Power point
... more or less able to bind the transcription machinery • Associated with most eukaryotic genes are multiple control elements, segments of noncoding DNA that serve as binding sites for transcription factors that help regulate transcription • Control elements and the transcription factors they bind are ...
... more or less able to bind the transcription machinery • Associated with most eukaryotic genes are multiple control elements, segments of noncoding DNA that serve as binding sites for transcription factors that help regulate transcription • Control elements and the transcription factors they bind are ...
Center for Eukaryotic Structural Genomics (CESG)
... Fusion protein vectors developed for high-throughput protein expression as part of the Protein Structure Initiative have been investigated for use in the expression and stabilization of human cyt b5, a monotopic membrane protein that must be attached to the cellular membrane for function. Expression ...
... Fusion protein vectors developed for high-throughput protein expression as part of the Protein Structure Initiative have been investigated for use in the expression and stabilization of human cyt b5, a monotopic membrane protein that must be attached to the cellular membrane for function. Expression ...
hypothesize that AraC can exist in 2 states, P1 and P2
... -CAP is a symmetrical dimer of two identical subunits -when bound to cAMP (low glucose, high cAMP), CAP is active and binds to a specific palindrome found upstream of genes that are controlled by catabolite repression -consensus: 5’-AAATGTGATCT-AGATCACATTT-3’ -DNA binding mediated by a HTH present i ...
... -CAP is a symmetrical dimer of two identical subunits -when bound to cAMP (low glucose, high cAMP), CAP is active and binds to a specific palindrome found upstream of genes that are controlled by catabolite repression -consensus: 5’-AAATGTGATCT-AGATCACATTT-3’ -DNA binding mediated by a HTH present i ...
Mahoney Abstract for Pathway to Independence Grant
... Mahoney Abstract for Pathway to Independence Grant Arteries and veins must have very distinct properties responsible for each vessel's unique functional demands. For example, arteries, unlike veins, transmit the pressure wave and constrict to maintain pressure and to control blood flow. We propose t ...
... Mahoney Abstract for Pathway to Independence Grant Arteries and veins must have very distinct properties responsible for each vessel's unique functional demands. For example, arteries, unlike veins, transmit the pressure wave and constrict to maintain pressure and to control blood flow. We propose t ...
Gene Section CLIC4 (chloride intracellular channel 4) Atlas of Genetics and Cytogenetics
... CLIC4 expression has also been shown to be upregulated in some tumours. In matched tissue arrays, CLIC4 was predominantly nuclear in normal epithelial tissues but not cancers. As tumours progressed CLIC4 expression became undetectable in tumour cells but increased in stromal cells. Sequence analysis ...
... CLIC4 expression has also been shown to be upregulated in some tumours. In matched tissue arrays, CLIC4 was predominantly nuclear in normal epithelial tissues but not cancers. As tumours progressed CLIC4 expression became undetectable in tumour cells but increased in stromal cells. Sequence analysis ...
Gene Expression Prokaryotes and Viruses
... • Expressed most of the time in most cells • Carry out important cellular functions ...
... • Expressed most of the time in most cells • Carry out important cellular functions ...
Poster
... from mutations in multiple genes. One candidate gene is T. T protein, a transcription factor found in a variety of animals including humans, is essential for correct embryonic development and guides the development of bone and cartilage from embryonic mesodermal tissue. T protein accumulates in the ...
... from mutations in multiple genes. One candidate gene is T. T protein, a transcription factor found in a variety of animals including humans, is essential for correct embryonic development and guides the development of bone and cartilage from embryonic mesodermal tissue. T protein accumulates in the ...
a higher level of chromatin structure.
... have identical DNA sequences, but one is inactivated and the other is not. An epigenetic state can usually be reversed; X chromosomes, for example, are reactivated prior to formation of gametes. Differences in disease susceptibility and longevity between genetically identical twins may be due, in pa ...
... have identical DNA sequences, but one is inactivated and the other is not. An epigenetic state can usually be reversed; X chromosomes, for example, are reactivated prior to formation of gametes. Differences in disease susceptibility and longevity between genetically identical twins may be due, in pa ...
8.4 Transcription - School District of La Crosse
... 8.4 Transcription • Transcription makes three types of RNA. – Messenger RNA (mRNA) carries the message that will be translated to form a protein. – Ribosomal RNA (rRNA) forms part of ribosomes where proteins are made. – Transfer RNA (tRNA) brings amino acids from the cytoplasm to a ribosome. ...
... 8.4 Transcription • Transcription makes three types of RNA. – Messenger RNA (mRNA) carries the message that will be translated to form a protein. – Ribosomal RNA (rRNA) forms part of ribosomes where proteins are made. – Transfer RNA (tRNA) brings amino acids from the cytoplasm to a ribosome. ...
