Section 7.2: Transcription: DNA
... 3. (a) The role of the promoter in transcription is to prepare a site where RNA polymerase can access and bind to the DNA strand. (b) The role of RNA polymerase is to read the DNA code and create a complementary RNA molecule. (c) The role of spliceosomes is to take part in eukaryotic post-transcript ...
... 3. (a) The role of the promoter in transcription is to prepare a site where RNA polymerase can access and bind to the DNA strand. (b) The role of RNA polymerase is to read the DNA code and create a complementary RNA molecule. (c) The role of spliceosomes is to take part in eukaryotic post-transcript ...
Chapter 6 From DNA to Protein: How Cell Read the Genome
... RNA splicing might occurred before or after polyadenylation ...
... RNA splicing might occurred before or after polyadenylation ...
Slide 1
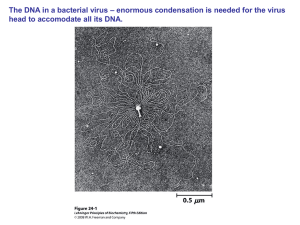
... In Escherichia coli the DNA is about 1 med mer long, while the cell is close to 1 μm. Here the DNA information also has to be read! ...
... In Escherichia coli the DNA is about 1 med mer long, while the cell is close to 1 μm. Here the DNA information also has to be read! ...
chap12studyguide
... 12.In E. coli, the lac operon controls the_________? 13.What are the parts of a Eukaryotic Chromosome? 14.Hox genes determine an animal’s __________? Completion Complete each statement. 15. According to the principle of ____________________, hydrogen bonds can form only between adenine and thymine, ...
... 12.In E. coli, the lac operon controls the_________? 13.What are the parts of a Eukaryotic Chromosome? 14.Hox genes determine an animal’s __________? Completion Complete each statement. 15. According to the principle of ____________________, hydrogen bonds can form only between adenine and thymine, ...
Dr Gisela Storz Biosketch
... Development in Bethesda, where she is a Senior Investigator. Dr. Storz has made contributions in multiple fields of molecular biology, including groundbreaking experiments on the sensing of oxidative stress ...
... Development in Bethesda, where she is a Senior Investigator. Dr. Storz has made contributions in multiple fields of molecular biology, including groundbreaking experiments on the sensing of oxidative stress ...
Messenger RNA profiling: a prototype method to supplant
... resembling mRNA structure but located in DNA Control: amplify DNA, look for ...
... resembling mRNA structure but located in DNA Control: amplify DNA, look for ...
Making Proteins
... The following tRNA has the anticodon UAC. What is the DNA base code for this tRNA? What amino acid would this tRNA carry? ...
... The following tRNA has the anticodon UAC. What is the DNA base code for this tRNA? What amino acid would this tRNA carry? ...
Making Proteins
... 1. Helicase does NOT unzip DNA at the gene of interest 2. RNA polymerase unwinds and matches RNA nucleotide bases to DNA, using one side as a template. 3. The mRNA strand is created. It now compliments the original DNA strand (G-C and A-U). 4. Ligase helps the strand of DNA to close and again. 5. mR ...
... 1. Helicase does NOT unzip DNA at the gene of interest 2. RNA polymerase unwinds and matches RNA nucleotide bases to DNA, using one side as a template. 3. The mRNA strand is created. It now compliments the original DNA strand (G-C and A-U). 4. Ligase helps the strand of DNA to close and again. 5. mR ...
Asymptotics of RNA Shapes: secondary structure
... models and novel algorithms to solve fundamental problems of molecular biology in the post-genome era. A central problem of structural biology concerns the algorithmic prediction of the structure of RNA and protein from only the nucleotide resp. amino acid sequence. In the context of RNA, nucleotide ...
... models and novel algorithms to solve fundamental problems of molecular biology in the post-genome era. A central problem of structural biology concerns the algorithmic prediction of the structure of RNA and protein from only the nucleotide resp. amino acid sequence. In the context of RNA, nucleotide ...
From Gene to Protein
... polymerase (I, II – used in RNA synthesis and III) • Bacteria have one kind – it makes not only mRNA but also other types of RNA • Bacteria have one chromosome and many plasmids. Information is constantly being sent to ribosomes for translation into proteins needed by the bacterial cell ...
