* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Gut Microbiota_ The Conductor in the Orchestra of Immune
Survey
Document related concepts
DNA vaccination wikipedia , lookup
Sociality and disease transmission wikipedia , lookup
Social immunity wikipedia , lookup
Transmission (medicine) wikipedia , lookup
Molecular mimicry wikipedia , lookup
Adoptive cell transfer wikipedia , lookup
Adaptive immune system wikipedia , lookup
Polyclonal B cell response wikipedia , lookup
Immune system wikipedia , lookup
Inflammatory bowel disease wikipedia , lookup
Schistosoma mansoni wikipedia , lookup
Cancer immunotherapy wikipedia , lookup
Immunosuppressive drug wikipedia , lookup
Innate immune system wikipedia , lookup
Transcript
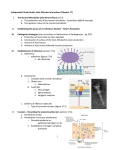
University of Groningen Gut Microbiota: The conductor in the Orchestra of immune-neuroendocrine communication El Aidy, Sahar Farouk Abdelsalam; Dinan, Timothy G; Cryan, John F Published in: Clinical Therapeutics DOI: 10.1016/j.clinthera.2015.03.002 IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it. Please check the document version below. Publication date: 2015 Link to publication in University of Groningen/UMCG research database Citation for published version (APA): El Aidy, S., Dinan, T. G., & Cryan, J. F. (2015). Gut Microbiota: The conductor in the Orchestra of immuneneuroendocrine communication. Clinical Therapeutics, 37(5), 954-67. DOI: 10.1016/j.clinthera.2015.03.002 Copyright Other than for strictly personal use, it is not permitted to download or to forward/distribute the text or part of it without the consent of the author(s) and/or copyright holder(s), unless the work is under an open content license (like Creative Commons). Take-down policy If you believe that this document breaches copyright please contact us providing details, and we will remove access to the work immediately and investigate your claim. Downloaded from the University of Groningen/UMCG research database (Pure): http://www.rug.nl/research/portal. For technical reasons the number of authors shown on this cover page is limited to 10 maximum. Download date: 12-06-2017 Clinical Therapeutics/Volume 37, Number 5, 2015 Review Article Gut Microbiota: The Conductor in the Orchestra of Immune–Neuroendocrine Communication Sahar El Aidy, PhD1,2; Timothy G. Dinan, PhD1,3; and John F. Cryan, PhD1,4 1 Laboratory of Neurogastroenterology, Alimentary Pharmabiotic Centre, University College Cork, Cork, Ireland; 2Department of Industrial Biotechnology, Genetic Engineering and Biotechnology Research Institute, Sadat City University, Sadat City, Egypt; 3Department of Psychiatry, University College Cork, Cork, Ireland; and 4Department of Anatomy and Neuroscience, University College Cork, Cork, Ireland ABSTRACT Purpose: It is well established that mammals are socalled super-organisms that coexist with a complex microbiota. Growing evidence points to the delicacy of this host–microbe interplay and how disruptive interventions could have lifelong consequences. The goal of this article was to provide insights into the potential role of the gut microbiota in coordinating the immune–neuroendocrine cross-talk. Methods: Literature from a range of sources, including PubMed, Google Scholar, and MEDLINE, was searched to identify recent reports regarding the impact of the gut microbiota on the host immune and neuroendocrine systems in health and disease. Findings: The immune system and nervous system are in continuous communication to maintain a state of homeostasis. Significant gaps in knowledge remain regarding the effect of the gut microbiota in coordinating the immune–nervous systems dialogue. Recent evidence from experimental animal models found that stimulation of subsets of immune cells by the gut microbiota, and the subsequent cross-talk between the immune cells and enteric neurons, may have a major impact on the host in health and disease. Implications: Data from rodent models, as well as from a few human studies, suggest that the gut microbiota may have a major role in coordinating the communication between the immune and Accepted for publication March 4, 2015. http://dx.doi.org/10.1016/j.clinthera.2015.03.002 0149-2918/$ - see front matter & 2015 Elsevier HS Journals, Inc. All rights reserved. 954 neuroendocrine systems to develop and maintain homeostasis. However, the underlying mechanisms remain unclear. The challenge now is to fully decipher the molecular mechanisms that link the gut microbiota, the immune system, and the neuroendocrine system in a network of communication to eventually translate these findings to the human situation, both in health and disease. (Clin Ther. 2015;37:954–967) & 2015 Elsevier HS Journals, Inc. All rights reserved. Key words: brain, enteric nervous system, gut microbiota, immune system. INTRODUCTION Our knowledge of the host–microbe interrelationship is accelerating because of the availability of rapidly expanding molecular techniques, especially in combination with the use of reductionist in vivo host models. A growing body of research continues to show that the normal mammalian structure and function are significantly dependent on their constant engagement in complex interactions with microbes.1 For example, the intestinal microbiota with its components and metabolites affects the host physiology in various ways to control energy homeostasis, gut barrier function, mucosal inflammation, and behavior.2–4 Subsequently, the host modifies many of the microbial activities, which Scan the QR Code with your phone to obtain FREE ACCESS to the articles featured in the Clinical Therapeutics topical updates or text GS2C65 to 64842. To scan QR Codes your phone must have a QR Code reader installed. Volume 37 Number 5 S. El Aidy et al. suggests a feedback mechanism that could influence the microbiota and drive a further cycle of biological changes to host physiology.5 This multidirectional interactive dialogue seems to strongly influence an expanding repertoire of human disorders, including obesity, depression, and irritable bowel syndrome; the goal is to decipher the tête-à-tête between the host and its commensal organisms.6,7 The present review highlights emerging evidence, which provides a framework for appreciating the impact of the host–microbiota interplay on health that is undoubtedly much broader than previously thought. This review outlines the impact of the gut microbiota on the immune system as well as the neuroendocrine system, both of which point to a potential role of the microbiota as crucial coordinators in the cross-talk