DISULFIDE GROUPS Disulfide bonds in proteins are
... Disulfide bonds in proteins are subject to cleavage by mild reducing agents (Section 8-1), by a few oxidizing agents (Section 8-6), or by nucleophilic displacement (Section 8-2). Of the procedures for cleavage, those involving reduction are probably the mildest and most specific. Many smaller protei ...
... Disulfide bonds in proteins are subject to cleavage by mild reducing agents (Section 8-1), by a few oxidizing agents (Section 8-6), or by nucleophilic displacement (Section 8-2). Of the procedures for cleavage, those involving reduction are probably the mildest and most specific. Many smaller protei ...
Classification of Protein 3D Structures Using Artificial Neural
... Three dimensional (3D) structures of a protein are determined by the amino acid sequence. Protein functions depend on their structures. Generally, the determination of protein structures aims at inferring functional information of the protein. With the incredible increase of sequenced protein inform ...
... Three dimensional (3D) structures of a protein are determined by the amino acid sequence. Protein functions depend on their structures. Generally, the determination of protein structures aims at inferring functional information of the protein. With the incredible increase of sequenced protein inform ...
Top down - The Fenyo Lab
... RED: triplicate experiments, FAl treated grindate BLACK: duplicated experiments, FAl treated cells (then ground) SCORE: Log Ion Current / Log protein abundance ...
... RED: triplicate experiments, FAl treated grindate BLACK: duplicated experiments, FAl treated cells (then ground) SCORE: Log Ion Current / Log protein abundance ...
Technical White Paper SOMAmer® Reagent Specificity
... those of non-specific interactions, a polyanionic competitor, present in excess, rapidly occupies binding sites freed by the dissociated non-cognate complexes and prevents their rebinding. One of the reasons a high degree of multiplexing is achievable in SOMAscan assay is the fact that, since all SO ...
... those of non-specific interactions, a polyanionic competitor, present in excess, rapidly occupies binding sites freed by the dissociated non-cognate complexes and prevents their rebinding. One of the reasons a high degree of multiplexing is achievable in SOMAscan assay is the fact that, since all SO ...
Bioinformatics and Functional Genomics, Chapter 8, Part 1
... of a protein. Its size is often 10 to 20 amino acids. Simple motifs include transmembrane domains and phosphorylation sites. These do not imply homology when found in a group of proteins. PROSITE (www.expasy.org/prosite) is a dictionary of motifs (there are currently 1600 entries). In PROSITE, a pat ...
... of a protein. Its size is often 10 to 20 amino acids. Simple motifs include transmembrane domains and phosphorylation sites. These do not imply homology when found in a group of proteins. PROSITE (www.expasy.org/prosite) is a dictionary of motifs (there are currently 1600 entries). In PROSITE, a pat ...
Class: Protein functional Annotation and Family Classification
... When an experiment yields a sequence (or a set of sequences), we need to find out as much as we can about this protein and its possible function from available data Especially important for poorly characterized or uncharacterized (“hypothetical”) proteins More challenging for large sets of sequences ...
... When an experiment yields a sequence (or a set of sequences), we need to find out as much as we can about this protein and its possible function from available data Especially important for poorly characterized or uncharacterized (“hypothetical”) proteins More challenging for large sets of sequences ...
Intrinsically unstructured proteins
... These proteins are characterized by an almost complete lack of folded structure and an extended conformation with high intramolecular flexibility and little secondary structure. These unusual features are apparent with most physicochemical techniques, which makes their detection straightforward (Box ...
... These proteins are characterized by an almost complete lack of folded structure and an extended conformation with high intramolecular flexibility and little secondary structure. These unusual features are apparent with most physicochemical techniques, which makes their detection straightforward (Box ...
Diversity of proteins
... • A _______ ________ may have a particular function • __________ between 2 domains provide crevices, grooves, and pockets on the surface of a protein for binding or catalytic sites ...
... • A _______ ________ may have a particular function • __________ between 2 domains provide crevices, grooves, and pockets on the surface of a protein for binding or catalytic sites ...
Cytochrome c regulates SET-mediated acetylation of the C
... 1H NMR - tracking Cc Met80-εCH3 – was used to assess the competition between p53-CTD and Cc for SET. Increasing concentrations of p53-CTD led to dissociation of Cc from SET and recovery of the Met80-εCH3 signal, indicating p53-CTD is able to compete with Cc for SET binding. 1D 19F NMR was then used ...
... 1H NMR - tracking Cc Met80-εCH3 – was used to assess the competition between p53-CTD and Cc for SET. Increasing concentrations of p53-CTD led to dissociation of Cc from SET and recovery of the Met80-εCH3 signal, indicating p53-CTD is able to compete with Cc for SET binding. 1D 19F NMR was then used ...
