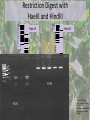
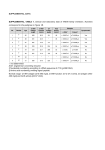
* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Biological Function of RMR2 in Maize: Genetic Study through
Genetically modified crops wikipedia , lookup
Chemical biology wikipedia , lookup
Bioinformatics wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
DNA vaccination wikipedia , lookup
Gene prediction wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Point mutation wikipedia , lookup
Molecular cloning wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Genetic engineering wikipedia , lookup
Designer baby wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
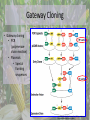
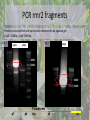
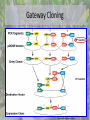
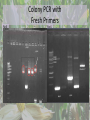
Biological Function of RMR2 in Maize: Genetic Study through Fluorescent Tagging of RMR2 Protein : Work In Progress Presented By Han Li Immediate Supervisor: Dr. Jacque Keele & Dr. Anding Luo Lab PI: Dr. Anne Sylvester Outline • Introduction • Experimental outline and procedures • Experimental results • Conclusion Central Dogma of Biology • DNA (deoxyribonucleic acid)- genetic code • TranscriptionDNAmRNA • TranslationmRNA protein This is followed by Mendelian inheritance New Discoveries in genetics • Silencing B-I* • Epigenetic regulation- heritable changes in gene function without a change in DNA sequence Genes can be regulated by: • • • • DNA Methylation Histone acetylation/deacetylation & methylation TGS and PTGS (Transcriptional Genomic Silencing and Post Transcriptional Genomic Silencing, Plants) dsRNA (RNAi) B’ X B-I* B-I*/B-I* B-I*/B’ B’ B-I*/B’ B’/B’ • Paramutation – An interaction between alleles of genes that leads to heritable changes in gene expression -Chandler, Vicki L. Cells (2007). 128: 641-645 -Griffiths AJF, Miller JH, Suzuki DT, et al. An Introduction to Genetic Analysis. 7th. 2000. -McGinnis KM et al., (2006) Genetics, 173:1637-1647. Role of RMR Protein • RMR2 (required to maintain repression 2) – Gene silencing the production of pigments – Epigenetics studies • Transcriptional gene silencing – Regulates RNA polymerase – DNA methylation – Mutation are defective in paramutations • RMR1 – Similar to RMR2 • Can be reversed by reintroducing the wild type protein McGinnis KM et al., (2006) Genetics, 173:1637-1647. Maize (Corn)- A common biological model used for genetic studies -Biello, David. That Burger You’re Eating Is Mostly Corn. Scientific American. Available from : http://www.scientificamerican.com/article.cfm?id=thatburger-youre-eating-is-mostly-corn --Corn. University of Illinois Extension: Watch your Garden Grow. Available from: http://urbanext.illinois.edu/veggies/corn.cfm -McGinnis KM et al., (2006) Genetics, 173:1637-1647. -ZeaMays. Wikipedia. Available from :http://en.wikipedia.org/wiki/File:ZeaMays.jpg RMR2 in Regulation of Gene Expression in Maize RMR2 RMR1 & RMR2 McGinnis KM et al., (2006) Genetics, 173:1637-1647. Aim of the project To study the role and expression of RMR2 on a molecular level – Fluorescent gene tagging • DNA sequence coding for fluorescent protein – Citrine yellow » Produced in the protein product Fluorescent marker Florescent tag Gene fragment 1 Gene fragment 2 Tether Protein product (rmr 