between these systems. MATERIALS AND METHODS Literature from a range of sources, including PubMed, Google Scholar, and MEDLINE, was searched to identify recent reports on the impact of the gut microbiota on the host immune and neuroendocrine systems in health and disease. All data were gathered for the latest available years. RESULTS The Host–Microbiota Dialogue Over the past decade, remarkable advances toward a better understanding of how the host–microbe interactome is linked with most pathways that affect health, disease, and aging were made possible with novel technologies; these include high-throughput DNA sequencing, bioinformatics, and gnotobiotic animal models.1 The intimate interactions between the host and its microbes that outnumber the host’s cells by 10 to 1 are required to stimulate the complete maturation of an efficient intestinal barrier that can promote niche colonization by commensals and opposes colonization by pathogens.4,8 Only microbial populations that are capable of establishing a mutualistic relation with the host can be maintained in the gut ecosystem,3 creating a habitat that exerts restrictive selection on its microbial inhabitants. This restrictive selection of specific microbial groups is illustrated by the relatively low phylum-level diversity observed in the microbiota of the gastrointestinal (GI) tract of many mammalian organisms, including mice and humans; this is dominated by the phyla of the May 2015 Bacteroidetes and the Firmicutes, with Proteobacteria, Actinobacteria, Fusobacteria, and Verrucomicrobia phyla present in relatively low abundance.9 In contrast, the diversity at the genus and species levels is enormous. Advances in metagenomic approaches helped to illustrate that despite the variation in species composition, the microbial communities encompass a relatively similar set of metabolic functions in healthy individuals, which are referred to as the “core microbiome.”10 Furthermore, diet and its nutritional value are partly shaped by (and they in turn can shape) the gut microbial community, supporting the notion that “we are what we eat,” a process that starts early in life.11,12 Indeed, this scenario is now being exploited for medical innovations along the diet–host interface in line with Hippocrates’ dictum that “Let food be thy medicine and medicine be thy food.” The primary individual microbiota colonize at birth. However, there is growing evidence that the in utero environment may not be sterile, as originally thought: bacteria such as Enterococcus faecalis, Staphylococcus epidermidis, and Escherichia coli have been isolated from the meconium of healthy neonates.13 The infant gut microbiota is more variable in its composition and less stable over time. In the first year of life, the infant intestinal tract progresses from sterility to extremely dense colonization, ending with a mixture of microbes that is relatively stable and largely similar to that found in the adult intestine.14,15 The composition of the gut microbiota is shaped by a variety of factors, including prenatal and postnatal variables as well as birth factors.16 Altering the intestinal microbiota, particularly during early life, has been shown to have lasting consequences on the host.17 For example, the variation in early-life environmental exposure that includes the method and place of delivery and a nourishing neonate regimen influences the microbiota of infants.18,19 The microbiota of infants delivered vaginally is dominated by microbial groups that colonize the vagina, whereas infants born by cesarean delivery have microbiota that more closely resembles those of the maternal skin community.16,20 However, the duration of the influence of these factors on the host remains obscure, with contradicting findings available to date. As a final age-related microbial shift, the elderly have a core microbiome different from that of younger adults, and the composition is directly correlated with health outcomes and the decline in the immune system.21 955 Clinical Therapeutics The gut is home to a diverse array of trillions of microbes that influence almost all aspects of human biology through their interactions with their host.1 Although the host provides the microbiota with the ecological niche and nutrients required of its survival, the indigenous microbiota, in turn, provides the host with full maturation of the innate and adaptive arms of the immune system, modulation of the nervous system function, nutrient absorption and fat distribution, contribution to digestion (eg, the ability of microbes to break down host nondigestible polysaccharides), the generation of short-chain fatty acids (SCFAs), metabolism of xenobiotics (which aids in protection from environmental toxins), and regulation of the gut motility.2,22–25 Colonization resistance is another protective function that the microbes provide to their host. This protective function results from a combination of various functions of commensals, which include their metabolic competition with the pathogen for the available nutrient resources. The microbiota provides colonization resistance; in addition, several of the host’s anatomical and physiological factors, including protective mucosal barrier, secretion of secretory immunoglobulin A (sIgA) and normal gastric motility, are also involved.26 This procedure promotes the establishment of an intestinal environment that prevents colonization with disease-inducing bacteria, which are abundant in unhealthy subjects.27 The Gut Microbiota Is Engaged in a Dynamic Interaction with the Host Immune System Initially, all microorganisms were considered as pathogens that are harmful and cause infectious diseases, and the host immune system should recognize and eliminate these invaders (non-self) while tolerating self-molecules to maintain homeostasis. However, the persistent association of host species with commensal organisms illustrates the major contribution of the gut microbiota in the education and regulation of the immune system.28 It is now well established that the immune system depends on colonization with a microbiota for its maturation.29 The initial microbial colonization is associated with prominent changes in mucosal and systemic immunity. For example, the physiologic phenomenon of maturation of the immune system, initiated within the fetal period, is dynamic in its character and expands in time through the first months (and even years) of a 956 child’s life; within the neonatal period, infancy, and early childhood, dysfunction of numerous components of the