No Slide Title
... and assortative mating) – Population is large (if appropriately chosen!) – No mutation (but there is) – No migration (but migration occurs) – No selection (but there can be selection) ...
... and assortative mating) – Population is large (if appropriately chosen!) – No mutation (but there is) – No migration (but migration occurs) – No selection (but there can be selection) ...
lecture26_Polymorphi..
... this is the observed frequency distribution from the complete sequencing a large population; however most SNP discovery projects sequence a small population and then consider the absence or presence of those previously discovered SNPs in large population; this is known to under-estimate the number o ...
... this is the observed frequency distribution from the complete sequencing a large population; however most SNP discovery projects sequence a small population and then consider the absence or presence of those previously discovered SNPs in large population; this is known to under-estimate the number o ...
chapter 11.3 ppt note sheet
... 4. If genes don’t flow between populations what could happen to the species in two nearby populations? ...
... 4. If genes don’t flow between populations what could happen to the species in two nearby populations? ...
Conservation genetics premises
... Conservation biology premises, relevant to genetics (by the end of this course, you should be prepared to support or refute any of these) 1. Fitness is directly related to genetic variation 2. Genetic variation is critical for long-term survival of species 3. The goal of conservation biology is to p ...
... Conservation biology premises, relevant to genetics (by the end of this course, you should be prepared to support or refute any of these) 1. Fitness is directly related to genetic variation 2. Genetic variation is critical for long-term survival of species 3. The goal of conservation biology is to p ...
Chapter 17
... frequency and interaction of alleles and genes in populations. *Microevolution can be studied by observing changes in the numbers and types of alleles in populations. ...
... frequency and interaction of alleles and genes in populations. *Microevolution can be studied by observing changes in the numbers and types of alleles in populations. ...
Tracing the Paths of the First Americans
... ancestors of today’s Native Americans, stem and archaeological evidence to confrom a single Asian source population. But clude that these Beringian populations the data also suggest that this population may might have become genetically isolated have become genetically quite diverse during from main ...
... ancestors of today’s Native Americans, stem and archaeological evidence to confrom a single Asian source population. But clude that these Beringian populations the data also suggest that this population may might have become genetically isolated have become genetically quite diverse during from main ...
When the caste die was cast - National Institute of Biomedical
... They picked Khatris from northern India, Brahmins from Bengal and Gujarat, Iyers and other Dravidian speakers from southern India, Marathas from Maharashtra and several tribes from central and southern India ...
... They picked Khatris from northern India, Brahmins from Bengal and Gujarat, Iyers and other Dravidian speakers from southern India, Marathas from Maharashtra and several tribes from central and southern India ...
PowerPoint Chapter 15
... predicted distribution of alleles in populations; the central theorem of population genetics. Establishes a set of conditions in a population where ...
... predicted distribution of alleles in populations; the central theorem of population genetics. Establishes a set of conditions in a population where ...
Clive Beggs` presentation
... startling result. Fundamental similarities in mitochondrial DNA in living humans suggested that we all contain genetic material from a single woman who was living in Africa around 200,000 years ago. ...
... startling result. Fundamental similarities in mitochondrial DNA in living humans suggested that we all contain genetic material from a single woman who was living in Africa around 200,000 years ago. ...
Chapter 9 Maintenance of Genetic Diversity
... diversity differs between large and small populations. Selection has a major impact in large populations. However, its impacts are greatly reduced in small populations where genetic drift has an increasingly important role. ...
... diversity differs between large and small populations. Selection has a major impact in large populations. However, its impacts are greatly reduced in small populations where genetic drift has an increasingly important role. ...
Ch 23 Notes
... constant in generations – UNLESS acted upon by agents* other than Mendelian segregation and recombination of alleles. What *agents can cause the gene pool to change? ...
... constant in generations – UNLESS acted upon by agents* other than Mendelian segregation and recombination of alleles. What *agents can cause the gene pool to change? ...
Stamatis Konstantinos
... divergent between Greek and other European hares, foreign nuclear genes should not be a serious handicap. Hence, in certain situations releasing programs might be tolerated under obeying all other non-genetic strict ...