Lecture 4, Exam III Worksheet Answers
... Thymine dimers within DNA. Two thymines beside each other are bonded together by a covalent bond. Nuclease. It is an enzyme that cleaves nucleic acids. Other enzymes that may be needed in this process are the enzymes needed to create DNA and link it together: primase, both DNA polymerases, and ligas ...
... Thymine dimers within DNA. Two thymines beside each other are bonded together by a covalent bond. Nuclease. It is an enzyme that cleaves nucleic acids. Other enzymes that may be needed in this process are the enzymes needed to create DNA and link it together: primase, both DNA polymerases, and ligas ...
Slide 1 - AccessPharmacy
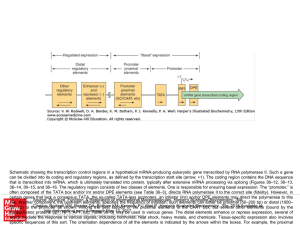
... Schematic showing the transcription control regions in a hypothetical mRNA-producing eukaryotic gene transcribed by RNA polymerase II. Such a gene can be divided into its coding and regulatory regions, as defined by the transcription start site (arrow; +1). The coding region contains the DNA sequenc ...
... Schematic showing the transcription control regions in a hypothetical mRNA-producing eukaryotic gene transcribed by RNA polymerase II. Such a gene can be divided into its coding and regulatory regions, as defined by the transcription start site (arrow; +1). The coding region contains the DNA sequenc ...
activators
... containing a collection of activators bound to an enhancer in such a way that stimulates transcription • The archetypal enhanceosome involves the IFNb enhancer with a structure that involves eight polypeptides bound cooperatively to an ...
... containing a collection of activators bound to an enhancer in such a way that stimulates transcription • The archetypal enhanceosome involves the IFNb enhancer with a structure that involves eight polypeptides bound cooperatively to an ...
SEMESTER II LSM4241 FUNCTIONAL GENOMICS
... SEMESTER II LSM4241 FUNCTIONAL GENOMICS Prerequisite: LSM3231 Workload: 30 lecture hours + 20 tutorial, self study and presentation hours This module aims to introduce selected topics on functional genomics. Areas covered include: the assignment of functions to novel genes following the genome-seque ...
... SEMESTER II LSM4241 FUNCTIONAL GENOMICS Prerequisite: LSM3231 Workload: 30 lecture hours + 20 tutorial, self study and presentation hours This module aims to introduce selected topics on functional genomics. Areas covered include: the assignment of functions to novel genes following the genome-seque ...
Computational-Skills..
... Protein/RNA factors for splicing and controlling organelles Organellar genomes (Plastid, Mitochondrion) Identify genetic elements for replication and transcription ...
... Protein/RNA factors for splicing and controlling organelles Organellar genomes (Plastid, Mitochondrion) Identify genetic elements for replication and transcription ...
Slide 1
... Transcription factor binding sites are not distributed uniformly in promoter regions The motif CGATGAG most frequently occurs between 60 and 100 nucleotides away from the transcription start site (where the code for a protein begins) ...
... Transcription factor binding sites are not distributed uniformly in promoter regions The motif CGATGAG most frequently occurs between 60 and 100 nucleotides away from the transcription start site (where the code for a protein begins) ...
by Tajekesa KP Blee, Nicola K. Gray, and Matthew
... Modulation of the cytoplasmic functions of mammalian post-transcriptional regulatory proteins by methylation and acetylation: a key layer of regulation waiting to be uncovered? by Tajekesa K.P. Blee, Nicola K. Gray, and Matthew Brook ...
... Modulation of the cytoplasmic functions of mammalian post-transcriptional regulatory proteins by methylation and acetylation: a key layer of regulation waiting to be uncovered? by Tajekesa K.P. Blee, Nicola K. Gray, and Matthew Brook ...
Operon
... metabolite that triggers transcription of the lac operon. Unlike allolactose, the sulfur (S) atom creates a chemical bond which is non-hydrolyzable by the cell, preventing the cell from "eating up" or degrading the inductant. IPTG induces activity of betagalactosidase, an enzyme that promotes lactos ...
... metabolite that triggers transcription of the lac operon. Unlike allolactose, the sulfur (S) atom creates a chemical bond which is non-hydrolyzable by the cell, preventing the cell from "eating up" or degrading the inductant. IPTG induces activity of betagalactosidase, an enzyme that promotes lactos ...
Poster
... and topoisomerase I) leaving the DNA free from both of them. The DNA is then electrophoresed on 1.2% agarose at 80 volts for 10 hours at 4°C. The gel is stained with ethidium bromide and photographed (See photo above). The lanes in the gel going from left to right have an increasing amount of histon ...
