Proteins - Science Learning Hub
... Context > Food Function and Structure > Teaching and Learning Approaches > Proteins ...
... Context > Food Function and Structure > Teaching and Learning Approaches > Proteins ...
Fluorescent Dyes and Proteins
... 1. What kind of structures are fluorescent 2. How to make and target fluorescent probes 3. Fluorescent probes for cellular structure and function 4. Using light to control cells ...
... 1. What kind of structures are fluorescent 2. How to make and target fluorescent probes 3. Fluorescent probes for cellular structure and function 4. Using light to control cells ...
RQ for Ex. 2
... A. If you remove the proteins from all the chromatin isolated from interphase cells, how many types of histones should you get? (1) (2) (3) (4) (5) (6) (7) (8) (>8) (depends how cells were grown). No explanation required. B. In the chromatin from the promoter region of HIV, there is a region of DNA ...
... A. If you remove the proteins from all the chromatin isolated from interphase cells, how many types of histones should you get? (1) (2) (3) (4) (5) (6) (7) (8) (>8) (depends how cells were grown). No explanation required. B. In the chromatin from the promoter region of HIV, there is a region of DNA ...
Soy Protein in Milk Replacers
... Problems with soy protein. One of the biggest problems with using soy proteins in milk replacers is the presence of anti-nutritional factors in soybeans. These include trypsin inhibitor, glycinin and βconglycinin. Trypsin inhibitor can reduce digestibility by binding trypsin, an enzyme in the digest ...
... Problems with soy protein. One of the biggest problems with using soy proteins in milk replacers is the presence of anti-nutritional factors in soybeans. These include trypsin inhibitor, glycinin and βconglycinin. Trypsin inhibitor can reduce digestibility by binding trypsin, an enzyme in the digest ...
THERAPUETIC DISCOVERY BY MODELLING
... offers a way to speed up discovery time and reduce costs, but such techniques have typically had low accuracy and need high resolution structures. We will capitalise on advances in computational protein structure prediction and protein docking to improve accuracy of target-based in silico compound s ...
... offers a way to speed up discovery time and reduce costs, but such techniques have typically had low accuracy and need high resolution structures. We will capitalise on advances in computational protein structure prediction and protein docking to improve accuracy of target-based in silico compound s ...
A green glow
... have been developed to study their function. One such technique is used to find out where and when a particular protein is produced – in the brain, the lower limbs or the digestive system for instance (Fig.3). More detailed questions as to the protein’s role in the cell, for example, can be answered ...
... have been developed to study their function. One such technique is used to find out where and when a particular protein is produced – in the brain, the lower limbs or the digestive system for instance (Fig.3). More detailed questions as to the protein’s role in the cell, for example, can be answered ...
10-30-ramnath
... The vacuolar protein sorting (VPS) pathway of Saccharomyces cerevisiae mediates transport of vacuolar protein precursors from the late Golgi to the lysosomelike vacuole. Sorting of some vacuolar proteins occurs via a prevacuolar endosomal compartment and mutations in a subset of VPS genes (the class ...
... The vacuolar protein sorting (VPS) pathway of Saccharomyces cerevisiae mediates transport of vacuolar protein precursors from the late Golgi to the lysosomelike vacuole. Sorting of some vacuolar proteins occurs via a prevacuolar endosomal compartment and mutations in a subset of VPS genes (the class ...
proteins
... • The tertiary structure is held together by bonds between the R groups of the amino acids in the protein, and so depends on what the sequence of amino acids is. There are three kinds of bonds involved: – hydrogen bonds, which are weak. – ionic bonds between R-groups with positive or negative charg ...
... • The tertiary structure is held together by bonds between the R groups of the amino acids in the protein, and so depends on what the sequence of amino acids is. There are three kinds of bonds involved: – hydrogen bonds, which are weak. – ionic bonds between R-groups with positive or negative charg ...
determination of dna sequence specificity of a dna
... another control of DNA alone. The gels are run for a period of time, and compared upon completion. ...
... another control of DNA alone. The gels are run for a period of time, and compared upon completion. ...
B2 Protein structure and function
... relatively cost and laborious procedure. • Computational methods will allow the prediction of both structure and possible function from simple amino acid sequence information. • Understanding of the true function of a protein still requires its isolation and biochemical and structure characterizatio ...
