* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Ch.05The Structure and Function of Large Biological Molecules
Protein–protein interaction wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Gene expression wikipedia , lookup
Citric acid cycle wikipedia , lookup
Photosynthetic reaction centre wikipedia , lookup
Epitranscriptome wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Genetic code wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Blood sugar level wikipedia , lookup
Peptide synthesis wikipedia , lookup
Fatty acid synthesis wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Phosphorylation wikipedia , lookup
Point mutation wikipedia , lookup
Fatty acid metabolism wikipedia , lookup
Proteolysis wikipedia , lookup
Metalloprotein wikipedia , lookup
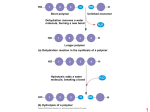
1 2 3 H Short polymer HO Dehydration removes a water molecule, forming a new bond HO 2 1 H Trioses (C3H6O3) Pentoses (C5H10O5) Hexoses (C6H12O6) Unlinked monomer 3 H 2O 4 Aldoses HO H Glyceraldehyde Longer polymer (a) Dehydration reaction in the synthesis of a polymer Ribose Glucose 1 2 3 4 Hydrolysis adds a water molecule, breaking a bond HO 1 2 3 H H 2O HO Galactose H Ketoses HO Dihydroxyacetone H Ribulose Fructose (b) Hydrolysis of a polymer 1 2 1–4 glycosidic linkage Glucose Glucose Maltose (a) Dehydration reaction in the synthesis of maltose 1–2 glycosidic linkage (a) Linear and ring forms (b) Abbreviated ring structure Glucose Fructose Sucrose (b) Dehydration reaction in the synthesis of sucrose 3 Chloroplast 4 Mitochondria Glycogen granules Starch (a) α and β glucose ring structures 0.5 µm α Glucose β Glucose 1 µm (b) Starch: 1–4 linkage of α glucose monomers Glycogen Amylose (b) Cellulose: 1–4 linkage of β glucose monomers Amylopectin (a) Starch: a plant polysaccharide (b) Glycogen: an animal polysaccharide 5 6 Cell walls Cellulose microfibrils in a plant cell wall Microfibril 10 µm 0.5 µm Cellulose molecules β Glucose monomer 7 8 Fatty acid (palmitic acid) Glycerol (a) Dehydration reaction in the synthesis of a fat Ester linkage (b) Chitin forms the exoskeleton of arthropods. (c) Chitin is used to make a strong and flexible surgical thread. (b) Fat molecule (triacylglycerol) 9 Hydrophilic head Structural formula of a saturated fat molecule 10 Stearic acid, a saturated fatty acid (a) Saturated fat Structural formula of an unsaturated fat molecule Hydrophobic tails (a) The structure of the chitin monomer. Oleic acid, an unsaturated fatty acid (a) Structural formula Choline Phosphate Glycerol Fatty acids Hydrophilic head Hydrophobic tails (b) Space-filling model (c) Phospholipid symbol cis double bond causes (b) Unsaturated fat bending 11 12 Hydrophilic head Hydrophobic tail WATER WATER 13 14 Substrate (sucrose) Glucose Enzyme (sucrase) OH H2O Fructose H O 15 16 Nonpolar α carbon Glycine (Gly or G) Valine (Val or V) Alanine (Ala or A) Methionine (Met or M) Leucine (Leu or L) Trypotphan (Trp or W) Phenylalanine (Phe or F) Isoleucine (Ile or I) Proline (Pro or P) Polar Serine (Ser or S) Threonine (Thr or T) Cysteine (Cys or C) Tyrosine (Tyr or Y) Asparagine Glutamine (Asn or N) (Gln or Q) Electrically charged Amino group Acidic Carboxyl group Aspartic acid Glutamic acid (Glu or E) (Asp or D) 17 Basic Lysine (Lys or K) Arginine (Arg or R) Histidine (His or H) 18 Peptide bond (a) Groove Groove Side chains Peptide bond (a) A ribbon model of lysozyme (b) A space-filling model of lysozyme Backbone (b) Amino end (N-terminus) Carboxyl end (C-terminus) 19 Antibody protein 20 Protein from flu virus Primary Structure Secondary Structure Tertiary Structure Quaternary Structure β pleated sheet +H N 3 Amino end Examples of amino acid subunits α helix 21 Hydrophobic interactions and van der Waals interactions 22 Polypeptide chain β Chains Polypeptide backbone Hydrogen bond Disulfide bridge Iron Heme α Chains Hemoglobin Ionic bond Collagen 23 24 Normal hemoglobin Primary structure 1 2 3 4 5 6 7 Secondary and tertiary structures Sickle-cell hemoglobin Primary structure Val His Leu Thr Pro Glu Glu β subunit Val His Leu Thr Pro Val Glu 1 2 3 4 5 6 7 Secondary and tertiary structures Exposed hydrophobic region β subunit Denaturation Normal hemoglobin (top view) Quaternary structure β Function α β α Quaternary structure β Sickle-cell hemoglobin α β α Function Molecules do not associate with one another; each carries oxygen. Molecules interact with one another and crystallize into a fiber; capacity to carry oxygen is greatly reduced. Normal protein 10 µm Red blood cell shape Normal red blood cells are full of individual hemoglobin moledules, each carrying oxygen. Denatured protein Renaturation 10 µm Red blood cell shape Fibers of abnormal hemoglobin deform red blood cell into sickle shape. 25 26 DNA 1 Synthesis of mRNA in the nucleus Correctly folded protein Polypeptide mRNA Cap NUCLEUS CYTOPLASM Hollow cylinder mRNA Chaperonin (fully assembled) Steps of Chaperonin 2 The cap attaches, causing the 3 Action: cylinder to change shape in such a way that it creates a 1 An unfolded polyhydrophilic environment for peptide enters the cylinder from one end. the folding of the polypeptide. 2 Movement of mRNA into cytoplasm via nuclear pore The cap comes off, and the properly folded protein is released. Ribosome 3 Synthesis of protein Amino acids Polypeptide 27 28 5' end 5ʹ′ end Nitrogenous bases Pyrimidines 5ʹ′C Sugar-phosphate backbones 3ʹ′C Base pair (joined by hydrogen bonding) Nucleoside Nitrogenous base Cytosine (C) Thymine (T, in DNA) Uracil (U, in RNA) Old strands Purines Phosphate group 5ʹ′C (b) Nucleotide 3ʹ′C 3' end Nucleotide about to be added to a new strand Sugar (pentose) 3' end Adenine (A) Guanine (G) Sugars 3ʹ′ end 5' end (a) Polynucleotide, or nucleic acid New strands Deoxyribose (in DNA) Ribose (in RNA) 5' end 3' end (c) Nucleoside components: sugars 5' end 29 3' end 30 31