ed. MJ Bronskill, P Sprawls. New York: Am. Assoc. Physicists Med. 24. Pettigrew R, Oshinski J, Chatzimavroudis G, Dixon W. 1999. MRI techniques for cardiovascular imaging. J. Magn. Reson. Imaging 10(5):590–601 25. McVeigh E. 1996. MRI of myocardial function: motion tracking techniques. Magn. Reson. Imaging 14:137–50 26. D’Cruz IA. 1996. Echocardiographic Anatomy: Understanding Normal and Abnormal Echocardiograms, pp. 312–34. Stamford, CT: Appleton & Lange 27. Lieberman A, Weiss J, Jugdutt B, Becker L, Bulkley B, et al. 1981. Twodimensional echocardiography and infarct size: relationship of regional wall motion and thickening to the extent of myocardial infarction in the dog. Circulation 63:739–46 27a. Pham DL, Xu C, Prince JL. 2000. Current methods in medical image segmentation. Annu. Rev. Biomed. Eng. 2:315–37 28. Mirsky I, Pasipoularides A. 1990. Clinical assessment of diastolic performance. Prog. Cardiovasc. Dis. V32(N4)291– 318 29. Jakob M, Hess O, Jenni R, Heywood J, Grimm J, Krayenbuehl H. 1991. Deter- 30. mination of left ventricular systolic wall thickness by digital subtraction angiography. Eur. Heart J. 12:573–82 Hexeberg E, Homans D, Bache R. 1995. Interpretation of systolic wall thickening. Can thickening of a discrete layer reflect fibre performance? Cardiovasc. Res. 29: 16–21 Lawson M, Johnson L, Coghlan L, Alami M, Tauxe E, et al. 1997. Correlation of thallium uptake with left ventricular wall thickness by cine magnetic resonance imaging in patients with acute and healed myocardial infarcts. Am. J. Cardiol. 80:434–41 Perrone-Filardi P, Bacharach S, Dilsizian V, Maurea S, Marinneto J, et al. 1992. Metabolic evidence of viable myocardium in regions with reduced wall thickness and absent wall thickening in patients with chronic ischemic left ventricular dysfunction. J. Am. Coll. Cardiol. 20:161–68 Curiel R, Laurienzo J, Unger E, Panza J. 1997. The magnitude of inotropic reserve is unrelated to basal systolic function or wall thickness in patients with chronic ischemic left ventricular dysfunction. Am. J. Cardiol. 80:783–86 Heupler S, Lauer M, Williams M, Shan K, Marwick T. 1997. Increased left ventricular cavity size, not wall thickness, potentiates myocardial ischemia. Am. Heart J. 133:691–97 Dong S, Macgregor J, Crawley A, McVeigh E, Belenkie I, et al. 1994. Left ventricular wall thickness and regional systolic function in patients with hypertrophic cardiomyopathy—a threedimensional tagged magnetic resonance imaging study. Circulation 90:1200–9 Pouleur H, Konstam M, Udelson J, Rousseau M. 1993. Changes in ventricular volume, wall thickness and wall stress during progression of left ventricular dysfunction. J. Am. Coll. Cardiol. 22:A43– A48 ? 31. 32. 33. 34. 35. 36. P1: FHY June 13, 2000 11:12 Annual Reviews AR106-16 Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. IMAGING 3-D CARDIAC FUNCTION 37. McCulloch A, Sung D, Wilson J, Pavelec R, Omens J. 1998. Flow-function relations during graded coronary occlusions in the dog: effects of transmural location and segment orientation. Cardiovasc. Res. 37: 636–45 38. Bazille A, Guttman M, McVeigh E, Zerhouni E. 1994. Impact of semi-automated versus manual image segmentation errors on myocardial strain calculation by MR tagging. Invest. Radiol. 