immune system is observed.30 Neonates have a decreased expression of costimulatory molecules, diminished dendritic cell (DC) differentiation, and impaired phagocytosis as well as a defective interaction between DCs, T lymphocytes, and regulatory T cells, impaired cytotoxic activity of T cells.31,32 Moreover, the suppressive activity of the transplacentally transmitted maternal IgG antibodies contributes to the deficiency of specific humoral response, including the low production of IgA in human neonates.33 Immune immaturities have also been reported in animal studies, in which germ-free (GF) mice and those in the first days of colonization share many traits, characterized by smaller Peyer’s patches and reduced cellularity of the lamina propria with lower Cd4þ and Cd8þ T cells.34,35 Plasma cells and intraepithelial lymphocytes are rare in the small intestinal mucosa, and sIgA levels are significantly lower with reduced expression of genes and activation markers of intestinal macrophages.35,36 sIgA, for instance, represents an intriguing example of how the microbiota mediates the host physiology through immune modulation. sIgA binds to luminal bacteria to prevent microbial translocation across the epithelial barrier,37 but it also influences the balance of immune and metabolic pathways in intestinal epithelial cells through a microbiota-dependent mechanism.38 In the absence of IgA, there is a shift toward the expression of genes involved in host defense, excessive production of antimicrobial proteins, and inflammatory-like responses to compensate for the deficiency in microbial compartmentalization via the sIgA. Instead, in the presence of IgA, the intestinal microbiota alters the expression of genes involved in lipid metabolism and storage, thereby shifting toward a homeostatic microbiota–immune–metabolic response.38,39 The initial engagement of the microbiota in the education of the host immune system appears to involve specific strains of the microbial community (pathobionts) of potential inflammatory allies, which survive and flourish in an environment enriched by the inflammatory process.25,40 To date, the few reports available suggest that these specific bacteria presumably activate mucosal immune priming for the bacterial sampling process to minimize their exposure to the systemic immune system by transient breaching of the epithelial barrier; they also stimulate the Volume 37 Number 5 S. El Aidy et al. production of antimicrobial proteins and a variety of immune pro-inflammatory and regulatory immune components.40–42 Elimination of the penetrant bacteria would thus lead to the induction of antigen presentation and processing molecules, innate and adaptive constituents that reside in the lamina propria. Hence, this initial close but regulated contact could benefit the host by strengthening its gut barrier, and it confers protection against true invading pathogens. Segmented filamentous bacteria represent an interesting example of such microbial–host initial immune engagement. In mice, these bacteria adhere transiently to the surface epithelium of the ileum and Peyer’s patches shortly after weaning43 to induce Th17 in the intestinal lamina propria via a mechanism that involves host production of serum amyloid A.44 These segmented filamentous bacteria induce Th17 locally in the GI tract; this effect was also shown to expand to the central nervous system (CNS),45 indicating that improving our understanding of the intestinal microbiota has therapeutic implications, not only for intestinal immunopathologies but also for systemic immune diseases. Combined transcriptional and immunohistochemical analyses of GF mice and ex-GF mice helped in detailing how the microbiota results in reprogramming of the intestinal mucosal immune responses, with the goal of initiating and maintaining a balanced immune response that leads to major molecular immune responses starting only few days after colonization.35,46 Cumulatively, the early life stage marks the most extensive changes in the host biology in response to the colonization of the microbial entities. At that point, the host encounters challenges, which, at the molecular level, elicit responses that involve genes associated with several illnesses (inflammatorylike response, metabolic disorder, and behavioral changes). Thus, abrupt shifts during the infant’s unique developmental path through this early unstable phase may have longer term health implications.47 Indeed, emerging evidence supports the concept of a critical window of development during early life that allows for the full-scale establishment of an adequate host–microbial homeostasis.19 After stabilization of the gut microbiota community, the immune system is continuously stimulated by several structural components of the microbial cells and responds by producing a wide range of lymphocytes and cytokines. In a state of homeostasis, the gut May 2015 microbiota stimulates a chronic state of low-level activation of the (innate) immune system in the host, in which bacterial particles stimulate intestinal macrophages and T cells to produce pro-inflammatory cytokines (eg, interleukin [IL]-1β, tumor necrosis factor-α, IL-18).48,49 The produced cytokines create a basal state of immune activation that starts at the intestinal mucosal surface and eventually affects the entire body. The adult human gut is believed to contain up to 1 g of lipopolysaccharides (LPS), and the low level exposure of immune cells to these bacterial cell wall components is essential in the establishment and maintenance of mucosal homeostasis.50 The specific mechanisms underlying host immune–microbe cross-talk remains unclear; nonetheless, a number of reports have been fundamental in revealing the mechanistic interaction that orchestrates this interaction between the host immune system and its microbiota. The regulation of the chemokine CXCL16, which is important for the invariant natural killer T-cell migration and homeostasis by the gut microbiota, represents an elegant example of this tightly regulated host–microbe dialogue. This chemokine regulation occurs via epigenetic changes (in particular, the reduction of methylation status of the CXCl16 gene), thereby reducing the number of invariant natural killer T cells in the lung and colonic lamina propria.51 Bacteroides fragilis is another well-studied example found to be critical for the normal function of the mammalian immune system via the production of a glycosphingolipid that inhibits natural killer T-cell proliferation in the colonic lamina propria.52 B