... divergent between Greek and other European hares, foreign nuclear genes should not be a serious handicap. Hence, in certain situations releasing programs might be tolerated under obeying all other non-genetic strict ...
15.2 PDQ - Biology with Radjewski
... 2. Explain, “natural selection acts on individuals, but populations evolve” • Changes that occur are developmental in a single organism over the course of a life cycle. • After breeding populations will evolve ...
... 2. Explain, “natural selection acts on individuals, but populations evolve” • Changes that occur are developmental in a single organism over the course of a life cycle. • After breeding populations will evolve ...
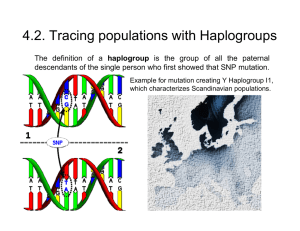
4.2. Tracing populations with Haplogroups
... Chronological development of mtDNA haplogroups U => 50,000 to 60,000 years ago (arose in Western Asia) H => 30,000 to 50,000 years ago (in the Near East - associated with Cro-Magnon in Europe) J => 45,000 years ago (in the Near East) X => over 30,000 years ago (in Caucasus) (Neanderthal???) I => 30 ...
... Chronological development of mtDNA haplogroups U => 50,000 to 60,000 years ago (arose in Western Asia) H => 30,000 to 50,000 years ago (in the Near East - associated with Cro-Magnon in Europe) J => 45,000 years ago (in the Near East) X => over 30,000 years ago (in Caucasus) (Neanderthal???) I => 30 ...
Analysis of Y chromosome lineages in native South American
... suggesting this value is not real. However at the Y-STRs level we found a small percentage of variation among populations which according to its associated P value ( P = 0.00098), seems to be real. When the five geographical regions are grouped into three populations, similar results are observed (T ...
... suggesting this value is not real. However at the Y-STRs level we found a small percentage of variation among populations which according to its associated P value ( P = 0.00098), seems to be real. When the five geographical regions are grouped into three populations, similar results are observed (T ...
Genetics and archaeogenetics of South Asia

The study of the genetics and archaeogenetics of the ethnic groups of South Asia aims at uncovering these groups' genetic history. The geographic position of India makes Indian populations important for the study of the early dispersal of all human populations on the Eurasian continent.According to the phylogeographic distribution of haplotypes observed among South Asian populations defined by social and linguistic criteria, the possibility arose of Y-DNA haplogroup F and mtDNA Haplogroup M might have originated in South Asia. The presence of several subclusters of F-M89 and K that are largely restricted to the Indian subcontinent is consistent with the scenario that a coastal (southern route) of early human migration out of Africa carried ancestral Eurasian lineages first to the coast of the Indian subcontinent, or that some of them originated there. Studies based on mtDNA variation have reported genetic unity across various Indian sub–populations. Conclusions of studies based on Y Chromosome variation and Autosomal DNA variation have been varied, although many researchers argue that most of the ancestral nodes of the phylogenetic tree of all the mtDNA types originated in the subcontinent. Recent genome studies appear to show evidence in support of the notion that modern south Asians (both Indo-Aryans and Dravidians) are a hybrid population descending from two genetically divergent populations referred to as the 'Ancestral North Indians' related to western eurasians and the 'Ancestral South Indians' who are not closely related to groups outside the subcontinent. It has been found that the ancestral node of the phylogenetic tree of all the mtDNA types typically found in Central Asia, the Middle East and Europe are also to be found in South Asia at relatively high frequencies. The inferred divergence of this common ancestral node is estimated to have occurred slightly less than 50,000 years ago. In India the major maternal lineages, or mitochondrial DNA Haplogroups, are M, R and U, whose coalescence times have been approximated to 50,000 BP.All major Y chromosome DNA haplogroups in the subcontinent are Haplogroup F's descendant haplogroups R (mostly R2a, R2 and R1a1), L, H and J (mostly J2). Many researchers have argued that Y-DNA Haplogroup R1a1 (M17) is of autochthonous Indian origin. However, proposals for a Central Asian origin for R1a1 are also quite common.