RNA PCR Kit (AMV)
... lead to pipetting errors. 3) Keep enzymes at -20℃ until just before use and return into the freezer promptly after use. 4) Use new disposable pipette tips to avoid contamination between samples. 5) PCR condition Optimum PCR condition varies depending on the thermal cycler used for PCR. It is re ...
... lead to pipetting errors. 3) Keep enzymes at -20℃ until just before use and return into the freezer promptly after use. 4) Use new disposable pipette tips to avoid contamination between samples. 5) PCR condition Optimum PCR condition varies depending on the thermal cycler used for PCR. It is re ...
Where is the root of the universal tree of life?
... The present crisis of molecular phylogeny will be overcome only if one recognises that the most important step in the analytical process is the search for good characters.(22) Since useful amino acid or nucleotidic positions are rare in sequence alignments, it will be important both to design new me ...
... The present crisis of molecular phylogeny will be overcome only if one recognises that the most important step in the analytical process is the search for good characters.(22) Since useful amino acid or nucleotidic positions are rare in sequence alignments, it will be important both to design new me ...
Nucleotide Sequence of the DNA Complementary to Avian (Chicken
... of the chicken hormone and the difference in sequence could be attributed to at least two deletion and insertion events. The first deletion involves residues 32-44 of the mammalian sequence, which is replaced by four amino acids in the chicken sequence (see Fig. 5). This occurs in the region where t ...
... of the chicken hormone and the difference in sequence could be attributed to at least two deletion and insertion events. The first deletion involves residues 32-44 of the mammalian sequence, which is replaced by four amino acids in the chicken sequence (see Fig. 5). This occurs in the region where t ...
Neema Bhukhan
... conserved non-coding transcript is located near the region of this gene suggests the possibility that it may have some functional contribution in the development of this gene. The evofold structure prediction with a score of 699 is located near the end of the same FOXP2 gene suggesting a similar rel ...
... conserved non-coding transcript is located near the region of this gene suggests the possibility that it may have some functional contribution in the development of this gene. The evofold structure prediction with a score of 699 is located near the end of the same FOXP2 gene suggesting a similar rel ...
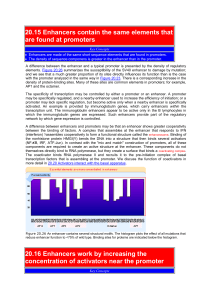
20.15 Enhancers contain the same elements that are
... the promoter (vicinity in this sense being a relative term). Two types of experiment illustrated in Figure 20.27 suggest that this is the case. A fragment of DNA that contains an enhancer at one end and a promoter at the other is not effectively transcribed, but the enhancer can stimulate transcript ...
... the promoter (vicinity in this sense being a relative term). Two types of experiment illustrated in Figure 20.27 suggest that this is the case. A fragment of DNA that contains an enhancer at one end and a promoter at the other is not effectively transcribed, but the enhancer can stimulate transcript ...
PCR
... Selection of an appropriate primer for reverse transcription depends on target mRNA size and the presence of secondary structure. For example, a primer that anneals specifically to the 3′-end of the transcript (a sequence-specific primer or oligo(dT) primer) may be problematic when reverse transcrib ...
... Selection of an appropriate primer for reverse transcription depends on target mRNA size and the presence of secondary structure. For example, a primer that anneals specifically to the 3′-end of the transcript (a sequence-specific primer or oligo(dT) primer) may be problematic when reverse transcrib ...
Three scientists who revealed the structure and workings of the
... was quick to laud the work of his long-time collaborator at Yale, Peter Moore. Ramakrishnan points to a series of lower resolution ‘snapshot’ structures produced by Harry Noller from the University of California, Santa Cruz; and the cryo-electron microscopy work of Joachim Frank from the Wadsworth c ...
... was quick to laud the work of his long-time collaborator at Yale, Peter Moore. Ramakrishnan points to a series of lower resolution ‘snapshot’ structures produced by Harry Noller from the University of California, Santa Cruz; and the cryo-electron microscopy work of Joachim Frank from the Wadsworth c ...
Locked Nucleic Acid - LNA™
... methylene bridge connecting the 2’-O atom with the 4’-C atom (see structure below). LNA™ nucleosides contain the six common nucleobases (T, C, G, A, U and mC) that appear in DNA and RNA and thus are able to form base-pairs according to standard Watson-Crick base pairing rules. Oligonucleotides incor ...