29:427–33 39. Levett J, Replogle R. 1979. Thermodilution cardiac output: a critical analysis and review of the literature. J. Surg. Res. 27(6): 392–404 40. Dodge H. 1971. Determination of left ventricular volume and mass. Radiol. Clin. N. Am. 9(3):459–67 41. Rogers W, Shapiro E, Weiss J, Buchalter M, Rademakers F, et al. 1991. Quantification of and correction for left ventricular systolic long-axis shortening by magnetic resonance tissue tagging and slice isolation. Circulation 84:721–31 42. Carr K, Engler R, Forsythe J, Johnson A, Gosink B. 1979. Measurement of left ventricular ejection fraction by mechanical cross-sectional echocardiography. Circulation 59(6):1196–206 43. Lawson M, Blackwell G, Davis N, Roney M, Dell’Italia L, Pohost G. 1996. Accuracy of biplane long-axis left ventricular volume determined by cine magnetic resonance imaging in patients with regional and global dysfunction. Am. J. Cardiol. 77:1098–104 44. Mogelvang J, Tokholm K, Saunamaki K, Reimer A, Stubgaard M, et al. 1992. Assessment of left ventricular volumes by magnetic resonance in comparison with radionuclide angiography, contrast angiography and echocardiography. Eur. Heart J. 13:1677–83 45. Amini A, Weymouth T, Jain R. 1990. Using dynamic programming for solving variational problems in vision. IEEE Trans. Pattern Anal. Mach. Intell. 12(9):855–67 453 46. Ranganath S. 1995. Contour extraction from cardiac MRI studies using snakes. IEEE Trans. Med. Imaging 14:328–38 47. Costa K, Hunter P, Wayne J, Waldman L, Guccione J, McCulloch A. 1996. A three-dimensional finite element method for large elastic deformations of ventricular myocardium. II. Prolate spheroidal coordinates. ASME J. Biomech. Eng. 118:464– 72 48. Young A, Hunter P. 1989. Epicardial surface estimation from coronary angiograms. Comput. Vis. Graph. Image Proc. 47:111– 27 49. Cohen L, Cohen I. 1993. Finite-element methods for active contour models and balloons for 2-D and 3-D images. IEEE Trans. Pattern Anal. Mach. Intell. 15:1131–47 50. Duncan J, Lee F, Smeulders A, Zaret B. 1991. A bending energy model for measurement of cardiac shape deformity. IEEE Trans. Med. Imaging 10(3):307–20 51. McInerney T, Terzopoulos D. 1995. A dynamic finite element surface model for segmentation and tracking in multidimensional medical images with application to cardiac 4D image analysis. Comput. Med. Imaging Graph. 19:69–83 52. Chen C, Huang T, Arrott M. 1994. Modeling, analysis, and visualization of left ventricle shape and motion by hierarchical decomposition. IEEE Trans. Pattern Anal. Mach. Intell. 16(4):342–56 53. O’Dell WG, Moore CC, Hunter WC, Zerhourni EA, McVeigh ER. 1995. Threedimensional myocardial deformations: calculation with displacement field fitting to tagged MR images. Radiology 195:829– 35 54. McEachen J, Duncan J. 1997. Shape-based tracking of left ventricular wall motion. IEEE Trans. Med. Imaging 16(3):270–83 55. Arts T, Hunter W, Douglas A, Muijtjens A, Reneman R, Corday E. 1992. Description of the deformation of the left ventricle by a kinematic model. J. Biomech. 