and Trp cage
... 2. Can we predict general ligand-receptor interactions from structural comparisons, models, and MSA’s? If residues are conserved in the receptors and ligands then these residues are critical for ligandreceptor interactions. 3. Which ligand residues interact with which receptor residues? The chemical ...
... 2. Can we predict general ligand-receptor interactions from structural comparisons, models, and MSA’s? If residues are conserved in the receptors and ligands then these residues are critical for ligandreceptor interactions. 3. Which ligand residues interact with which receptor residues? The chemical ...
Widening the reach of structural biology
... be limited to moderate resolutions of 6–8 Å, too low to give atomic detail, except in cases where crystal structures could be docked into the EM envelope. Spectacular advances have, however, come in the past two years with the development of new detectors (direct electron detectors) that have liter ...
... be limited to moderate resolutions of 6–8 Å, too low to give atomic detail, except in cases where crystal structures could be docked into the EM envelope. Spectacular advances have, however, come in the past two years with the development of new detectors (direct electron detectors) that have liter ...
Concept review: Chromatography (applied to protein purification)
... • 1. Cell disruption should be performed at cold temperatures. Keep the sample on ice as much as possible and use chilled solutions. This will decrease the activity of the proteases for the simple reasons that all chemical reactions occur more slowly at low temperature. • 2. Add protease inhibitors ...
... • 1. Cell disruption should be performed at cold temperatures. Keep the sample on ice as much as possible and use chilled solutions. This will decrease the activity of the proteases for the simple reasons that all chemical reactions occur more slowly at low temperature. • 2. Add protease inhibitors ...
Gene Section MAPK12 (mitogen activated protein kinase 12) -
... protein level increases when myoblast differentiate into myotubes endogenous (Tortorella et al., 2003; Cuenda and Cohen, 1999). Moreover, it has been shown that over-expression of p38gamma in skeletal muscle cells leads to differentiation from myoblast to myotubes, and that a dominant-negative mutan ...
... protein level increases when myoblast differentiate into myotubes endogenous (Tortorella et al., 2003; Cuenda and Cohen, 1999). Moreover, it has been shown that over-expression of p38gamma in skeletal muscle cells leads to differentiation from myoblast to myotubes, and that a dominant-negative mutan ...
Efficient Isolation and Identification of Intracellular Protein
... The HaloTag Pull-Down method is capable of isolating large multiprotein structural complexes such as the NPC 107-160 (4) as well as smaller regulatory protein complexes such as the NFκB complex (5). Recovered protein partners can either be analyzed by Western blotting if binding partners are kno ...
... The HaloTag Pull-Down method is capable of isolating large multiprotein structural complexes such as the NPC 107-160 (4) as well as smaller regulatory protein complexes such as the NFκB complex (5). Recovered protein partners can either be analyzed by Western blotting if binding partners are kno ...
aLFQ: an R-package for estimating absolute protein quantities from
... 1 INTRODUCTION A variety of quantitative proteomic methods have been established to measure the relative abundance of proteins across samples. Although relative quantification methods are useful to compare the same proteins between multiple biological samples, they do not provide the possibility to ...
... 1 INTRODUCTION A variety of quantitative proteomic methods have been established to measure the relative abundance of proteins across samples. Although relative quantification methods are useful to compare the same proteins between multiple biological samples, they do not provide the possibility to ...
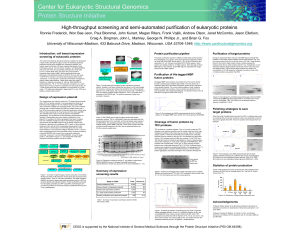
High-throughput screening and semi
... Design of expression plasmid The cloned genes are under the control of a T5 phage based promoter that is IPTG or lactose inducible. The cloned genes are fused to an N-terminal (His)n-tagged (n=6 or 8) maltose binding protein (MBP which enhances solubility and expression levels), and a TEV protease c ...
... Design of expression plasmid The cloned genes are under the control of a T5 phage based promoter that is IPTG or lactose inducible. The cloned genes are fused to an N-terminal (His)n-tagged (n=6 or 8) maltose binding protein (MBP which enhances solubility and expression levels), and a TEV protease c ...
presentation source
... enzymes • broken by reducing agents, e.g., mercaptoethanol and dithiothreitol – CH2-S - S-CH2 - ------> -CH2 -SH + HS-CH2- ...
... enzymes • broken by reducing agents, e.g., mercaptoethanol and dithiothreitol – CH2-S - S-CH2 - ------> -CH2 -SH + HS-CH2- ...