... methylene bridge connecting the 2’-O atom with the 4’-C atom (see structure below). LNA™ nucleosides contain the six common nucleobases (T, C, G, A, U and mC) that appear in DNA and RNA and thus are able to form base-pairs according to standard Watson-Crick base pairing rules. Oligonucleotides incor ...
Supplementary Information (doc 82K)
... from sucrose gradient fractions corresponding to RNPs (F1 to F2), 40S subunit (F3), 60S subunit (F4), 80S monosome (F5 and F6) and polysomes (F7 to F12), resolved on 10% urea-acrylamide gel and analyzed by northern blot hybridization with the 173 nt ss-DNA probe corresponding to nucleotides 41-213 o ...
... from sucrose gradient fractions corresponding to RNPs (F1 to F2), 40S subunit (F3), 60S subunit (F4), 80S monosome (F5 and F6) and polysomes (F7 to F12), resolved on 10% urea-acrylamide gel and analyzed by northern blot hybridization with the 173 nt ss-DNA probe corresponding to nucleotides 41-213 o ...
The tryptophan biosynthetic pathway
... trpL mRNA stalls at one of its two Trp codons. This permits the RNA antiterminator structure to form, which prevents formation of the terminator. Transcription then continues into the operon’s structural genes. An attenuator site, in effect, is a DNA sequence where a choice is made by RNA polymerase ...
... trpL mRNA stalls at one of its two Trp codons. This permits the RNA antiterminator structure to form, which prevents formation of the terminator. Transcription then continues into the operon’s structural genes. An attenuator site, in effect, is a DNA sequence where a choice is made by RNA polymerase ...
Computational Biology - Bioinformatik
... In addition to their role in the RNAi pathway, siRNAs also act in RNAi-related pathways, e.g., as an antiviral mechanism or in shaping the chromatin structure of a genome. ...
... In addition to their role in the RNAi pathway, siRNAs also act in RNAi-related pathways, e.g., as an antiviral mechanism or in shaping the chromatin structure of a genome. ...
Slide 1
... Summary of my argument Selection acts at the time of addition of new amino acids to the code. The new amino acid is assigned to codons that formerly coded for an amino acid with similar properties. This minimizes disruption to existing genes. ...
... Summary of my argument Selection acts at the time of addition of new amino acids to the code. The new amino acid is assigned to codons that formerly coded for an amino acid with similar properties. This minimizes disruption to existing genes. ...
Biochimica et Biophysica Acta
... 3.1. Preparation of the two-plasmid system A bacterial two-plasmid system has been previously described for studying RNA–protein interactions [21]. In this system, a first plasmid contains a modified lacZ reporter gene with an RNA-binding element located close to the translation initiation region. A s ...
... 3.1. Preparation of the two-plasmid system A bacterial two-plasmid system has been previously described for studying RNA–protein interactions [21]. In this system, a first plasmid contains a modified lacZ reporter gene with an RNA-binding element located close to the translation initiation region. A s ...
LIN-28 co-transcriptionally binds primary let
... Priscilla M Van Wynsberghe, Zoya S Kai, Katlin B Massirer, Victoria H Burton, Gene W Yeo & Amy E Pasquinelli Nature structural & molecular biology, VOLUME 18, 302-308, MARCH 2011 ...
... Priscilla M Van Wynsberghe, Zoya S Kai, Katlin B Massirer, Victoria H Burton, Gene W Yeo & Amy E Pasquinelli Nature structural & molecular biology, VOLUME 18, 302-308, MARCH 2011 ...
Ribosome stalls at trp codons, allowing 2+3 pairing Transcription
... •Contains complementary sequences that can form hairpin structures when transcribed into RNA ...
... •Contains complementary sequences that can form hairpin structures when transcribed into RNA ...
Activity: Invasion of the Snorks
... the base pairing rules are the same as us. 3. Code for the missing complementary strand of DNA for the Snork. 4. Getting back to the mRNA sample and using a codon chart, translate the mRNA into the amino acid sequence. Remember that AUG is always the start codon and it signifies the beginning of eac ...
... the base pairing rules are the same as us. 3. Code for the missing complementary strand of DNA for the Snork. 4. Getting back to the mRNA sample and using a codon chart, translate the mRNA into the amino acid sequence. Remember that AUG is always the start codon and it signifies the beginning of eac ...
Chapter 3. The Beginnings of Genomic Biology
... deriving from the phosphate groups making up the helices, positively charged ionic species within cells are attracted to these molecules. These positively charged molecules can be small ions such as K+ and Mg++, or they can be larger positively charged proteins, and/or other larger molecular species ...