25(10): 1119–27 ? P1: FHY June 13, 2000 Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. 454 11:12 Annual Reviews O’DELL ¥ AR106-16 MCCULLOCH 56. Moon M, Ingels N, Daughters G, Stinson B, Hansen D, Miller D. 1994. Alterations in left ventricular twist mechanics with inotropic stimulation and volume loading in human subjects. Circulation 89:142–50 57. Hansen D, Daughters G, Alderman E, Ingels NJ, Miller D. 1988. Torsional deformation of the left ventricular midwall in human hearts with intramyocardial markers: regional heterogeneity and sensitivity to the inotropic effects of abrupt rate changes. Circulation Res. 62:941–52 58. Walley K, Grover M, Raff G, Benge J, Hannaford B, Glantz S. 1982. Left ventricular dynamic geometry in the intact and open chest dog. Circulation Res. 50:573–89 59. Hashima A, Young A, McCulloch A, Waldman L. 1993. Nonhomogeneous analysis of epicardial strain distributions during acute myocardial ischemia in the dog. J. Biomech. 26:19–35 60. Smaill B, Hunter P. 1991. Material properties of passive myocardium. In Theory of Heath: Biomechanics, Biophysics, and Nonlinear Dynamics of Cardiac Function, pp. 1–30. New York: Springer-Verlag 61. Douglas A, Hunter W, Wiseman M. 1990. Inhomogeneous deformation as a source of error in strain measurements derived from implanted markers in the canine left ventricle. J. Biomech. 23:331–41 62. Lugo-Olivieri C, Moore C, Poon E, Lima J, McVeigh E, Zerhouni E. 1994. Temporal evolution of three dimensional deformation in the ischemic human left ventricle: assessment by MR tagging. Proc. Soc. Magn. Reson. 1:1482 63. MacKey S, Potel M, Rubin J. 1982. Graphics methods for tracking three-dimensional heart wall motion. Comput. Biomed. Res. 15:455–73 64. McCulloch A, Smaill B, Hunter P. 1989. Regional left ventricular epicardial deformation in the passive dog heart. Circulation Res. 64:721–33 65. Douglas A, Rodriguez E, O’Dell W, Hunter W. 1991. Unique strain history during ejec- 66. 67. tion in canine left ventricle. Am. J. Physiol. Heart Circ. Physiol. 260:H1596–611 Waldman L, Fung Y, Covell J. 1985. Transmural myocardial deformation in the canine left ventricle. Circulation Res. 57: 152–63 Omens J, May K, McCulloch A. 1991. Transmural distribution of threedimensional strain in the isolated arrested canine left ventricle. Am. J. Physiol. Heart Circ. Physiol. 261:H918–28 Zerhouni E, Parish D, Rogers W, Yang A, Shapiro E. 1988. Human heart: tagging with MR imaging—a method for noninvasive assessment of myocardial motion. Radiology 169:59–63 Mosher T, Smith M. 1990. A DANTE tagging sequence for the evaluation of translational sample motion. Magn. Reson. Med. 15:334–39 Fischer S, McKinnon G, Maier S, Boesiger P. 1993. Improved myocardial tagging contrast. Magn. Reson. Med. 30:191–200 Axel L, Dougherty L. 1989. MR imaging of motion with spatial modulation of magnetization. Radiology 171:841–45 Bolster B, McVeigh E, Zerhouni E. 1990. Myocardial tagging in polar coordinates with striped tags. Radiology 177:769–72 Moore C, Reeder S, McVeigh E. 1994. Tagged MR imaging in a deforming phantom: photographic validation. Radiology 