Moonlighting proteins—an update
... proteins make use of the general methods seen in previously identified moonlighting proteins to switch between functions, although the details differ, for example how interacting with a different protein partner or cofactor results in a conformational change that then affects function. For some of the n ...
... proteins make use of the general methods seen in previously identified moonlighting proteins to switch between functions, although the details differ, for example how interacting with a different protein partner or cofactor results in a conformational change that then affects function. For some of the n ...
Protein Structure Prediction
... Gene Ontology - flexibility • Imagine • protein 1 phosphorylates protein 2 • protein 2 binds to protein 3 (which then binds to DNA) • proteins 1, 2, or 3 may be coded on nearby genes • makes sense in terms of regulation / protein production • different metabolic functions • part of same "cellular p ...
... Gene Ontology - flexibility • Imagine • protein 1 phosphorylates protein 2 • protein 2 binds to protein 3 (which then binds to DNA) • proteins 1, 2, or 3 may be coded on nearby genes • makes sense in terms of regulation / protein production • different metabolic functions • part of same "cellular p ...
Phase behaviour and transitions of peptides and proteins
... My research is focused on the application of theoretical computational tools developed in soft condensed matter physics to investigate the phase behaviour and transitions of complex systems of biomolecules. From a purely statistical mechanical point of view an ensemble of many peptides and proteins ...
... My research is focused on the application of theoretical computational tools developed in soft condensed matter physics to investigate the phase behaviour and transitions of complex systems of biomolecules. From a purely statistical mechanical point of view an ensemble of many peptides and proteins ...
Budding yeast Saccharomyces cerevisiae as a model to study
... glutaredoxin Grx5 localized in mitochondria protected S. cerevisiae cells against oxidative damage caused by menadione and hydrogen peroxide (RodríguezManzaneque et al., 1999). Deletion of the gene coding for Grx5 significantly increased the levels of protein carbonyls; besides, several specific pro ...
... glutaredoxin Grx5 localized in mitochondria protected S. cerevisiae cells against oxidative damage caused by menadione and hydrogen peroxide (RodríguezManzaneque et al., 1999). Deletion of the gene coding for Grx5 significantly increased the levels of protein carbonyls; besides, several specific pro ...
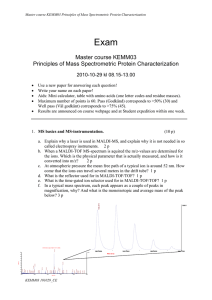
Master course KEMM03 Principles of Mass Spectrometric Protein
... b. In the Mascot output table below, two peptides contain missed cleavage sites. Which peptides and what cleavage sites? Answer with peptide masses, and indicate where in each peptide the missed cleavage site is. Use one-letter code and amino acid number in sequence. 2p ...
... b. In the Mascot output table below, two peptides contain missed cleavage sites. Which peptides and what cleavage sites? Answer with peptide masses, and indicate where in each peptide the missed cleavage site is. Use one-letter code and amino acid number in sequence. 2p ...
Proteomics
... – The digestion enzyme is Trypsin, with 0 missed cleavages – The result is a list of theoretical peptides that can be compared to the measured masses ...
... – The digestion enzyme is Trypsin, with 0 missed cleavages – The result is a list of theoretical peptides that can be compared to the measured masses ...
pick your protein
... 45 g per drink. The ideal dose of protein is 20 – 40 g per serving (1). If the product is 100% protein or has very little carbohydrates, a recommended intake method is to mix the protein powder with almond milk, juice, or a sport drink for some additional carbohydrates, if additional carbohydrates a ...
... 45 g per drink. The ideal dose of protein is 20 – 40 g per serving (1). If the product is 100% protein or has very little carbohydrates, a recommended intake method is to mix the protein powder with almond milk, juice, or a sport drink for some additional carbohydrates, if additional carbohydrates a ...
Towards the atomic level protein sequence analysis
... Proteins differ in the arrangements of 20 naturally occurring amino acids. This difference in protein sequence can also be captured at atom level. Carbon is the only element that contributes towards the hydrophobic interactions that drives the protein to carry out its biochemical reactions. Understa ...
... Proteins differ in the arrangements of 20 naturally occurring amino acids. This difference in protein sequence can also be captured at atom level. Carbon is the only element that contributes towards the hydrophobic interactions that drives the protein to carry out its biochemical reactions. Understa ...
A Acidic amino acids: Those whose side chains can carry a negative
... Beta-conformation: Stable “zig-zag” conformation of protein chain which can align with sections of chain in same conformation and cross-link efficiently via hydrogen bonds between the peptide linkages. One of the classic “secondary” structures. Beta sheet: Two or more sections of protein chain in be ...
