* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download A Model for Measurement Error for Gene Expression Arrays
Perceptual control theory wikipedia , lookup
Inverse problem wikipedia , lookup
Generalized linear model wikipedia , lookup
Computer simulation wikipedia , lookup
Gene prediction wikipedia , lookup
Psychometrics wikipedia , lookup
Error detection and correction wikipedia , lookup
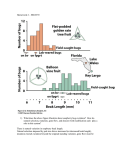
ECS 289A Presentation Jimin Ding • • • • • • • • Problem & Motivation Two-component Model Estimation for Parameters in above model Define low and high level gene expression Comparing expression levels Limitations of the model and method Other possible solutions References A Model for Measurement Error for Gene Expression Arrays David Rocke & Blythe Durbin Journal of Computational Biology Nov.2001 Problem & Motivation • Statistical inference for data need assumption of normality with constant variance --- So hypothesis testing for the difference between control and treatment need equal variance (not depending on the mean of the data); • Measurement error for gene expression rises proportionately to the expression level --- So linear regression fails and log transformation has been tried; • However, for genes whose expression level is low or entirely unexpressed, the measurement error doesn’t go down proportionately Example --- So log transformation fails by inflating the variance of observations near background, and two component model is introduced. Example: Mice From: Barosiewics etatl, 2000 From Durbin et.al 2002 back Two-Component Model y e • • • • Y is the intensity measurement is the expression level in arbitrary units is the mean intensity of unexpressed genes Error term: ~ N (0, ) ~ N (0, ) Estimation for background ( & ) y e ~ N (0, ), ~ N (0, ) • Estimation of background using negative controls • Estimation of background with replicate measurements Detail • Estimation of background without replicate Estimation of & with replicate measurements • Begin with a small subset of genes with low intensity (10%) ˆ xB ˆ S B m 1 2 si (ni 1) n m i 1 • Define a new subset consisting of genes whose intensity values are in [ xB 2sB , xB 2sB ] • Repeat the first and second steps until the set of genes does not change. Estimation of the High-level RSD • The variance of intensity in two-component model: , where • At high expression level, only multiple error term is noticeable, so the ratio of the variation to the mean is a constant, i.e. RSD= s • For each replicated gene that is at high level, compute the mean ̂ i of the y ̂ and the standard deviation si of log( y ˆ ) • Then use the pooled standard deviation to estimate : Define “high” and “low” 2 0.9 2 2 2 s / 3s • Low expression level: Most of the variance is due to the additive error component. 95% CI: (ˆ 1.96 Var(ˆ ) , ˆ 1.96 Var(ˆ ) ) 2 s2 0 .9 2 2 2 s 3 / s • High expression level: Most of the variance is due to the multiplicative error component. 95% CI: Comparing Expression Levels • Common method: standard t-test on ratio of expression for treatment and control (low level), or its logarithm (high level). • Problem: Less effective when gene is expressed at a low level in one condition and high in the other: Solution consider treatment and control are correlated • Model: • Variation: Background: High-level RSD: Hypothesis testing (Comparison) • • • Assume the data have been adjusted: Testing: (Gene has same expression level at Control and treatment) Then using the following approximate variance to do standard t-test for log ratio of raw data: Limitations • No theoretical result for above estimations. (Consistency and asymptotical distribution) • Cutoff point of high level and low level is fairly artificial • The convergence of estimation of background information is heavily dependent on data and initial selection Literature & Other Possible Solutions for Measurement Error • Chen et al. (1997): measurement error is normally distributed with constant coefficient of variation (CV)—in accord with experience • Ideker et al.(2000) introduce a multiplicative error component (normal) • Newton et al. (2001) propose a gamma model for measurement error. • Durbin et al.(2002) suggest transformation g ( y) ln[ y ( y ) 2 c ] , where c 2 / s2 • Huber et al.(2002) introduce transformation h( x) ar sinh( a bx) References • • • • Blythe Durbin, Johanna Hardin, Douglas Hawkins, and David Rocke. “A variancestabilizing transformation from gene-expression microarray data”, Bioinformatics, ISMB, 2002. Chen. Y., Dougherty, E.R. and Bittner, M.L.(1997) “Ratio-based decisions and the quantitative analysis of cDNA microarray images”, J.Biomed. Opt.,2,364374 Wolfgang Huber, Anja von Heydebreck,Martin Vingron (Dec.2002) “Analysis of microarray gene expression data”, Preprint Wolfgang Huber, Anja von Heydebreck, Holger S¨ultmann, Annemarie Poustka, and Martin Vingron. “Variance stablization applied to microarray data calibration and to the quantification of differential expression”, Bioinformatics, 18 Suppl. 1:S96–S104, 2002. ISMB 2002.