190:765–69 Young A, Axel L, Dougherty L, Bogen D, Parenteau C. 1993. Validation of tagging with MR imaging to estimate material deformation. Radiology 188:101–8 Guttman M, Prince J, McVeigh E. 1994. Tag and contour detection in tagged MR images of the left ventricle. IEEE Trans. Med. Imaging 13:74–88 Atalar E, McVeigh E. 1994. Optimum tag thickness for the measurement of motion by MRI. IEEE Trans. Med. Imaging 13: 152–60 Kraitchman D, Young A. 1995. Semiautomatic tracking of myocardial motion ? 68. 69. 70. 71. 72. 73. 74. 75. 76. 77. P1: FHY June 13, 2000 11:12 Annual Reviews AR106-16 IMAGING 3-D CARDIAC FUNCTION 78. 79. Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. 80. 81. 82. 83. 84. 85. 86. 87. 88. in MR tagged images. IEEE Trans. Med. Imaging 14:422–33 Spencer A. 1980. Continuous Mechanics. London/New York: Longman Pelc NJ, Dragnova M, Pelc LR. 1995. Tracking of cyclic motion with phasecontrast cine MR velocity data. J. Magn. Reson. Imaging 5:339–45 Drangova M, Zhu Y, Bowman B, Pelc N. 1998. In vitro validation of myocardial motion tracking from phase-contrast velocity data. Magn. Reson. Imaging 16:863– 70 McDicken W, Hoskins P, Moran C, Sutherland G. 1996. New technology in echocardiography. 1. Doppler techniques. Heart 75:2–8 Fleming A, Xia X, McDicken W, Sutherland G, Fenn L. 1994. Myocardial velocity gradients detected by Doppler imaging. Br. J. Radiol. 67:679–88 Fedele F, Trambaiolo P, Magni G, DeCastro S, Cacciotti L. 1998. New modalities of regional and global left ventricular function analysis: state of the art. Am. J. Cardiol. 81:49G–57G Tessler F, Kimmesmith C, Sutherland M, Schiller V, Perrella R, Grant E. 1990. Interobserver and intra-observer variability of Doppler peak velocity measurements— an in vitro study. Ultrasound Med. Biol. 16:653–57 Phillips P, Kadi A, von Ramm O. 1995. Feasibility study for a two-dimensional diagnostic ultrasound velocity mapping system. Ultrasound Med. Biol. 21:217–29 Picot P, Rickey D, Mitchell R, Rankin R, Fenster A. 1993. 3-Dimensional colour doppler imaging. Ultrasound Med. Biol. 19:95–104 Fischer S, McKinnon G, Scheidegger M, Prins W, Maier S, Boesiger P. 1994. True myocardial motion tracking. Magn. Reson. Med. 31:401–13 Wedeen V, Weisskoff R, Reese T, Beache G, Poncelet B, et al. 1995. Motionless movies of myocardial strain-rates using 89. 90. stimulated echoes. Magn. Reson. Med. 33: 401–8 Young A, Axel L. 1992. Three-dimensional motion and deformation of the heart wall: estimation with spatial modulation of magnetization—a model-based approach. Radiology 185(1):241–47 O’Dell WG, McVeigh ER. 1998. A modal description of LV motion during ejection. Proc. Int. Soc. Magn. Reson. Med. 2:876 Denney TJ, McVeigh E. 1997. Model-free reconstruction of three-dimensional myocardial strain from planar tagged MR images. J. Magn. Reson. 7(5):799–810 Croisille P, Moore C, Judd R, Lima J, Arai M, et al. 1999. Differentiation of viable and nonviable myocardium by the use of three-dimensional tagged MRI in 2-dayold reperfused canine infarcts. Circulation 99:284–91 Streeter D, Hanna W. 1973. Engineering mechanics for successive states in canine left ventricular myocardium. II: Fiber angle and sarcomere length. Circ. Res. 33:656– 64 LeGrice I, Smaill B, Chai L, Edgar S, Gavin J, Hunter P. 1995. Laminar structure of the heart: ventricular myocyte arrangement and connective tissue architecture in the dog. Am. J. Physiol. Heart Circ. Physiol. 