* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download IGEM2006-UCSF-Powerpoint
Survey
Document related concepts
E. coli long-term evolution experiment wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
Non-coding DNA wikipedia , lookup
Molecular cloning wikipedia , lookup
Molecular evolution wikipedia , lookup
Biosynthesis wikipedia , lookup
DNA supercoil wikipedia , lookup
Community fingerprinting wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Transformation (genetics) wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Transcript
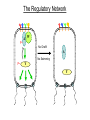
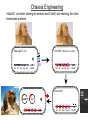
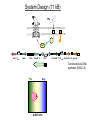
Remote Control of Bacterial Chemotaxis UCSF iGEM Team 2006 Patrick Visperas Matthew Eames Eli Groban Ala Trusina Christopher Voigt, Tanja Kortemme, Chao Tang, Chris Rao Goal Remote Control of a Bacterial Machine Start How Bacterial Chemotaxis Works Bacteria Swim Up Gradients How Chemotaxis is Observed Goulian Motility Assay (U Penn) Glide CCW CW concentration gradient No gradient ? Tumble The Regulatory Network A W P~ No CheW A No Swimming P~ Y Y Binding Partners •Orthogonal binding pair Tsr-CheW (Liu et al, 1991) •We mapped these mutations onto the Tar-Chew complex Tar CheW Tsr mutation can be mapped to Tar Gradient Aspartate • point mutation in Tar* reduces motility to approximately 40% that of wild-type Part Design • New orthogonal signaling pairs “Codon randomization” is used to reuse scaffolds without fear of recombination Chassis Engineering • Build E. coli strain lacking all sensors and CheW, but retaining the other chemotaxis proteins UU1250 (Parkinson, U Utah) Wild-type E. coli tar tsr tap trg aer cheW tar tsr tap trg aer cheW UUDcheW cheW* tar* tar tsr tap trg aer cheW Device Characterization: The Fim-Switch INPUT [ara] OUTPUT MCS Ptrc* rfp IRR IRL gfp FimE FimE IRL IRR T. Ham, et al (2006) Tested in UU1250 35 30 2 Possible Problems 25 1. The plasmid is unstable 20 15 2. The strain contains FimE/ FimB 10 5 RED + ara % bacteria - ara 0 0 0.3 1.2 5 1 2 3 4 5 6 7 8 GREEN [arabinose] (mM) System Design (11 kB) Phe CheY araC PBAD fimE T1x4 CheW r2 Ptrc* r2 CheW* T1 PlacIQ r0 phe-tar r0 asp-tar* Constructed via DNA synthesis (DNA 2.0) Phe Asp - arabinose System Design (11 kB) Asp CheY araC PBAD fimE T1x4 CheW r2 Ptrc* r2 CheW* T1 PlacIQ r0 phe-tar r0 asp-tar* Constructed via DNA synthesis (DNA 2.0) Phe Asp - arabinose Phe Asp + arabinose Results Gradient Gradient Aspartate Phenylalanine Conclusions • Orthogonal pairs are a rapid method to built signaling pathways that can operate simultaneously • Chassis developed to rapidly screen for new proteinprotein interactions • Limited by the switch performance in this chassis • Directional control could also be achieved by nonchemical inputs (light, radiation, etc)