Unit 8 - Macromolecules Processes
... Deoxyribose Nucleic Acid double helix sugar-phosphate backbone made up of subunits called NUCLEOTIDES ...
... Deoxyribose Nucleic Acid double helix sugar-phosphate backbone made up of subunits called NUCLEOTIDES ...
Chapter_9_Student
... During gel electrophoresis, DNA fragments become separated because a) multiple copies of DNA are made. b) smaller DNA molecules move faster than larger fragments. ...
... During gel electrophoresis, DNA fragments become separated because a) multiple copies of DNA are made. b) smaller DNA molecules move faster than larger fragments. ...
DNA to RNA practice
... Finally, follow the process as you complete the amino acid sequence from the tRNA sequence. Make sure to break them down into codons first. Read the chart and write the first three letters of the amino acid followed by a dash. Ex. Leu – Arg - Pro ...
... Finally, follow the process as you complete the amino acid sequence from the tRNA sequence. Make sure to break them down into codons first. Read the chart and write the first three letters of the amino acid followed by a dash. Ex. Leu – Arg - Pro ...
DNA and Chromosomes
... almost exactly the same. However, it has one important differences: It is possible that the new strand of DNA will have a few “errors”. For example, maybe a G was matched up with a T, instead of a C. Or, maybe a whole section of DNA got moved to a new area. ...
... almost exactly the same. However, it has one important differences: It is possible that the new strand of DNA will have a few “errors”. For example, maybe a G was matched up with a T, instead of a C. Or, maybe a whole section of DNA got moved to a new area. ...
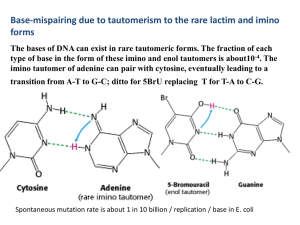
4 . The imino tautomer of adenine can pair with cytosine
... deaminated to hypoxanthine, cytosine to uracil, and guanine to xanthine. Hypoxanthine pairs with cytosine, inducing a mutation of A-T to G-C. It is likely that mutations in DNA repair genes will lead to the accumulation of mutations throughout the genome. In time, genes important in controlling cell ...
... deaminated to hypoxanthine, cytosine to uracil, and guanine to xanthine. Hypoxanthine pairs with cytosine, inducing a mutation of A-T to G-C. It is likely that mutations in DNA repair genes will lead to the accumulation of mutations throughout the genome. In time, genes important in controlling cell ...
Ch. 12 Introduction to Biotechnology
... • “Golden rice” has been genetically modified to contain beta-carotene ...
... • “Golden rice” has been genetically modified to contain beta-carotene ...
AP Biology – Evolution Unit
... The DNA molecule consists of two strands that wrap around each other to form a long, twisted ladder called a double helix. The structure of DNA was deduced in 1956 by two scientists named Watson and Crick. Structure of DNA Draw and label the four nucleotides of DNA. Label the deoxyribose sugar (five ...
... The DNA molecule consists of two strands that wrap around each other to form a long, twisted ladder called a double helix. The structure of DNA was deduced in 1956 by two scientists named Watson and Crick. Structure of DNA Draw and label the four nucleotides of DNA. Label the deoxyribose sugar (five ...
iProof™ High-Fidelity DNA Polymerase - Bio-Rad
... 3% DMSO which aids in template denaturation. Further optimization of DMSO should be made in 2% increments. In some cases, DMSO may be used to help relax supercoiled plasmid DNA. High DMSO concentrations (10%) will require lowering the annealing temperature by 5.5–6.0°C. Other PCR additives such as f ...
... 3% DMSO which aids in template denaturation. Further optimization of DMSO should be made in 2% increments. In some cases, DMSO may be used to help relax supercoiled plasmid DNA. High DMSO concentrations (10%) will require lowering the annealing temperature by 5.5–6.0°C. Other PCR additives such as f ...
Molecular genetics
... group) is added to the 5’ end of RNA after splicing. RNA cap determines the site of translation. PolyA tailing is the process by which a long tail of Adenine residue is added to the 3’ end of m-RNA during splicing. Ribozymes are RNA molecules act as enzymes. RNase P is a Ribozyme. 9. Recombinant DNA ...
... group) is added to the 5’ end of RNA after splicing. RNA cap determines the site of translation. PolyA tailing is the process by which a long tail of Adenine residue is added to the 3’ end of m-RNA during splicing. Ribozymes are RNA molecules act as enzymes. RNase P is a Ribozyme. 9. Recombinant DNA ...
Review-Qs-for-modern-genetics
... 1. The main enzyme involved in DNA replication is RNA polymerase. FALSE – DNA polymerase. 2. To determine the amino acid, look up the three base anticodon on the genetic dictionary FALSE – codon. 3. Ligase joins DNA fragments of the lagging strand. TRUE 4. DNA polymerase lengthens the new strands fr ...
... 1. The main enzyme involved in DNA replication is RNA polymerase. FALSE – DNA polymerase. 2. To determine the amino acid, look up the three base anticodon on the genetic dictionary FALSE – codon. 3. Ligase joins DNA fragments of the lagging strand. TRUE 4. DNA polymerase lengthens the new strands fr ...
AUTOMATED DNA SEQUENCING, MegaBACE 1000
... Because the slab gel electrophoresis was still manual and needed several steps, modern capillary systems were introduced. These systems are currently the fastest system available they use the same basic electrophoresis and detection method technology as the slab ...
... Because the slab gel electrophoresis was still manual and needed several steps, modern capillary systems were introduced. These systems are currently the fastest system available they use the same basic electrophoresis and detection method technology as the slab ...
CHAPTER 10: DNA,RNA & Protein Synthesis
... 2. Elongation- continued as ribosome moves the distance of 1 codon on mRNA 3. Elongation is built with new tRNAs attaching each amino acid as it reads the codons on the mRNA. 4. Termination- ribosome reaches “stop” codon on the mRNA 5. Disassembly – each piece is free. ...
... 2. Elongation- continued as ribosome moves the distance of 1 codon on mRNA 3. Elongation is built with new tRNAs attaching each amino acid as it reads the codons on the mRNA. 4. Termination- ribosome reaches “stop” codon on the mRNA 5. Disassembly – each piece is free. ...
Agarose gel electrophoresis

Agarose gel electrophoresis is a method of gel electrophoresis used in biochemistry, molecular biology, and clinical chemistry to separate a mixed population of DNA or proteins in a matrix of agarose. The proteins may be separated by charge and/or size (isoelectric focusing agarose electrophoresis is essentially size independent), and the DNA and RNA fragments by length. Biomolecules are separated by applying an electric field to move the charged molecules through an agarose matrix, and the biomolecules are separated by size in the agarose gel matrix.Agarose gels are easy to cast and are particularly suitable for separating DNA of size range most often encountered in laboratories, which accounts for the popularity of its use. The separated DNA may be viewed with stain, most commonly under UV light, and the DNA fragments can be extracted from the gel with relative ease. Most agarose gels used are between 0.7 - 2% dissolved in a suitable electrophoresis buffer.