fragilis polysaccharide A, conversely, interacts with the Toll-like receptor (TLR)-2 on DCs to induce colonic regulatory T cells (Tregs).53 Colonic Tregs have been demonstrated to also be induced by a group of Clostridium species, presumably via bacterial production of SCFAs.54 Strikingly, the difference between Treg stimulation with B fragilis and Clostridium species is that the latter can induce Tregs in both healthy and inflamed tissues; however, B fragilis induces Tregs only in inflamed tissues.28 These results indicate that commensals, once they escape their normal habitats and interact with the host in a different context, become pathogenic. Collectively, reports to date confirm the hygiene hypothesis, which has recently been reformulated as the “Old Friends” hypothesis; it postulates that 957 Clinical Therapeutics colonization with a “healthy” microbiota during the vulnerable developmental period exerts effects that may decrease susceptibility to diseases, whereas its absence or dysbiosis, as in antibiotic treatment in childhood, may have reverse effects.55 This concept is highlighted in the expanding repertoire of human diseases that are associated with changes in the intestinal microbiota, such as inflammatory bowel disease, allergic disorders, autoimmunity, and major depression.56 Gut Microbiota Talk to the Brain: Language Is an Enigma For 4150 years, it has been clear that there is a bidirectional gut–brain interaction, which regulates homeostasis. However, the concept of a microbiome– gut–brain axis is a recent phenomenon, and the mechanisms that underlie this axis are only slowly being resolved. Communication between the gut microbiota and the gut–brain axis can occur via multiple conduits that include gut-secreted neuropeptides, vagal nerve, sensory nerves, and immune mediators.57 The brain can influence the commensals directly via the receptor-mediated signaling and signaling molecules released in the gut lumen from immune cells or epithelial cells (in particular, enteroendocrine cells) or indirectly via changes in the intestinal motility and secretion.58 Similarly, the enteric bacteria have been shown to profoundly affect brain function and behavior.59 On the molecular level, GF mice exhibit an altered expression of genes involved in neuropeptide production,39 which are involved in brain development and behavior.60,61 On the behavioral level, GF mice exhibit increased spontaneous motor activity compared with that of conventional mice60 and become more exploratory (more like their donors) after fecal transfer from mice with anxiety-like behavior.62 Lactobacillus rhamnosus (JB-1), when fed to mice, reduces stress-induced corticosterone hormone levels. Moreover, the same strain makes the mice less anxious, an effect which seems to be dependent on the vagus nerve.63 Emerging data point to the involvement of bacterial components and by-products in the dialogue with the gut–brain axis. The mammalian behavior and neuroendocrine responses are dramatically affected by the metabolites circulating in the blood, many of which are dependent on the microbiota for their synthesis.64 Even cerebral metabolites are influenced by the commensals 958 through the microbiota–gut–brain axis, demonstrating how closely connected the gut microbiota is with brain health and disease, development, attenuation, learning, memory, and behavior.65 The gut microbiota is capable of altering the levels of metabolites that serve as precursors for the synthesis of the signaling molecules; for example, neuropeptides such as glutamine and tyrosine, as revealed from experiments performed in gnotobiotic animals.40,66,67 Although these metabolites can communicate the message around the entire body, our knowledge of which bacterial species produce which metabolites or display which components (and which of these synthesized products can bring about the host–microbial equilibrium) is limited (apart from the capsular polysaccharide A and glycophospholipid in B fragilis that exert potent immunoregulatory and neurologic effects42,68). For instance, we do not know which bacteria are involved in the synthesis of the bacterial metabolite hippurate, which is known to be modulated by diet, stress, and disease and is widely detected in several mammalian species.69 Bacterial LPS and peptidoglycan components are possible candidates in communicating the host–microbe interplay. For example, LPS can be carried directly to the brain via the blood circulation to act on the TLRs in the brain to give rise to neuroinflammatory processes, thereby modify the brain function.70 These outcomes show that the physiologic effect of the commensals is not limited to the GI tract but extends to the systemic immune system and the brain.57 SCFAs such as acetate, propionate, and n-butyrate, produced during fermentation of nondigestible polysaccharides, also represent probable candidates to be involved in setting up the balance.71 N-butyrate especially has been the focus of many studies aiming to unravel its overall impact on the human physiology. It has been shown, in in vitro experiments and in mice, to regulate energy homeostasis, stimulate leptin production in adipocytes, and induce the secretion of several neuropeptides, such as glucagon-like peptide-1. This action is proposed to modulate insulin secretion, lipid and glucose metabolism, and food intake, and it also exhibits antiinflammatory effects.72,73 On the level of the CNS, n-butyrate reportedly has profound effects on mood and behavior.74 Through their effect on gastric motility and intestinal transit stimulation, SCFAs result in an elevation in serotonin release, as reported in an in vitro colonic mucosal system.75 Therefore, SCFAs provide a Volume 37 Number 5 S. El Aidy et al. perfect example to illustrate the connectivity between the microbiota and the gut–brain axis.76–78 For example, Wang et al79 highlighted changes in the fecal concentrations of the SCFAs in children with autism spectrum disorders, suggesting that altered production of these microbial metabolites, which were shown to exhibit neuroactivity, may be a mechanism by which bacteria can alter brain function. Together, these findings suggest that the modification of the amount of dietary fiber across the microbe–host food network would influence the composition of the microbiota, which in turn modulates the luminal concentrations of intestinal SCFAs, a potential target for restoring the host overall physiology. In fact, an unbalanced diet that is poor in dietary fiber content (and thus does not require complex