38:H571–82 Basser P, Mattiello J, LeBihan D. 1994. Estimation of the effective self-diffusion tensor from the NMR spin echo. J. Magn. Reson. Ser. B 103:247–54 Hsu E, Muzikant A, Matulevicius S, Penland R, Henriquez C. 1998. Magnetic resonance myocardial fiber-orientation mapping with direct histological correlation. Am. J. Physiol. Heart Circ. Physiol. 43: H1627–34 Scollan D, Holmes, A, Winslow R, Forder J. 1998. Histological validation of myocardial microstructure obtained from diffusion tensor magnetic resonance imaging. Am. J. Physiol. Heart Circ. Physiol. 44:H2308– 18 ? 91. 92. 93. 94. 95. 96. 97. 455 P1: FHY June 13, 2000 Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. 456 11:12 Annual Reviews O’DELL ¥ AR106-16 MCCULLOCH 98. Reese T, Weisskoff R, Smith R, Rosen B, Dinsmore R, WeDeen V. 1995. Imaging myocardial fiber architechture in vivo with magnetic resonance. Magn. Reson. Med. 34:786–91 99. Suga H. 1990. Ventricular energetics. Physiol. Rev. 70:247–77 100. McCulloch A. 1995. In Cardiac Biomechanics, ed. JD Bronzino, pp. 418–439. Boca Raton, FL: CRC Press 101. Krouskop T, Doughtery D, Vinson F. 1987. A pulsed Doppler ultrasonic system for making non-invasive measurements of the mechanical properties of soft tissue. J. Rehabil. Res. Dev. 24:1–8 102. Muthupillai R, Lomas D, Rossman P, Greenleaf J, Manduca A, Ehman R. 1995. Magnetic resonance elastography by direct visualization of propagating acoustic strain waves. Science 269:1854– 57 103. McCulloch A, Waldman L, Rogers J, Guccione J. 1992. Large-scale finite element analysis of the beating heart. Crit. Rev. Biomed. Eng. 20:427–49 104. Moulton M, Creswell L, Downing S, Actis R, Szabo B, Pasque M. 1996. Myocardial material property determination in the in vivo heart using magnetic res- 105. 106. onance imaging. Int. J. Cardiol. Imaging 12:153–67 Guccione J, McCulloch A, Waldman L. 1991. Passive material properties of intact ventricular myocardium determined from a cylindrical model. ASME J. Biomech. Eng. 113:42–55 O’Dell WG, Hunter WC, McVeigh ER, McCulloch AD. 1997. Determination of material properties for the passively filling canine left ventricle. Ann. Biomed. Eng. 25:(Suppl. 1):S-27 Duncan J, Shi P, Constable T, Sinusas A. 1998. Physical and geometrical modeling for image-based recovery of left ventricular deformation. Prog. Biophys. Mol. Biol. 69:333–51 WeDeen V, Tseng W, Reese T, Kantor H, Weisskoff R. 1998. Magnetic resonance imaging finds myocardial fiber shortening and reduced cross-fiber shortening in hypertrophic cardiomyopathy. Circulation 98(17):I514 (Suppl.) Budinger TF, Berson A, McVeigh ER, Pettigrew RI, Pohost GM, et al. 1998. Cardiac MR imaging: report of a working group sponsored by the National Heart, Lung, and Blood Institute. Radiology 208:573–76 ? 107. 108. 