... polymerase (I, II – used in RNA synthesis and III) • Bacteria have one kind – it makes not only mRNA but also other types of RNA • Bacteria have one chromosome and many plasmids. Information is constantly being sent to ribosomes for translation into proteins needed by the bacterial cell ...
DNA Transcription & Protein Translation
... 1. To explain how DNA and RNA code for proteins and determine traits. 2. To investigate and understand common mechanisms of protein synthesis. ...
... 1. To explain how DNA and RNA code for proteins and determine traits. 2. To investigate and understand common mechanisms of protein synthesis. ...
Practice Exam- KEY - mvhs
... should eb the same (no introns were cut out). However, if the RNA is shorter than the DNA then you could conclude that it went through some post-translational modifications after transcription. 10. a) OUTER b) Most of the amino acids in this section are either polar or charged, so they will be attra ...
... should eb the same (no introns were cut out). However, if the RNA is shorter than the DNA then you could conclude that it went through some post-translational modifications after transcription. 10. a) OUTER b) Most of the amino acids in this section are either polar or charged, so they will be attra ...
I - 國立彰化師範大學圖書館
... (1). How to map the start site of transcription (+1) of the X gene? (2). Assuming that there is a single metal responsive element (MRE) in the X gene, which fragment should involved? Explain why? (3). If you get the MRE sequence, how would you confirm your data by other experiments? (4). Please writ ...
... (1). How to map the start site of transcription (+1) of the X gene? (2). Assuming that there is a single metal responsive element (MRE) in the X gene, which fragment should involved? Explain why? (3). If you get the MRE sequence, how would you confirm your data by other experiments? (4). Please writ ...
DNA Replication, RNA Molecules and Transcription
... the DNA, NTPs to serve as the building blocks and donors of bonding energy, and a buffered salt solution for optimal RNA polymerase activity. (A) DNA polymerases polymerize deoxyribonucleotides; RNA polymerase polymerizes ribonucleotides. (B) DNA polymerases require a primer to provide a 3’ OH group ...
... the DNA, NTPs to serve as the building blocks and donors of bonding energy, and a buffered salt solution for optimal RNA polymerase activity. (A) DNA polymerases polymerize deoxyribonucleotides; RNA polymerase polymerizes ribonucleotides. (B) DNA polymerases require a primer to provide a 3’ OH group ...
Document
... Translation- Translating language of nucleic acids (base sequences) into language of proteins (amino acids) 1. Gene on DNA carries code to make protein a. Code written in language with only 4 “letters”, the nitrogen bases A,C,G,U ...
... Translation- Translating language of nucleic acids (base sequences) into language of proteins (amino acids) 1. Gene on DNA carries code to make protein a. Code written in language with only 4 “letters”, the nitrogen bases A,C,G,U ...
Sample Questions for EXAM III
... 2. Glucose is a repressor molecule. 3. Catabolite-activating protein-cAMP complex is required for transcription, and its level is low when glucose is present. 4. The CAP binding site is activated. ...
... 2. Glucose is a repressor molecule. 3. Catabolite-activating protein-cAMP complex is required for transcription, and its level is low when glucose is present. 4. The CAP binding site is activated. ...
1 Genetics 301 Sample Second Midterm Examination Solutions
... peptide bond- bond which joins amino acids in forming a polypeptide chain. wobble pairing- unusual hydrogen bond pairing between bases in the tRNA anticodon and the mRNA codon which allows a single tRNA to pair with more than one codon. Okazaki fragment- small, single strand fragments of DNA which a ...
... peptide bond- bond which joins amino acids in forming a polypeptide chain. wobble pairing- unusual hydrogen bond pairing between bases in the tRNA anticodon and the mRNA codon which allows a single tRNA to pair with more than one codon. Okazaki fragment- small, single strand fragments of DNA which a ...
In the nucleus
... The codon of mRNA is translated into the amino acid sequence of a protein. Step 1- Processed mRNA leaves the nucleus and enters the cytoplasm and joins with 2 ribosome subunits. The mRNA start codon (AUG) signals a tRNA molecule carrying methionine and attaches at the anticodon at the P site. St ...
... The codon of mRNA is translated into the amino acid sequence of a protein. Step 1- Processed mRNA leaves the nucleus and enters the cytoplasm and joins with 2 ribosome subunits. The mRNA start codon (AUG) signals a tRNA molecule carrying methionine and attaches at the anticodon at the P site. St ...