metabolic interactions between the microbial members) would result in lower microbial diversity and in the imbalanced host–microbial relationship that is observed in the emerging Western lifestyle–associated human diseases.80 In a recent attempt to unravel which bacterial metabolites can bring about the host–microbial balance, Williams et al81 used a combination of genetics and biochemistry to identify a key microbial enzyme that converts dietary tryptophan to the neuropeptide tryptamine (ie, tryptophan decarboxylase enzyme, which was found to be present in several bacteria that colonize 10% of the human population). This finding suggests that the gut microbiota can sequester tryptophan from the diet and alter its metabolites in the host, eventually resulting in altered brain levels of the neuropeptide serotonin, which ultimately affects brain function.82 Similarly, although the gut microbiota can alter the metabolism of several other neuropeptides, the limited knowledge available regarding the microbial mechanisms hinders efforts to close this gap in research. The assumption by Williams et al is supported by the elevated levels of tryptophan in GF animals compared with their conventional and conventionalized counterparts.66,67 Serotonin release from enterochromaffin epithelial cells is stimulated by tryptamine83 and is a key regulator of the gut motility and secretion.84 Recently, it has been shown that gut microbiota, acting through SCFAs, is an important determinant of enteric serotonin production and homeostasis. The gut microbiota from ex-GF mice colonized with human gut microbiota and conventionally raised mice the colonic rate limiting for mucosal serotonin synthesis; tryptophan hydroxylase 1 mRNAs May 2015 as well as the neuroendocrine secretion gene; chromogranin A with no effect on the serotonin transporter or serotonin catabolic gene; monoamine oxidase A.85 Interestingly, the reported effects did not result from an increase in the number of the cells producing serotonin but rather through the action of SCFAs. Changes in the levels of serotonin were also linked to the pathology of GI disorders, in which modulation of serotonin secretion is proposed as a treatment given its significant role in the enteric nervous system (ENS) and gut motility.86 Bacterial by-products that come in contact with the gut epithelium stimulate enteroendocrine cells (EECs) to produce several neuropeptides such as peptide YY, neuropeptide Y, cholecystokinin, glucagon-like peptide-1 and -2, and substance P.87 As an example, the gut senses the bacterial by-product SCFAs through G protein– coupled receptors 41 and 43 expressed on EECs.88 Upon their secretion by EECs, the neuropeptides presumably diffuse throughout the lamina propria, which is occupied with a variety of immune cells, until they reach the bloodstream or act on the vagal nerve or intrinsic sensory neurons.89,90 However, the exact mechanisms and whether the neuropeptides have a direct contribution in the bidirectional communication between the microbiota and CNS are still unknown. Recently, Bohorquez et al91 demonstrated a direct communication between EECs and neurons innervating the small intestine and colon that the paracrine transmission. The authors discovered a neuroepithelial circuit that can act as both sensory channels for food and as gut microbiota to communicate the message from the lumen to the ENS and CNS and vice versa. This physical innervation of sensory EECs suggests an accurate temporal transfer of the sensory signals originating in the gut lumen with a real-time modulatory feedback onto the EECs. Nonetheless, neuropeptides remain an intriguing way by which the gut microbiota initiates an effective dialogue with the host. Several reports have shown the changes in neuropeptides and neurotransmitters in response to changes in the gut microbiota composition. For example, Lactobacillus acidophilus stimulates the expression of cannabinoid and opioid receptors in rodents and in in vitro human epithelial cell cultures, thereby reducing experimentally evoked visceral pain.92 Antibiotic-induced microbial dysbiosis is associated with elevated levels of substance P in the colon.93 Not only does the host respond to the gut microbes by 959 Clinical Therapeutics producing neuropeptides, a variety of different components of intestinal microbiota can also produce a plethora of neurochemicals, which act to reduce inflammation, dampen the stress response, and potentially improve mood.94,95 For example, the metabolism of commensals has been shown to produce histamines and catecholamines, both of which can stimulate the ENS and the CNS via the primary afferent fibers of the vagal nerve.77,96 Dopamine is also produced in large quantities by a variety of bacteria including E coli, Bacillus cereus, Bacillus subtilis, and Staphylococcus aureus.97 However, the host produces enzymes that degrade these bacterial neurochemicals. For example, dopamine is silenced by the host-produced enzyme monoamine oxidase.57 It is tempting to speculate that the microbiota has evolved gene analogues to the host signaling molecules such as neuropeptides to manipulate its host to create a mutualistic relationship; for the host to prevent that manipulation in which it conflicts with the host’s fitness interests, however, corresponding host genes have evolved to eradicate the microbial attempts to invade. In fact, the same strategy is emphasized in the detoxification of enzymes evolved by mammals to eliminate gut-derived toxins and xenobiotics by rendering them more polar, enabling them to be easily secreted in the urine and thereby modifying both gut microbial products and drugs.23 For example, the compound 4-cresyl, which is formed from the purification of tyrosine (the precursor of the neuropeptide dopamine) by colonic microbiota, is converted into a more polar form (4-cresyl sulfate, an abundant urinary microbial metabolite) by host sulfotransferases.98 This microbial influence on the process of drug detoxification may also explain individual variations in certain drug responses, owing to the enormous interpersonal variations in the gut microbiota species.6,99 In fact, when rats were administered the antipsychotic olanzapine in the presence of a cocktail of broad-spectrum antibiotics, the Firmicutes phylum was suppressed and the relative abundance of Bacteroidetes was increased while at the same time ameliorating the olanzapine-induced weight gain, uterine fat deposition, plasma free fatty acid levels, and macrophage infiltration of adipose tissue.100 