109. Annual Review of Bimedical Engineering Volume 2, 2000 Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. CONTENTS PIERRE M. GALLETTI: A Personal Reflection, Robert M. Nerem 1 PHYSICOCHEMICAL FOUNDATIONS AND STRUCTURAL DESIGN OF HYDROGELS IN MEDICINE AND BIOLOGY, N. A. Peppas, Y. Huang, M. Torres-Lugo, J. H. Ward, J. Zhang 9 BIOENGINEERING MODELS OF CELL SIGNALING, Anand R. Asthagiri, Douglas A. Lauffenburger 31 FUNDAMENTALS OF IMPACT BIOMECHANICS: Part I Biomechanics of the Head, Neck, and Thorax, Albert I. King 55 INJURY AND REPAIR OF LIGAMENTS AND TENDONS, Savio L.-Y. Woo, Richard E. Debski, Jennifer Zeminski, Steven D. Abramowitch, Serena S. Chan Saw, MS, James A. Fenwick 83 ELECTROPHYSIOLOGICAL MODELING OF CARDIAC VENTRICULAR FUNCTION: From Cell to Organ, R. L. Winslow, D. F. Scollan, A. Holmes, C. K. Yung, J. Zhang, M. S. Jafri 119 CRYOSURGERY, Boris Rubinsky 157 CELL MECHANICS: Mechanical Response, Cell Adhesion, and Molecular Deformation, Cheng Zhu, Gang Bao, Ning Wang 189 MICROENGINEERING OF CELLULAR INTERACTIONS, Albert Folch, Mehmet Toner 227 QUANTITATIVE MEASUREMENT AND PREDICTION OF BIOPHYSICAL RESPONSE DURING FREEZING IN TISSUES, John C. Bischof 257 MICROFABRICATED MICRONEEDLES FOR GENE AND DRUG DELIVERY, Devin V. McAllister, Mark G. Allen, Mark R. Prausnitz 289 CURRENT METHODS IN MEDICAL IMAGE SEGMENTATION, Dzung L. Pham, Chenyang Xu, Jerry L. Prince 315 ANTIBODY ENGINEERING, Jennifer Maynard, George Georgiou 339 NEW CURRENTS IN ELECTRICAL STIMULATION OF EXCITABLE TISSUES, Peter J. Basser, Bradley J. Roth 377 TWO-PHOTON EXCITATION FLUORESCENCE MICROSCOPY, Peter T. C. So, Chen Y. Dong, Barry R. Masters, Keith M. Berland IMAGING THREE-DIMENSIONAL CARDIAC FUNCTION, W. G. O'Dell, A. D. McCulloch 399 431 THREE-DIMENSIONAL ULTRASOUND IMAGING, Aaron Fenster, Donal B. Downey 457 BIOPHYSICAL INJURY MECHANISMS IN ELECTRICAL SHOCK TRAUMA, Raphael C. Lee, Dajun Zhang, Jurgen Hannig 477 WAVELETS IN TEMPORAL AND SPATIAL PROCESSING OF BIOMEDICAL IMAGES, Andrew F. Laine 511 MICRODEVICES IN MEDICINE, Dennis L. Polla, Arthur G. Erdman, William P. Robbins, David T. Markus, Jorge Diaz-Diaz, Raed Rizq, Yunwoo Nam, Hui Tao Brickner, Amy Wang, Peter Krulevitch NEUROENGINEERING MODELS OF BRAIN DISEASE, Leif H. Finkel EXTRACORPOREAL TISSUE ENGINEERED LIVER-ASSIST DEVICES, Emmanouhl S. Tzanakakis, Donavon J. Hess, Timothy D. Sielaff, Wei-Shou Hu 551 577 607 Annu. Rev. Biomed. Eng. 2000.2:431-456. Downloaded from arjournals.annualreviews.org by UNIVERSITY OF ROCHESTER LIBRARY on 07/27/09. For personal use only. MAGNETIC RESONANCE STUDIES OF BRAIN FUNCTION AND NEUROCHEMISTRY, Kâmil Ugurbil, Gregor Adriany, Peter Andersen, Wei Chen, Rolf Gruetter, Xiaoping Hu, Hellmut Merkle, Dae-Shik Kim, Seong-Gi Kim, John Strupp, Xiao Hong Zhu, Seiji Ogawa 633 INTERVENTIONAL AND INTRAOPERATIVE MAGNETIC RESONANCE IMAGING, J. Kettenbach, D. F. Kacher, S. K. Koskinen, Stuart G. Silverman, A. Nabavi, Dave Gering, Clare M. C. Tempany, R. B. Schwartz, R. Kikinis, P. M. Black, F. A. Jolesz 661 CARTILAGE TISSUE REMODELING IN RESPONSE TO MECHANICAL FORCES, Alan J. Grodzinsky, Marc E. Levenston, Moonsoo Jin, Eliot H. Frank 691 IN VIVO NEAR-INFRARED SPECTROSCOPY, Peter Rolfe 715