2) Jackson, David. Sylvester, Anne. Chan, Agnes. Etc. Maize Cell genomics DB (2010). Available from: http://maize.jcvi.org/cellgenomics/ Experimental outline • Use Multi site Gateway cloning to make the construct – – – – – Primer design Amplification of Rmr2 DNA fragments BP reaction Sequencing LR reaction • Introduce the construct into Agrobacterium tumefaciens • Transform maize at the Plant Transformation Facility (ISU) – Stable transgenic plant Gateway Cloning • Gateway cloning • PCR (polymerase chain reaction) • Plasmids • Special flanking sequences User Manual for MultiSite Gateway Pro (2006) available from: http://www.invitrogen.com/vntigateway Designing primers for Gateway cloning • Determining the insertion position – The best position to insert the fluorescent protein is right after the start codon • Should incorporate regulatory regions – 3000 bp upstream – 800 bp downstream Promoter tag Terminator • Primer 3 (http://frodo.wi.mit.edu/) – p1: GGGG ACA AGT TTG TAC AAA AAA GCA GGC Tct agc cac ttg gct gta ctg tg – p4: GGGG AC AAC TTT GTA TAG AAA AGT TGG GTG cat ggt acc ggc ggt ctt gg – p3: GGGG ACA ACT TTG TAT AAT AAA GTT GAG ccg gtc ctc cgc tcc ccg t – p2: GGGG AC CAC TTT GTA CAA GAA AGC TGG GTA ctt gcc ggt gca gta gag tt Steve Rozenand Helen J. Skaletsky. Primer3. (2000) available at http://fokker.wi.mit.edu/primer3/ PCR rmr2 fragments Products are amplified and results were observed on an agarose gel: p1p4~ 3200bp p2p3~3800bp Fig 1 p1p4 Fig 2 p2p3 p2p3 4000 3000 4000 3000 p1 p4 Fluorescent tag p2 p3 Gateway Cloning User Manual for MultiSite Gateway Pro (2006) available from: http://www.invitrogen.com/vntigateway Gel purification and BP reaction Fig 3 P2P3 P1P4 •Bands were cut out and gel purified. 4000 3000 •1 µL of each product was loaded on a gel •BP reaction was conducted to introduce the DNA fragments into the pDONOR vector. •Kanamycin resistance •The vectors are then introduced into the kanamycin sensitive bacteria (E. coli) through transformation. •Heat shock Colony PCR • Bacteria were plated on kanamycin plates and were grown at 37 ◦C overnight. • Colonies are then subjected to colony PCR – Amplify the DNA sequences in the plasmid – Plasmid specific primers Fig 3 Fig 4 Samples 1-12 p1p4 Control “-” Samples 13-23 p1p4 Colony PCR with Fresh Primers Fig 5 Fig 6 p2p3 p1p4 5000 4000 3000 5000 4000 3000 Control “+” p2p3 p1p4 Restriction Digest with HaeIII and HindIII site2 HaeIII HaeIII HaeIII HaeIII HaeIII HaeIII End HaeIII HaeIII HaeIII HaeIII 2581 1172 2933 667 213 359 3118 2990 2955 376 229 Mass % 45 16 11 9 7 4 4 1 1 1 1 New DNANew DNA Size site1 Size site1 1 Start Start HaeIII 1172 2131409 229 HaeIII 3470 359505 HindIII 376 HaeIII 429 HaeIII 667 HindIII HaeIII 2581 667352 HaeIII 6 291 HaeIII 376 212 1 1172 HaeIII Start 130 HaeIII 229 128 HaeIII 2990 35 HaeIII 2955 22 HaeIII 2933 17 HaeIII 359 16HaeIII HaeIII 213 2581 site2 1 HaeIII 3471 HaeIII 3900 HaeIII HaeIII HaeIII HaeIII End HaeIII HaeIII HaeIII HaeIII Hae III 2933 HaeIII 2955 2990 3118 End site2 HindIII 2581 HindIII 1172 End 2933 667 213 359 3118 2990 2955 376 229 New DNA Size site1 % 1 Start Mass % Mass 1 Start Start 3471 89 HaeIII 3470 45 213 229 HindIII 359 3900 11 HaeIII 429 16 376 HindIII 3906 0 HaeIII 6 11 667 9 7 1172 HaeIII 4 4 1 1 1 1 2581 HaeIII 3471 HindIII 2933 HaeIII 2955 2990 3900 End HindIII 3906 3118 End 1 3471 3900 site2 HindIII HindIII End Hind III 3471 3900 3906 Mass % 1 Start 89 11 0 3471 HindIII 3900 End HindIII 3906 4000 3000 1000 400 P2P3 P1P4 Davis, M. Wayne. APE. Version 2.0.44. 