These findings of a microbiota-dependent, olanzapine-induced weight gain have recently been confirmed in GF mice, and in in vitro studies point to a direct interaction between olanzapine and microbes.101 960 Gut Microbiota Coordinates the Immune– Neuroendocrine Cross-Talk It has been an evolving subject since the beginning of modern medicine that the immune system and the brain, the 2 major adaptive systems of the body, talk to each other through a network of direct- and indirect-signaling pathways that involve the hypothalamic–pituitary–adrenal axis and the sympathetic nervous system. It is crucial for the nervous system to communicate with the immune system to maintain the host homeostasis because any challenge to the immune system would affect body temperature, visceral pain, and reflex withdrawal, all of which lead to pain sensation and behavioral changes but also negatively impact the immune system.102 However, assessing the host responses toward its commensal inhabitants, which are reflected in the complex interrelationships between nutrient metabolism, the immune system, and the neuro-endocrine system that occur at many levels (ranging from endocrine signaling to direct sensing of bacteria and their products), strongly suggests that the microbiota is an instrumental component of the psychoneuroimmunology network (Figure).103 Neuropeptides may be 1 of the signals by which the microbiota coordinates this communication network because neurons, epithelial cells, microbial cells, and immune cells can all produce and respond to them by expressing the appropriate receptors. For example, at the start of a local immune response (eg, during the initial microbial colonization), migration of immature DCs to lymph nodes was found to be mediated via α1-adrenergic receptors, emphasizing the early effects of the sympathetic nervous system.104 Moreover, various immune cells travel through the blood, and when they come within sensing distance of a given neuropeptide, they begin to chemotactically orient toward it; they then communicate with other immune cells in the adaptive arm (eg, B and T lymphocytes) to ensure a well-coordinated immune response. Neuropeptides are produced by the host second brain: the ENS (as well as the CNS), which functions to regulate blood flow and GI motility, partly through its interaction with the immune system and gut microbiota.105 Any changes in the luminal chemistry are sensed by the ENS, which transfers this information to the intrinsic neurons to cause the release of excitatory or inhibitory neuropeptides from a subset of cells named the intestinal cells of Cajal in the myenteric Volume 37 Number 5 S. El Aidy et al. Microbiota Microbiota signaling GCPRs Neuropeptides Microbial signaling Enteroendocrine signaling Dendritic cell Epithelium ChA T ChAT T cell Immune regulation Mood and behaviour Intestinal motility and secretion Barrier function Neuropeptide release Microbiota signaling; Excitatory/ inhibitory signals -microbial products, -microbial compounds, -neuropeptide analogues Mucosal immune signaling, Endocrine signaling Vagal nerve activation B cell Mucosa Submucosa Enteric neurons Smooth muscle Ma Immune–neuroendocrine signaling cro ph age Excitatory/ inhibitory signals Gut motility Muscularis Figure. The multidirectional dialogue between the gut microbiota, the (mucosal) immune system, and the neuroendocrine system. Within the gastrointestinal tract, the intestinal microbiota (via microbial signals) stimulates the immune system and the enteric nervous system, which in turn modulate the functionality of the central nervous system by various means of communication, including vagus nerve activation and cytokine release. In response, the brain modulates these multiple signaling pathways via the hypothalamic–pituitary–adrenal axis and sympathetic and vagal efferents. GCPRs ¼ G proteincoupled receptors. plexus.106 A recent study showed that in a state of homeostasis, the GI motility is regulated via the interactions between the ENS, microbiota, and a class of macrophages that reside in close proximity to the myenteric plexus and intestinal cells of Cajal, named muscularis macrophages (MMs). Muller et al107 unraveled the mechanisms regulating the cross-talk between the gut microbiota, the MM May 2015 immune cells, and eventually the ENS, in which the gut microbiota (particularly its component LPS) stimulates the production of the cytokine colonystimulating factor 1 in MMs, followed by the production of bone morphogenetic protein (BMP) 2; this in turn stimulates the ENS via activation of the receptors of BMP2 and downstream signaling molecules, resulting in modulation of the gut motility. 961 Clinical Therapeutics Subsequent to illustrating that MMs are sensitive to changes in the luminal environment after antibiotic treatment, the question remains whether the underlying mechanisms use a combination of downstream immune and neuronal signaling in the mucosa, including signals provided by EECs. Another plausible mechanism by which the gut microbiota with its components and accumulating metabolites can play a critical role in shuttling information with the host nervous and immune systems is through signaling pathways. These pathways include warning signals similar to those produced during injury or inflammation. In this case, the brain may receive the sentinel signals from the peripheral afferent nerves and elicit neural reflexes, which in turn regulate the immune responses to avoid any unwarranted inflammatory response. This scenario resembles what happens in endotoxemia, in which the signaling is conducted to the brain by the vagus nerve, and it responds by activating the efferent cholinergic signaling, which prevents the immune cells from producing inflammatory cytokines via the stimulation of their cholinergic receptors.108 Immune cells such as macrophages, neutrophils, DCs, and B and T lymphocytes not only produce neuropeptides but also express receptors to the neuropeptides. For example, acetylcholine, the principle vagal neuropeptide that attenuates the release of pro-inflammatory cytokines tumor necrosis factor-α, IL-1β, IL-6, and IL-18 (but not IL-10), is released from a subset of CD4þ T cells that transfer the signal to other immune cells through the activation of α7-nicotinic acetylcholine receptors on macrophages.109 Acetylcholine is also produced by a specific type of T cells known as ChATþ T cells, whose activation by norepinephrine, via β2-adrenergic receptors, results in the production of proinflammatory and anti-inflammatory cytokines.110 