* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Biochemistry of Sulfur
Artificial gene synthesis wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Catalytic triad wikipedia , lookup
Signal transduction wikipedia , lookup
Biosynthesis wikipedia , lookup
Magnesium transporter wikipedia , lookup
Photosynthesis wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Western blot wikipedia , lookup
Gaseous signaling molecules wikipedia , lookup
Light-dependent reactions wikipedia , lookup
Electron transport chain wikipedia , lookup
Photosynthetic reaction centre wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
Sulfur dioxide wikipedia , lookup
Sulfur cycle wikipedia , lookup
Microbial metabolism wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
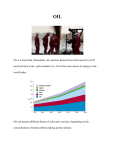
PD Dr. Arnulf Kletzin Institut für Mikrobiologie und Genetik TU Darmstadt Schnittspahnstraße 10 64287 Darmstadt Germany Sulfur Oxidation and Reduction in Acidianus ambivalens Numerous (micro-) organisms oxidize sulfur (S˚) and inorganic sulfur compounds (ISCs) for energy conservation. The oxidation of S˚ to sulfuric acid proceeds in at least two steps, often more involving intermediates like sulfite, thiosulfate, tetrathionate, etc. (Fig. 1). Thermoacidophilic Archaea oxidize S˚ with a cytoplasmic sulfur-disproportionating enzyme. The products thiosulfate and sulfite are oxidized by membrane-bound oxidoreductases. Figure 1: 1: The The biological biological sulfur sulfur cycle cycle Figure shown by by reactions reactions involving involving inorganic inorganic shown sulfur compounds. compounds. Enzymes: Enzymes: 1, 1, polypolysulfur sulfide reductase; reductase; 2, 2, sulfur sulfur reductase; reductase; 3, 3, sulfide sulfide: quinone quinone oxidoreductase oxidoreductase or or sulfide: sulfide:cytochrome cc oxidoreductase; oxidoreductase; 4, 4, sulfide:cytochrome sulfur oxygenase; oxygenase; 5, 5, sulfite: sulfite: acceptor acceptor sulfur oxidoreductase; 6, 6, ATP ATP sulfurylase; sulfurylase; 7, 7, ATP ATP oxidoreductase; sulfurylase or or adenylylsulfate: adenylylsulfate: phosphate phosphate sulfurylase adenylyltransferase; 8, 8, adenylylsulfate adenylylsulfate adenylyltransferase; (APS) reductase; reductase; 9, 9, sulfite sulfite reductase; reductase; 10, 10, (APS) tetrathionate reductase; reductase; 11, 11, thiosulfate: thiosulfate: tetrathionate acceptor oxidoreductase; oxidoreductase; 12, 12, Sox Sox acceptor complex; 13, 13, sulfur sulfur oxygenase oxygenase reductase; reductase; complex; 14, thiosulfate thiosulfate reductastase; reductastase; 15, 15, 14, tetrathionate hydrolase hydrolase (there (there is is also also aa tetrathionate trithionate hydrolase hydrolase in in addition); addition); 16, 16, OOtrithionate acetylserin or or O-phosphoserine O-phosphoserine sulfhydrosulfhydroacetylserin lases; 17, 17, cysteine cysteine desulfurase. desulfurase. Gray Gray lases; circles denote denote disproportionation disproportionation reactreactcircles ions. Reactions Reactions of of cysteine cysteine breakdown breakdown in in ions. eukaryotes and and of of phosphoadenosinphosphoadenosineukaryotes phosphosulfate formation formation and and reduction reduction phosphosulfate are omitted. omitted. are Sulfur Oxidation, Sulfur Oxygenase Reductase: The initial enzyme in the S˚ oxidation pathway is unique in several aspects. The soluble protein was termed sulfur oxygenase reductase (SOR) because it is the only S˚-disproportionating enzyme. Sulfite, thiosulfate and hydrogen sulfide are products. Oxygen is required for activity but no organic cofactors: (Eq. 1) 4 S˚ + O2 + 4 H2O ---- > 2 HSO3- + 2 H2S + 2 H+ Optimal activity was observed at 85 ˚C and pH 7, maximal activity at 108 ˚C. Three conserved cysteine residues are present in the known SORs. Site-directed mutagenesis showed that one of these was indispensable, while mutagenesis of the other two resulted in reduced activities. EPR spectroscopy and redox titration showed that the SOR contains a mononuclear non-heme iron center in the high-spin Fe3+ state and with a low reduction potential (Eo' = -268 mV). It was intriguing to find that the reduction potential was more than 300 mV lower than usually found for this type of iron