... relatively cost and laborious procedure. • Computational methods will allow the prediction of both structure and possible function from simple amino acid sequence information. • Understanding of the true function of a protein still requires its isolation and biochemical and structure characterizatio ...
Insights into membrane protein function from molecular modelling
... perception of pain. Moreover, single non-synonymous nucleotide polymorphisms (SNPs) in the corresponding gene have been linked to affective mood disorders such as major depression. In a collaboration between the laboratories of Lin-Hua Jiang and Steve Baldwin at Leeds, we have attempted to identify ...
... perception of pain. Moreover, single non-synonymous nucleotide polymorphisms (SNPs) in the corresponding gene have been linked to affective mood disorders such as major depression. In a collaboration between the laboratories of Lin-Hua Jiang and Steve Baldwin at Leeds, we have attempted to identify ...
Sample Preparation I (Protein Purification)
... Our life is maintained by molecular network systems ...
... Our life is maintained by molecular network systems ...
tutorial10_3D_structure
... • Structural alignment attempts to establish homology between two or more protein structures based on their 3D conformation. • Structural alignment often implies evolutionary relationships between proteins with low seq-id. ...
... • Structural alignment attempts to establish homology between two or more protein structures based on their 3D conformation. • Structural alignment often implies evolutionary relationships between proteins with low seq-id. ...
Slide 1
... • to reach “Similarity page” label a protein in the list in the “Samples page” and click on the icon “Similarity” on the left side • the upper panel provides a vertical list of peptides assigned to this protein, and a horizontal list of accession numbers of additional proteins, where this peptide is ...
... • to reach “Similarity page” label a protein in the list in the “Samples page” and click on the icon “Similarity” on the left side • the upper panel provides a vertical list of peptides assigned to this protein, and a horizontal list of accession numbers of additional proteins, where this peptide is ...
MY FAVORITE PROTEIN Activity - Center for Biophysics and
... significance of your protein. o At least 1 VMD representation of your protein which highlights interesting features of your protein's structure, such as its binding/active site and features contributing to its stability, like hydrogen bonds (for example in alpha helices & beta sheets), disulfide bri ...
... significance of your protein. o At least 1 VMD representation of your protein which highlights interesting features of your protein's structure, such as its binding/active site and features contributing to its stability, like hydrogen bonds (for example in alpha helices & beta sheets), disulfide bri ...
A1982NF37500001
... presence of protein. (Incidentally, the en2 zyme uses the a-side of DPNH. ) “As perhaps the most studied protein, serum albumin serves as a model for other proteins and has great importance and interest in its own right. There are many clinical, commercial, and laboratory applications which require ...
... presence of protein. (Incidentally, the en2 zyme uses the a-side of DPNH. ) “As perhaps the most studied protein, serum albumin serves as a model for other proteins and has great importance and interest in its own right. There are many clinical, commercial, and laboratory applications which require ...
8 M Guanidine Hydrochloride Solution Buffered, pH - Sigma
... phosphates, or carboxyl groups, and therefore, is compatible with mass spectrometric procedures. Guanidine hydrochloride is commonly used as a denaturant, because of its ability to break hydrogen bonds between amino acid residues. By breaking these bonds, the 3D conformation of the protein is unfold ...
... phosphates, or carboxyl groups, and therefore, is compatible with mass spectrometric procedures. Guanidine hydrochloride is commonly used as a denaturant, because of its ability to break hydrogen bonds between amino acid residues. By breaking these bonds, the 3D conformation of the protein is unfold ...
bodylogix.com gnc.ca bodylogix.com gnc.ca gnc.ca
... this revolutionary water is packed with health benefits: 10 grams of high-quality whey protein to help with metabolism and energy; 5 grams of fibre to keep you feeling full longer; electrolytes to help rehydrate, recover and rebuild after activity; and a balanced assortment of vitamins. Contains onl ...
... this revolutionary water is packed with health benefits: 10 grams of high-quality whey protein to help with metabolism and energy; 5 grams of fibre to keep you feeling full longer; electrolytes to help rehydrate, recover and rebuild after activity; and a balanced assortment of vitamins. Contains onl ...
TD11 Identification of in vivo substrates of GroEL Nature 1999, 402
... -compare to ESI (used in Walsh study from TD3) - better accuracy w/ ESI - MALDI generally easier for high MW samples - MALDI requires smaller sample size - Both are used commonly for proteomics Ⅷ. ...