The location of ChATþ T cells is interesting: they are present in a low percentage in the spleen but are much more abundant in the Peyer’s patches,111 which are surrounded by enteric ganglia and are directly innervated by enteric and parasympathetic neurons whose fibers transverse in close proximity to immune cells. The high abundance of ChATþ T cells in Peyer’s patches suggests that their stimulation by microbial components exhibits both defensive and regulatory roles at the gut mucosal surface, in which trillions of microbiota reside. Acetylcholine receptor is also expressed on B cells (named ChATþB cells); similar 962 to ChATþT cells (but only in mucosal-associated lymphoid tissues and not in other peripheral lymphoid organs), they release acetylcholine when stimulated by another neuropeptide (ie, cholecystokinin).112 Intriguingly, it has been shown that the expression of ChATþB cells begins after birth, following the microbial colonization via MyD88-dependent TLR pathways in a transit manner. The assumption was evident by the reduction in Ach receptor expression after antibiotic treatment. The specific function of the ChATþB cells to control local recruitment of neutrophils during endotoxemia as shown in this study seems crucial to prevent severe tissue damage that is associated with the neutrophil recruitment or dysregulation of their proteolytic enzymes, as observed in cases of GI inflammation.113 Together, these evolving data support the key role of the gut microbes in orchestrating the existing immune–neuroendocrine dialogue. When Intimate Friends Turn into Foes The gut microbiota is normally thought of as including the “friendly” bacteria that perform many beneficial functions, until dysbiosis occurs. In this situation, the imbalanced gut microbiota represents an environmental alteration and may become harmful to the host. Microbial dysbiosis is characterized by reduced diversity and the outgrowth of pathobionts, which can lead to inadequate immune functioning and inflammatory responses by an imbalance between regulatory and pro-inflammatory lymphocytes, GI motility and permeability, and subsequent changes in metabolism and extra-intestinal milieus, including the behavior and CNS functions. Therefore, the host may encounter major challenges, which are linked at the molecular level to diverse, complex diseases (eg, inflammatory bowel disease and metabolic, neurologic, and cardiovascular disorders).114 Altered exposure to the microbiota and its components has been associated with separately modified immune, endocrine, and CNS functions, and it may therefore disrupt the normal immune–neuroendocrine network. For example, elevated levels of inflammatory response during microbial dysbiosis appear to have direct effects on the tryptophan metabolic pathway. Dysregulation of the kynurenine arm in the tryptophan metabolic pathway, which is observed in many disorders of the brain and the GI tract,66 involves the catalytic enzyme indoleamine-2,3-dioxygenase,40 the activity of which is induced by inflammatory mediators and by Volume 37 Number 5 S. El Aidy et al. corticosteroids.115 These findings suggest that alterations in the production of neuropeptides by dysbiosis would in turn alter the neuroendocrine–immune signaling or act directly on the CNS. The gut microbiota also impacts brain health in several ways.116 Exposure of GF adult mice to the gut microbiota decreased permeability of the blood–brain barrier.117 Bacteria and their components such as LPS cause leakage of the gut barrier and the subsequent stimulation of the innate immune response and inflammation. Excessive production of cytokines is associated with systemic inflammation and high plasma levels of proinflammatory and anti-inflammatory cytokines, along with salivary and plasma cortisol and norepinephrine levels. Elevated levels of cytokines and neuropeptides were shown to be associated with disrupted sleep, depressed mood, increased anxiety levels, impaired long-term memory for emotional stimuli, and elevated visceral pain sensitivity.116,118 Particularly in elderly patients, elevated levels of the gut microbiota– delivered LPS, and the resulting endotoxemia, are more frequent as a result of small intestinal bacterial overgrowth that is caused by disturbed gut motility and increased intestinal permeability.119 Indeed, gut microbiota alterations have been described in several diseases with altered GI motility, raising the urgency that increased attention be given to the role of the gut microbiome in modulating GI function. Patients with schizophrenia are also known to have high gut bacterial translocation, and the presence of CD14 as a translocator marker (which was found to be associated with the bacterial component C-reactive protein) tripled the risk of schizophrenia.120 The host responds to specific microbes with antibodies or antigen-specific cellular immune responses. Thereby, when microbial dysbiosis occurs, CNS dysfunction could result from the autoimmune reaction caused by molecular mimicry between bacteria and hostself proteins, as in case of multiple sclerosis.121 Multiple sclerosis, a devastating autoimmune disease that leads to progressive deterioration of neurologic function, has been shown to be profoundly influenced by the gut microbiota via its ability to stimulate proinflammatory T-cell responses during experimental-associated encephalomyelitis.45 Moreover, the commensal microbes were shown to be essential in triggering immune processes that led to a relapsing-remitting autoimmune disease driven by myelin-specific CD4þT cells.122 The same study demonstrated that the recruitment and activation of autoantibody-producing B cells from the May 2015 endogenous immune repertoire depend on the availability of the target auto-antigen, myelin oligodendrocyte glycoprotein, and commensal microbiota. This finding again highlights the crucial role of the gut microbiota in coordinating the neuro–endocrine–immune cross-talk. CONCLUSIONS Over the past decade, preclinical trials, as well as a few clinical studies, have highlighted the crucial role of the gut microbiota in maintaining host homeostasis. Analogously, microbiota dysbiosis has been associated with a wide range of host disorders, including inflammatory bowel disorders, neuroendocrine disorders, and behavioral disorders. This growing body of research illustrates the significant direct effect that microbes have on the host immune and neuroendocrine systems. Nonetheless, the underlying molecular mechanisms that orchestrate this dialogue are far less