centers and low enough to explain the S˚ reducing activity of the enzyme (Eo' [H2S/S˚] = -270 mV). Figure 2: Electron Micrograph of the SOR (Reinhard Rachel, Regensburg) Figure 3: Molecular model of the SOR. A, Cartoon representation with α-helical regions in blue and βsheet in violett, Fe atom red balls. B, Surface representation with the subunits coloured differently. C, Section with the surface coloured according to charge (blue=positive, red = negative). SOR structure and mechanism: Hollow globular particles of 15.5 nm in diameter appeared in electron microscopic pictures of the purified Ac. ambivalens SOR (Fig. 2). X-ray crystallographic analysis to 1.7 Å resolution showed that the SOR is a spherical homo-icosatetramer (i.e. 24 subunits). It surrounds an empty cavity with a diameter of 71-107 Å Fig. (Fig. 3) and a molecular mass of 844.000 for the native SOR (871.000 for the recombinant). Each subunit consisted of a β-barrel core surrounded by α-helices. The iron center was coordinated by two histidine ligands, a bidentate glutamate and two water molecules in a structural motif, which is known as "2-His 1-Carboxylate facial triad" (Fig. 4). Mutation of any of the three iron ligands to alanine resulted in the complete loss of activity and iron-binding capabilities. The residue C31 was modified to a persulfide. The iron center and the three cysteines are buried in a pocket in the interior of each monomer. Some conclusions for the reaction mechanism could be derived from the structure. The core active site is composed of the iron site and the modified C31. It is accessible only from the interior cavity. Substrate entry has to proceed through the hydrophobic channels along the 4-fold axes of the sphere (Fig. 2). S˚ is most probably bound covalently to C31. The linear sulfur chain is aligned to the iron site and replaces the water ligands, poising the iron site for oxygen binding and activation. Figure 4: The catalytic pocket of the SOR containing the conserved cysteines and the iron. (A) Stereo view of the mononuclear non-heme iron center with Fe ligands, Fe ion; red, waters; ball-and-stick, protein ligands. (B) Cavity surface representation of the catalytic pocket with Cys and Fe highlighted; gray arrow, cavity entrance. (C) Identification of an additional sulfur atom at Cys31. Seite: 2/6 Thiosulfate Oxidation: A tetrathionate-forming, membrane-bound thiosulfate:quinone oxidoreductase (TQO) was isolated from aerobically grown Ac. ambivalens cells. Optimal activity was observed at 85 ˚C and pH 5. The 102 kDa glycosylated holoenzyme had a α2β2 stoichiometry. Oxygen electrode measurements showed an electron transport from thiosulfate to molecular oxygen via the terminal heme copper quinol:oxygen oxidoreductase (Fig. 5). The fate of the tetrathionate formed by the Ac. ambivalens TQO has not been investigated yet when the organism grows on S˚. However, there is a possibility that a thiosulfate/tetrathionate cycle exists. Tetrathionate is unstable in the presence of strong reductants and is reduced to thiosulfate in vitro at high temperatures. Therefore, H2S and sulfite might re-reduce tetrathionate formed by the TQO and thus feed electrons indirectly from the S˚ disproportionation reaction catalyzed by the SOR into the quinone pool (Fig. 5). Figure 5: Hypothetical model of S˚ oxidation in A. ambivalens derived from known enzyme activities (in italics) and possible non-enzymic reactions: SAOR, sulfite:acceptor oxidoreductase; TQO, thiosulfate:quinone oxidoreductase; SOR, sulfur oxygenase reductase; APS, adenylylsulfate; APAT, adenylylsulfate:phosphate adenylyltransferase, CQ, caldariella quinone. Seite: 3/6 Anaerobic Sulfur Reduction with Hydrogen as Electron Donor: Many microorganisms reduce sulfur when growing anaerobically. Several metabolic types with S˚ as electron acceptor can be distinguished. Many heterotrophs require the addition of S˚ to the media as terminal electron acceptors. The products are either CO2 or small organic compounds like acetate. H2S is produced in the presence and H2 in the absence of S˚. Alternatively, numerous S˚ reducers like Acidianus ambivalens gain energy from H2 oxidation; ATP and reduction equivalents are used for CO2 fixation: H2 + S˚ ---- > H2S (or HS-, depending on the pH) Acidianus ambivalens sulfur reductase and hydrogenase: A sulfur reductase (SR; H2:S˚ oxidoreductase) purified from membrane fractions of anaerobically grown Ac. ambivalens cells showed activity in the presence of a co-purified hydrogenase. The A. ambivalens hydrogenase encoded by a polycistronic cluster including genes for a NiFe and an FeS subunit was rather dissimilar to other hydrogenases (Fig. 6). The Isp1 protein is probably the membrane anchor. The FeS subunit contained a leader peptide with a twin-arginine translocation motif (TAT) not present in the mature protein suggesting a transport across the membrane (Fig. 7). Fe and Ni were present in membrane and in enriched hydrogenase fractions in accordance with the observed sequence similarity. The Ac. ambivalens SR gene cluster consisted of five ORFs, sreABCDE (Fig. 6). The deduced amino acid sequences of sreA encoding a 110 kDa subunit visible on SDS gels and sreB showed similarity to the molybdopterin and the FeS subunits of the DMSO/Nitrate reductase family, respectively. SreA also contained a twin arginine motif suggesting an export by the TAT pathway. The sreC gene encoded a protein with 10 predicted transmembrane helices, sreD an unknown polyferredoxin with 26 cysteine residues and sreE a small system-specific chaperone. Molybdenum, but not tungsten was found in solubilized membrane fractions suggesting that the SR is a molybdoprotein in accordance with the observed sequence similarity. Figure 6: Gene clusters encoding the NiFe hydrogenase and the SR in Ac. ambivalens. Small black circles indicate cysteinecontaining sequence motifs potentially coordinating FeS clusters; Ni and Mo indicate potential metal binding sites; m, predicted transmembrane protein. RR twin arginine protein translocation motif, the tat cluster encodes proteins required for twin arginine translocation pathway; HynS, HynL, small and large subunit of the hydrogenase; isp1, membrane anchor; hyp, hydrogenase maturation genes/proteins; hoxM, protease of HynL maturation. Seite: 4/6 The predicted orientation and the molecular composition of the Ac. ambivalens SR deduced from the biochemical results and the sequence analysis are similar to the W. succinogenes PSR (Fig. 7). Both enzymes consist of homologous catalytic and electron transfer subunits (SreA/PsrA and SreB/PsrB, respectively) and non-homologous membrane anchors (SreC/PsrC). The hydrogenases also have comparable quaternary structures; they are composed of at least three subunits each, the homologous Ni-containing catalytic (HynL/HynB; Fig. 7), the FeS-containing electron transfer subunits (HynS/HynA) and the non-homologous membrane anchors (Isp1/HynC). The only other probably orthologous sreABCDE gene clusters have been identified in the genomes of Su. solfataricus and Su. islandicus (but not in Su. acidocaldarius and Su. tokodaii) suggesting that they should grow by heterotrophic anaerobic S˚ respiration; however, this has not been demonstrated yet. Phylogenetic analysis showed that other homologs of SreA are present in the genomes of other facultative sulfur reducers and that they form a branch in the DMSO reductase family tree together with other enzymes with unrelated substrates. It can be concluded that these organisms have a mutually similar set of membrane-bound SRs with related but diverse molybdopterin and electron transfer subunits while additional subunits might vary. The membrane-bound hydrogenases diversify in a similar way, although the analysis is more complicated because NiFe hydrogenases fall into four subfamilies, which are all realized in Archaea. It was interesting to see that the Sulfolobus species sequenced so far do not have hydrogenase genes, showing that they should not be able to grow lithotrophically with H2. Figure 7: Schematic representation of the sulfur and polysulfide respiration in Ac. ambivalens and Wolinella succinogenes. The Ac. ambivalens model was developed in analogy to Wo. succinogenes from the results of the sequence comparison and the biochemical data. Homologous subunits are shown in identical shading. CM cytoplasmic membrane, OM, outer membrane, MQ, menaquinone, SQ, sulfolobus quinone. Seite: 5/6