PPT NOTES_AP Biology Chapter 17 Notes
... • Enzymes in the eukaryotic nucleus _________________ pre-mRNA before the genetic messages are dispatched to the cytoplasm • During RNA processing, both ___________ of the primary transcript are usually altered • Also, usually some interior parts of the molecule are ________________, and the other p ...
... • Enzymes in the eukaryotic nucleus _________________ pre-mRNA before the genetic messages are dispatched to the cytoplasm • During RNA processing, both ___________ of the primary transcript are usually altered • Also, usually some interior parts of the molecule are ________________, and the other p ...
Topic 3 The Chemistry of Life - wfs
... acid, the codon – anticodon match allows a very specific protein or polypeptide to be produced. 14. A particular sequence on a DNA molecule that codes for a specific protein is called a gene. This gene will always code for the formation of the same protein during the process of its transcription and ...
... acid, the codon – anticodon match allows a very specific protein or polypeptide to be produced. 14. A particular sequence on a DNA molecule that codes for a specific protein is called a gene. This gene will always code for the formation of the same protein during the process of its transcription and ...
DNA, RNA, and Protein
... mRNA docks on ribosome. Its 1st codon is AUG tRNA with met binds via its anticodon UAC. tRNA with its amino binds to 2nd codon. Ribosome detaches met from 1st tRNA. Peptide bond forms between met & 2nd amino acid. First tRNA exits the ribosome & 3rd tRNA enters. Elongation continues until reaches st ...
... mRNA docks on ribosome. Its 1st codon is AUG tRNA with met binds via its anticodon UAC. tRNA with its amino binds to 2nd codon. Ribosome detaches met from 1st tRNA. Peptide bond forms between met & 2nd amino acid. First tRNA exits the ribosome & 3rd tRNA enters. Elongation continues until reaches st ...
Transcription
... • Chemical signals turn gene for a specific protein on. • Enzymes attach to DNA at the gene’s location and unzip only where that gene is on the DNA. – DNA A T C G ...
... • Chemical signals turn gene for a specific protein on. • Enzymes attach to DNA at the gene’s location and unzip only where that gene is on the DNA. – DNA A T C G ...
Randy Carroll
... Helicase: The chains made in DNA separated by Enzymes. Mutation: An error in the replication process of DNA. Nitrogen-Containing Base: An atom surround by oxygen that contains nitrogen. ...
... Helicase: The chains made in DNA separated by Enzymes. Mutation: An error in the replication process of DNA. Nitrogen-Containing Base: An atom surround by oxygen that contains nitrogen. ...
Nucleoside Phosphoramidate Monoesters: Potential
... Formation of RNA Polymerase II pre-initiation complex ...
... Formation of RNA Polymerase II pre-initiation complex ...
Polyadenylation
Polyadenylation is the addition of a poly(A) tail to a messenger RNA The poly(A) tail consists of multiple adenosine monophosphates; in other words, it is a stretch of RNA that has only adenine bases. In eukaryotes, polyadenylation is part of the process that produces mature messenger RNA (mRNA) for translation. It, therefore, forms part of the larger process of gene expression.The process of polyadenylation begins as the transcription of a gene finishes, or terminates. The 3'-most segment of the newly made pre-mRNA is first cleaved off by a set of proteins; these proteins then synthesize the poly(A) tail at the RNA's 3' end. In some genes, these proteins may add a poly(A) tail at any one of several possible sites. Therefore, polyadenylation can produce more than one transcript from a single gene (alternative polyadenylation), similar to alternative splicing.The poly(A) tail is important for the nuclear export, translation, and stability of mRNA. The tail is shortened over time, and, when it is short enough, the mRNA is enzymatically degraded. However, in a few cell types, mRNAs with short poly(A) tails are stored for later activation by re-polyadenylation in the cytosol. In contrast, when polyadenylation occurs in bacteria, it promotes RNA degradation. This is also sometimes the case for eukaryotic non-coding RNAs.mRNA molecules in both prokaryotes and eukaryotes have polyadenylated 3'-ends, with the prokaryotic poly(A) tails generally shorter and less mRNA molecules polyadenylated.