... Beta-conformation: Stable “zig-zag” conformation of protein chain which can align with sections of chain in same conformation and cross-link efficiently via hydrogen bonds between the peptide linkages. One of the classic “secondary” structures. Beta sheet: Two or more sections of protein chain in be ...
Extraction, Purification and Analysis of Anti cancer activity of Ricin
... Based on the above work different observations were made. Regarding the purification of the ricin protein, the maximum concentration of pure protein is obtained in the elute 3. When the elutes were tested against the Colon cancer cell lines, the observations showed that the ricin protein is capable ...
... Based on the above work different observations were made. Regarding the purification of the ricin protein, the maximum concentration of pure protein is obtained in the elute 3. When the elutes were tested against the Colon cancer cell lines, the observations showed that the ricin protein is capable ...
Overview of Protein Structure • The three
... among chemical polymers in that they adopt one folded conformation in solution. Steric repulsion between neighboring amino acids partially restricts the number of conformations accessible to a peptide chain. Weak noncovalent interactions among neighboring residues in the folded state determine the n ...
... among chemical polymers in that they adopt one folded conformation in solution. Steric repulsion between neighboring amino acids partially restricts the number of conformations accessible to a peptide chain. Weak noncovalent interactions among neighboring residues in the folded state determine the n ...
L -2 Sample preparation Before crystallization (first step
... Reducing agents are substances that cause other chemical species to be reduced or gain electrons. In order for reducing agents to cause the gaining of electrons on some other chemical species they must undergo oxidation and prevent the oxidation of free sulfhydryl residues (cysteines) in proteins. ...
... Reducing agents are substances that cause other chemical species to be reduced or gain electrons. In order for reducing agents to cause the gaining of electrons on some other chemical species they must undergo oxidation and prevent the oxidation of free sulfhydryl residues (cysteines) in proteins. ...
Salon service™
... distribution as hydrolyzed keratin obtained from wool or human hair. These low molecular weight proteins penetrate deep into the hair shaft to provide internal and external moisture while enhancing the hair’s healthy appearance and shine. Ammonium Hydroxide, Disodium Phosphate and Phosphoric Acid ad ...
... distribution as hydrolyzed keratin obtained from wool or human hair. These low molecular weight proteins penetrate deep into the hair shaft to provide internal and external moisture while enhancing the hair’s healthy appearance and shine. Ammonium Hydroxide, Disodium Phosphate and Phosphoric Acid ad ...
mirror of label in #2
... Whey is the preferred protein source in sports and bodybuilding nutrition because it contains superior quality Branched Chain Amino Acids — made up of Leucine, Isoleucine and Valine — which are important for the maintenance of muscle tissue.◊ Unlike some other incomplete protein sources, Body Fortre ...
... Whey is the preferred protein source in sports and bodybuilding nutrition because it contains superior quality Branched Chain Amino Acids — made up of Leucine, Isoleucine and Valine — which are important for the maintenance of muscle tissue.◊ Unlike some other incomplete protein sources, Body Fortre ...
Replicate OPM - MultiscaleLab
... Create a python module which given a PDB from the protein data bank returns a HTMD molecule which contains the PDB plus one or two slabs of dummy atoms representing the position of the membrane leaflet (similarly to OPM http://opm.phar.umich.edu/). Write a short report describing the algorithm used ...
... Create a python module which given a PDB from the protein data bank returns a HTMD molecule which contains the PDB plus one or two slabs of dummy atoms representing the position of the membrane leaflet (similarly to OPM http://opm.phar.umich.edu/). Write a short report describing the algorithm used ...
Bimolecular fluorescence complementation

Bimolecular fluorescence complementation (also known as BiFC) is a technology typically used to validate protein interactions. It is based on the association of fluorescent protein fragments that are attached to components of the same macromolecular complex. Proteins that are postulated to interact are fused to unfolded complementary fragments of a fluorescent reporter protein and expressed in live cells. Interaction of these proteins will bring the fluorescent fragments within proximity, allowing the reporter protein to reform in its native three-dimensional structure and emit its fluorescent signal. This fluorescent signal can be detected and located within the cell using an inverted fluorescence microscope that allows imaging of fluorescence in cells. In addition, the intensity of the fluorescence emitted is proportional to the strength of the interaction, with stronger levels of fluorescence indicating close or direct interactions and lower fluorescence levels suggesting interaction within a complex. Therefore, through the visualisation and analysis of the intensity and distribution of fluorescence in these cells, one can identify both the location and interaction partners of proteins of interest.