12 from: http://biologylabs.utah. edu/jorgensen/wayne d/ape/ Work Progress and Conclusion All steps prior to the BP reaction were successful Bacteria have the plasmid Survived on the kanamycin plates Indicated that plasmid is present Most of the bands are smaller than expected Restriction digest confirmed that fragments of Rmr2 were inserted Possible reasons: Recombination alteration Bacteria excreting plasmids out Dimers Maize sequence repeats Future Trouble Shooting • • • Use a different bacteria vector Utilize only PCR reactions New primers • Avoid Maize genome issues: common repeats, GC rich (different physical structure) Acknowledgment Special thanks to: Dr. Anne Sylvester Dr. Jacque Keele Dr. Anding Luo Rest of Sylvester’s Lab EPSCoR NSF Questions? Work Cited • • • • • • • • • • • • Biello, David. That Burger You’re Eating Is Mostly Corn[internet]. Scientific American. [12 November 2008; 29 March 2012]. Available from : http://www.scientificamerican.com/article.cfm?id=that-burger-youre-eating-is-mostly-corn Chandler, Vicki L. Paramutation: From Maize to Mice. Cells [internet]. [2007; cited 12 April 2012]. 128: 641-645. Available from: ftp://ftpmips.gsf.de/plasmar/epigenetics/Cell%20review%20Issue/Paramutation%20%20From%20Maize%20to%20Mice.pdf Commonly Used Promoters [internet]. African Biosafety Network Expertise. [2010; cited 29 March 2012] available from :http://www.nepadbiosafety.net/for-regulators/resources/subjects/biotechnology/commonly-used-promoters Corn [internet]. University of Illinois Extension: Watch your Garden Grow. [2012: 10 April 2012]. Available from: http://urbanext.illinois.edu/veggies/corn.cfm Corn and Sorgnum Pictures/corn ears [internet]. Texas A&M. [1 March 2002; cited 29 March 2012]. Available from: http://soilcrop.tamu.edu/photogallery/cornsorghum+/pages/corn%20ears.htm Davis, M. Wayne. APE [internet download]. Version 2.0.44. 12 April 2012. Available from: http://biologylabs.utah.edu/jorgensen/wayned/ape/ Jackson, David. Sylvester, Anne. Chan, Agnes. Etc. Characterizing Sub-cellular Compartments in Maize Using Fluorescent Protein Tagging Lines. Maize Cell genomics DB. [18 December 2010; cited 20 April 2012]. Available from: http://maize.jcvi.org/cellgenomics/ McGinnis KM, Springer C, Lin Y, Carey CC, Chandler V (2006) Transcriptionally silenced transgenes in maize are activated by three mutations defective in paramutation. Genetics, 173:1637-47. Griffiths AJF, Miller JH, Suzuki DT, et al. An Introduction to Genetic Analysis. 7th. New York: W. H. Freeman; 2000. Steve Rozenand Helen J. Skaletsky (2000) Primer3 on the WWW for general users and for biologist programmers.In: Krawetz S, Misener S (eds)Bioinformatics Methods and Protocols: Methods in Molecular Biology.Humana Press, Totowa, NJ, pp 365-386 Source code available at http://fokker.wi.mit.edu/primer3/ User Manual for MultiSite Gateway Pro (Invitrogen 2006) Using gateway technology to simultaneously clone multiple DNA fragments. Available from: http://www.invitrogen.com/vntigateway ZeaMays [Internet]. Wikipedia. [2012; cited 29 March 2012]. Available from :http://en.wikipedia.org/wiki/File:ZeaMays.jpg