Slide 1 - Elsevier
... action of the Ca2+-calmodulin complex on many enzymes, and calcium has some direct effects on enzymes such as PKC and calpain. CaM kinase, Ca2+-calmodulin-dependent kinase. Copyright © 2014 Elsevier Inc. All rights reserved. ...
... action of the Ca2+-calmodulin complex on many enzymes, and calcium has some direct effects on enzymes such as PKC and calpain. CaM kinase, Ca2+-calmodulin-dependent kinase. Copyright © 2014 Elsevier Inc. All rights reserved. ...
Protein Structure - Particle Sciences
... can be quite a challenge, but without a good understanding of the nature of protein structure and the conformational characteristics of the specific protein being formulated, the results can be ruinous. This technical brief aims to give the reader a quick overview of protein structure. It will also ...
... can be quite a challenge, but without a good understanding of the nature of protein structure and the conformational characteristics of the specific protein being formulated, the results can be ruinous. This technical brief aims to give the reader a quick overview of protein structure. It will also ...
Hydrogen Bonds, Hydrophobicity Forces and the Character of the
... – The decrease in hydrogen-bond energy per amino acid, Ehb /N, with decreasing temperature gets more rapid with increasing chain length, as shown in Figure 5a. This implies that the three-helix protein makes more stable secondary structure than the one- and two-helix segments. It turns out that the ...
... – The decrease in hydrogen-bond energy per amino acid, Ehb /N, with decreasing temperature gets more rapid with increasing chain length, as shown in Figure 5a. This implies that the three-helix protein makes more stable secondary structure than the one- and two-helix segments. It turns out that the ...
Signaling Through Scaffold, Anchoring, and Adaptor Proteins
... 35 to 40 residues that also bind proline-rich motifs, commonly with the consensus PPXY or PPLP (Fig. 2E) (25). The WW domains of the E3 ubiquitin protein ligase Nedd4 bind such proline-rich motifs in an amiloridesensitive epithelial Na1 channel, likely leading to channel degradation (Fig. 3A) (26). ...
... 35 to 40 residues that also bind proline-rich motifs, commonly with the consensus PPXY or PPLP (Fig. 2E) (25). The WW domains of the E3 ubiquitin protein ligase Nedd4 bind such proline-rich motifs in an amiloridesensitive epithelial Na1 channel, likely leading to channel degradation (Fig. 3A) (26). ...
GroEL and GroES - ETH - D-INFK - TI
... Finally, hydrolysis of ATP in the cis ring allows for the weakening of the structure. Binding of the ATP in the trans ring causes the disassembly of the cis assembly, allowing for the release of the GroES and substrate protein. In the end, the ring is left in original state. It needs to be noted th ...
... Finally, hydrolysis of ATP in the cis ring allows for the weakening of the structure. Binding of the ATP in the trans ring causes the disassembly of the cis assembly, allowing for the release of the GroES and substrate protein. In the end, the ring is left in original state. It needs to be noted th ...
interrpo_nov16
... Protein signatures Alternatively, model the pattern of conserved amino acids at specific positions within a multiple sequence alignment • Patterns • Profiles • Profile HMMs Use these models (signatures) to infer relationships with the characterised sequences from which the alignment was constructed ...
... Protein signatures Alternatively, model the pattern of conserved amino acids at specific positions within a multiple sequence alignment • Patterns • Profiles • Profile HMMs Use these models (signatures) to infer relationships with the characterised sequences from which the alignment was constructed ...
Gene Section SOCS2 (suppressor of cytokine signaling 2) in Oncology and Haematology
... Diagram representing the structure of SOCS proteins. At least eight proteins belonging to the SOCS family of proteins are shown (upper panel). They are characterized by the presence of an SH2 central domain and the SOCS box domain at the C-terminus. A small domain called kinase inhibitory region (KI ...
... Diagram representing the structure of SOCS proteins. At least eight proteins belonging to the SOCS family of proteins are shown (upper panel). They are characterized by the presence of an SH2 central domain and the SOCS box domain at the C-terminus. A small domain called kinase inhibitory region (KI ...
Protein domain

A protein domain is a conserved part of a given protein sequence and (tertiary) structure that can evolve, function, and exist independently of the rest of the protein chain. Each domain forms a compact three-dimensional structure and often can be independently stable and folded. Many proteins consist of several structural domains. One domain may appear in a variety of different proteins. Molecular evolution uses domains as building blocks and these may be recombined in different arrangements to create proteins with different functions. Domains vary in length from between about 25 amino acids up to 500 amino acids in length. The shortest domains such as zinc fingers are stabilized by metal ions or disulfide bridges. Domains often form functional units, such as the calcium-binding EF hand domain of calmodulin. Because they are independently stable, domains can be ""swapped"" by genetic engineering between one protein and another to make chimeric proteins.