... deriving from the phosphate groups making up the helices, positively charged ionic species within cells are attracted to these molecules. These positively charged molecules can be small ions such as K+ and Mg++, or they can be larger positively charged proteins, and/or other larger molecular species ...
Molecular Mechanisms of Long Noncoding RNAs
... Archetype I: as signals, lncRNA expression can faithfully reflect the combinatorial actions of transcription factors (colored ovals) or signaling pathways to indicate gene regulation in space and time. Archetype II: as decoys, lncRNAs can titrate transcription factors and other proteins away from ch ...
... Archetype I: as signals, lncRNA expression can faithfully reflect the combinatorial actions of transcription factors (colored ovals) or signaling pathways to indicate gene regulation in space and time. Archetype II: as decoys, lncRNAs can titrate transcription factors and other proteins away from ch ...
Molecular Biology of Transcription and RNA Processing
... molecular biology researchers focused on identifying and describing the molecules and mechanisms responsible for conveying the genetic message of DNA. RNA was known to be chemically similar to DNA and present in abundance in all cells, but its diversity and biological roles remained to be discovered ...
... molecular biology researchers focused on identifying and describing the molecules and mechanisms responsible for conveying the genetic message of DNA. RNA was known to be chemically similar to DNA and present in abundance in all cells, but its diversity and biological roles remained to be discovered ...
What are enzymes and how do they work
... termination and 2) how the structure of the release factor impacts its function. Key concepts: The release factor’s role is to catalyze the hydrolysis of the bond between the last tRNA and the completed protein. The release factor does not have an anticodon or carry an amino acid, but it is the same ...
... termination and 2) how the structure of the release factor impacts its function. Key concepts: The release factor’s role is to catalyze the hydrolysis of the bond between the last tRNA and the completed protein. The release factor does not have an anticodon or carry an amino acid, but it is the same ...
Slide 1
... Use RNAi to characterize regulatory function in protein secretion areA is a positively acting regulatory gene which has been shown to be essential for activating genes encoding enzymes, permeases, needed to acquire nitrogen for the environment areA has recently been shown in Aspergillus to play a p ...
... Use RNAi to characterize regulatory function in protein secretion areA is a positively acting regulatory gene which has been shown to be essential for activating genes encoding enzymes, permeases, needed to acquire nitrogen for the environment areA has recently been shown in Aspergillus to play a p ...
Stages of Translation (Biol 200 Sp2015): KEY Initiation
... ii. __ The large and small ribosomal subunits separate and fall off the mRNA___ 3. What is the sequence of the codon that indicates the end of this protein? ___UAG________ (This is called a "STOP codon". This is one of 3 possible STOP codons.) 4. The release factor binds to the A-site relatively wea ...
... ii. __ The large and small ribosomal subunits separate and fall off the mRNA___ 3. What is the sequence of the codon that indicates the end of this protein? ___UAG________ (This is called a "STOP codon". This is one of 3 possible STOP codons.) 4. The release factor binds to the A-site relatively wea ...
Polyadenylation
Polyadenylation is the addition of a poly(A) tail to a messenger RNA The poly(A) tail consists of multiple adenosine monophosphates; in other words, it is a stretch of RNA that has only adenine bases. In eukaryotes, polyadenylation is part of the process that produces mature messenger RNA (mRNA) for translation. It, therefore, forms part of the larger process of gene expression.The process of polyadenylation begins as the transcription of a gene finishes, or terminates. The 3'-most segment of the newly made pre-mRNA is first cleaved off by a set of proteins; these proteins then synthesize the poly(A) tail at the RNA's 3' end. In some genes, these proteins may add a poly(A) tail at any one of several possible sites. Therefore, polyadenylation can produce more than one transcript from a single gene (alternative polyadenylation), similar to alternative splicing.The poly(A) tail is important for the nuclear export, translation, and stability of mRNA. The tail is shortened over time, and, when it is short enough, the mRNA is enzymatically degraded. However, in a few cell types, mRNAs with short poly(A) tails are stored for later activation by re-polyadenylation in the cytosol. In contrast, when polyadenylation occurs in bacteria, it promotes RNA degradation. This is also sometimes the case for eukaryotic non-coding RNAs.mRNA molecules in both prokaryotes and eukaryotes have polyadenylated 3'-ends, with the prokaryotic poly(A) tails generally shorter and less mRNA molecules polyadenylated.