... -compare to ESI (used in Walsh study from TD3) - better accuracy w/ ESI - MALDI generally easier for high MW samples - MALDI requires smaller sample size - Both are used commonly for proteomics Ⅷ. ...
Effect of sol-gel encapsulation on the spectroscopic and
... new composite materials incorporate the chemically selective functionality and reactivity of different biomolecules in solid state form and have led to the perspective of developing a new class of solid state chemical and biomedical sensors (1). The use of sol-gel derived silicate materials for prot ...
... new composite materials incorporate the chemically selective functionality and reactivity of different biomolecules in solid state form and have led to the perspective of developing a new class of solid state chemical and biomedical sensors (1). The use of sol-gel derived silicate materials for prot ...
Ethanol production will have to increase to meet government
... Protein is needed to build enzymes, antibodies and some hormones. Proteins are also needed for blood clotting, wound healing and water balance. “Proteins are long chains of amino acids,” Hermann said. “There are 20 amino acids and the body cannot make nine of these amino acids. These amino acids are ...
... Protein is needed to build enzymes, antibodies and some hormones. Proteins are also needed for blood clotting, wound healing and water balance. “Proteins are long chains of amino acids,” Hermann said. “There are 20 amino acids and the body cannot make nine of these amino acids. These amino acids are ...
Antiporter-lika proteinsubenheter i andningskedjans Komplex I
... All organisms need energy to survive. The energy comes either from sunlight (photosynthesis) or from burning food molecules (respiration), but in both cases the energy is converted to ATP by the organism. ATP is an energy stock that can be used by a multitude of processes in the cell. Eukaryote aero ...
... All organisms need energy to survive. The energy comes either from sunlight (photosynthesis) or from burning food molecules (respiration), but in both cases the energy is converted to ATP by the organism. ATP is an energy stock that can be used by a multitude of processes in the cell. Eukaryote aero ...
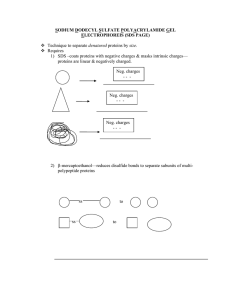
SDS Electrophoresis
... But then separated subunits/polypeptides will be linear & negative due to SDS treatment 3) Heat—to further denature proteins 4) polyacrylamide—gel matrix that acts as size sorter 5) electrophoresis, using electric field with positive anode and negative cathode, all proteins are attracted to bottom ...
... But then separated subunits/polypeptides will be linear & negative due to SDS treatment 3) Heat—to further denature proteins 4) polyacrylamide—gel matrix that acts as size sorter 5) electrophoresis, using electric field with positive anode and negative cathode, all proteins are attracted to bottom ...
Copper(II) - Sigma
... 6. Dihazi, H., et al., One-step purification of recombinant yeast 6-phosphofructo-2-kinase after the identification of contaminants by MALDI-TOF MS. Protein Expr. Purif., 21(1), 201-209 (2001). 7. Fernandes, S., et al., Affinity extraction of dye- and metal ion-binding proteins in polyvinylpyrrolido ...
... 6. Dihazi, H., et al., One-step purification of recombinant yeast 6-phosphofructo-2-kinase after the identification of contaminants by MALDI-TOF MS. Protein Expr. Purif., 21(1), 201-209 (2001). 7. Fernandes, S., et al., Affinity extraction of dye- and metal ion-binding proteins in polyvinylpyrrolido ...
Bimolecular fluorescence complementation

Bimolecular fluorescence complementation (also known as BiFC) is a technology typically used to validate protein interactions. It is based on the association of fluorescent protein fragments that are attached to components of the same macromolecular complex. Proteins that are postulated to interact are fused to unfolded complementary fragments of a fluorescent reporter protein and expressed in live cells. Interaction of these proteins will bring the fluorescent fragments within proximity, allowing the reporter protein to reform in its native three-dimensional structure and emit its fluorescent signal. This fluorescent signal can be detected and located within the cell using an inverted fluorescence microscope that allows imaging of fluorescence in cells. In addition, the intensity of the fluorescence emitted is proportional to the strength of the interaction, with stronger levels of fluorescence indicating close or direct interactions and lower fluorescence levels suggesting interaction within a complex. Therefore, through the visualisation and analysis of the intensity and distribution of fluorescence in these cells, one can identify both the location and interaction partners of proteins of interest.