... and topoisomerase I) leaving the DNA free from both of them. The DNA is then electrophoresed on 1.2% agarose at 80 volts for 10 hours at 4°C. The gel is stained with ethidium bromide and photographed (See photo above). The lanes in the gel going from left to right have an increasing amount of histon ...
Epigenetics - Current Issues in Human Genetics
... Holt. (2007). Epigenetics:Environmental factors can alter the way our genes are expressed, making even identical twins different. PBS. NOVA. Junko, et. al. (2009). Transgenerational Rescue of a Genetic Deficit in LTP and Memory Formation by Juvenile Enrichment. Journal of Neuroscience. 1496-1502. ...
... Holt. (2007). Epigenetics:Environmental factors can alter the way our genes are expressed, making even identical twins different. PBS. NOVA. Junko, et. al. (2009). Transgenerational Rescue of a Genetic Deficit in LTP and Memory Formation by Juvenile Enrichment. Journal of Neuroscience. 1496-1502. ...
Biosynthesis and degradation of proteins
... IAPs are proteins that block apoptosis by binding to and inhibiting caspases. The apoptosis-stimulating protein Smac antagonizes the effect of IAPs on caspases. ...
... IAPs are proteins that block apoptosis by binding to and inhibiting caspases. The apoptosis-stimulating protein Smac antagonizes the effect of IAPs on caspases. ...
Protein misfolding associated to mild modifications of local cellular pH
... and the hydrophobic cavities present in the native state of the protein. This means that misfolding could be associated with intermediate folding states, and protonation of residues. Even though natural pathological mutants show higher tendency to aggregate as amyloid-like structures, wt apoA-I also ...
... and the hydrophobic cavities present in the native state of the protein. This means that misfolding could be associated with intermediate folding states, and protonation of residues. Even though natural pathological mutants show higher tendency to aggregate as amyloid-like structures, wt apoA-I also ...
Regulation of Gene Expression
... Comparison of the same genes in different types of tissues shows that the genes are usually more heavily methylated in cells where they are not expressed. In addition, de-methylating certain inactive genes (removing their extra methyl groups) turns them on. ...
... Comparison of the same genes in different types of tissues shows that the genes are usually more heavily methylated in cells where they are not expressed. In addition, de-methylating certain inactive genes (removing their extra methyl groups) turns them on. ...
How Genes Are Regulated
... the same proteins. Prokaryotic organisms express the entire DNA they encode in every cell, but not necessarily all at the same time. Proteins are expressed only when they are needed. Eukaryotic organisms express a subset of the DNA that is encoded in any given cell. In each cell type, the type and a ...
... the same proteins. Prokaryotic organisms express the entire DNA they encode in every cell, but not necessarily all at the same time. Proteins are expressed only when they are needed. Eukaryotic organisms express a subset of the DNA that is encoded in any given cell. In each cell type, the type and a ...
Tumor Necrosis Factor alpha
... Description: TNF-alpha is a homotrimer with a subunit molecular mass of 17 kDa and that it plays a major role in growth regulation, differentiation, inflammation, viral replication, tumorigenesis, and autoimmune diseases; and in viral, bacterial, fungal, and parasitic infections. Besides inducing he ...
... Description: TNF-alpha is a homotrimer with a subunit molecular mass of 17 kDa and that it plays a major role in growth regulation, differentiation, inflammation, viral replication, tumorigenesis, and autoimmune diseases; and in viral, bacterial, fungal, and parasitic infections. Besides inducing he ...
Histone acetylation and deacetylation

Histone acetylation and deacetylation are the processes by which the lysine residues within the N-terminal tail protruding from the histone core of the nucleosome are acetylated and deacetylated as part of gene regulation. Histone acetylation and deacetylation are essential parts of gene regulation. These reactions are typically catalysed by enzymes with ""histone acetyltransferase"" (HAT) or ""histone deacetylase"" (HDAC) activity. Acetylation is the process where an acetyl functional group is transferred from one molecule (in this case, Acetyl-Coenzyme A) to another. Deacetylation is simply the reverse reaction where an acetyl group is removed from a molecule.Acetylated histones, octameric proteins that organize chromatin into nucleosomes and ultimately higher order structures, represent a type of epigenetic marker within chromatin. Acetylation removes the positive charge on the histones, thereby decreasing the interaction of the N termini of histones with the negatively charged phosphate groups of DNA. As a consequence, the condensed chromatin is transformed into a more relaxed structure that is associated with greater levels of gene transcription. This relaxation can be reversed by HDAC activity. Relaxed, transcriptionally active DNA is referred to as euchromatin. More condensed (tightly packed) DNA is referred to as heterochromatin. Condensation can be brought about by processes including deacetylation and methylation; the action of methylation is indirect and has no effect upon charge.