certain than what is required to open new avenues of therapeutic intervention. A better understanding of the causative mechanisms that govern the microbiota– host interactions is urgently needed to provide novel avenues to rationally intervene in disease situations, using either microbial or dietary interventions, with the goal of correcting situations of microbial dysbiosis and thereby restoring homeostasis. Despite the availability of an enormous array of data from preclinical studies, limited information is available on how these findings may translate to the human situation in health and disease. The huge interindividual variation among human subjects, not only in terms of microbiota composition and its genome but also diet, sex-related differences, genetics, and environmental factors, adds to the complexity of deciphering the host–microbe interplay in clinical studies. Another major challenge in clinical studies is that the food industry does not have as strong of a clinical trials culture as there has been traditionally in the pharma sector, and the cost of large-scale trials is often prohibitive. ACKNOWLEDGMENTS Drs. Dinan and Cryan are supported by Science Foundation Ireland (grant no. 07/CE/B1368 and 12/ RC/2273) and by the Irish Health Research Board, Health Research Awards (HRA_POR/2011/23) and (HRA_POR/2012/32). Sahar El Aidy contribution: literature search, figure creation, writing. Timothy G. Dinan and John Cryan: critical reviewing. 963 Clinical Therapeutics CONFLICTS OF INTEREST The authors’ research at the Alimentary Pharmabiotic Centre is funded by Science Foundation Ireland, through the Irish Government’s National Development Plan in collaboration with a variety of pharmaceutical and food industries. The authors have indicated that they have no other conflicts of interest regarding the content of this article. REFERENCES 1. Blaser MJ. The microbiome revolution. J Clin Invest. 2014;124: 4162–4165. 2. Cryan JF, Dinan TG. Mind-altering microorganisms: the impact of the gut microbiota on brain and behaviour. Nat Rev Neurosci. 2012;13:701–712. 3. Tremaroli V, Backhed F. Functional interactions between the gut microbiota and host metabolism. Nature. 2012;489:242–249. 4. Hooper LV, Littman DR, Macpherson AJ. Interactions between the microbiota and the immune system. Science. 2012;336:1268–1273. 5. Sharon G, et al. Specialized metabolites from the microbiome in health and disease. Cell Metab. 2014;20:719–730. 6. Patterson AD, Turnbaugh PJ. Microbial determinants of biochemical individuality and their impact on toxicology and pharmacology. Cell Metab. 2014;20:761–768. 7. Rogers GB. The human microbiome: opportunities and challenges for clinical care. Intern Med J. 2014. [E-pub ahead of print]. 8. Walter J, Ley R. The human gut microbiome: ecology and recent evolutionary changes. Annu Rev Microbiol. 2011;65:411–429. 9. Eckburg PB, et al. Diversity of the human intestinal microbial flora. Science. 2005;308:1635–1638. 10. Qin J, et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature. 2010;464:59–65. 11. Kau AL, et al. Human nutrition, the gut microbiome and the immune system. Nature. 2011;474:327–336. 12. Tilg H, Moschen AR. Food, immunity, and the microbiome. Gastroenterology. 2015. [E-pub ahead of print]. 13. Jimenez E, et al. Staphylococcus epidermidis: a differential trait of the fecal microbiota of breast-fed infants. BMC Microbiol. 2008;8:143. 14. Palmer C, et al. Development of the human infant intestinal microbiota. PLoS Biol. 2007;5:e177. 15. Nylund L, et al. Intestinal microbiota during early life— impact on health and disease. Proc Nutr Soc. 2014;73: 457–469. 964 16. Mueller NT, et al. The infant microbiome development: mom matters. Trends Mol Med. 2015;21:109–117. 17. Cox LM, et al. Altering the intestinal microbiota during a critical developmental window has lasting metabolic consequences. Cell. 2014;158:705–721. 18. Penders J, et al. Factors influencing the composition of the intestinal microbiota in early infancy. Pediatrics. 2006;118:511–521. 19. Borre YE, et al. Microbiota and neurodevelopmental windows: implications for brain disorders. Trends Mol Med. 2014;20:509–518. 20. Johnson CL, Versalovic J. The human microbiome and its potential importance to pediatrics. Pediatrics. 2012;129:950–960. 21. Claesson MJ, et al. Gut microbiota composition correlates with diet and health in the elderly. Nature. 2012;488:178–184. 22. El Aidy S, et al. Intestinal colonization: how key microbial players become established in this dynamic process: microbial metabolic activities and the interplay between the host and microbes. Bioessays. 2013;35:913–923. 23. Holmes E, et al. Therapeutic modulation of microbiotahost metabolic interactions. Sci Transl Med. 2012;4. 137rv6. 24. Blumberg R, Powrie F. Microbiota, disease, and back to health: a metastable journey. Sci Transl Med. 2012;4. 137rv7. 25. Chung H, et al. Gut immune maturation depends on colonization with a host-specific microbiota. Cell. 2012;149:1578–1593. 26. Hooper LV, Macpherson AJ. Immune adaptations that maintain homeostasis with the intestinal microbiota. Nat Rev Immunol. 2010;10:159–169. 27. Veiga P, et al. Bifidobacterium animalis subsp. lactis fermented milk product reduces inflammation by altering a niche for colitogenic microbes. Proc Natl Acad Sci U S A. 2010;107:18132–18137. 28. Surana NK, Kasper DL. Deciphering the tete-a-tete between the microbiota and the immune system. J Clin Invest. 2014;124:4197–4203. 29. Kostic AD, Howitt MR, Garrett WS. Exploring hostmicrobiota interactions in animal models and humans. Genes Dev. 2013;27:701–718. 30. Newburg DS, Walker WA. Protection of the neonate by the innate immune system of developing gut and of human milk. Pediatr Res. 2007;61:2–8. 31. Velilla PA, Rugeles MT, Chougnet CA. Defective antigenpresenting cell function in human neonates. Clin Immunol. 2006;121:251–259. 32. Kotiranta-Ainamo A, Rautonen J, Rautonen N. Imbalanced cytokine secretion in newborns. Biol Neonate. 2004;85:55–60. 33. Keles S, et al. Transient hypogammaglobulinemia and unclassified hypogammaglobulinemia: ‘similarities and differences’. Pediatr Allergy Immunol. 2010;21:843–851. Volume 37 Number 5 S. El Aidy et al. 34. Wei B, et al. Commensal microbiota and CD8þ T cells shape the formation of invariant NKT cells. J Immunol. 2010;184:1218– 1226. 35. El Aidy S, et al. Temporal and spatial interplay of microbiota and intestinal mucosa drive establishment of immune homeostasis in conventionalized mice. Mucosal Immunol. 2012;5:567–579. 36. Mikkelsen HB, et al. Macrophages in the small intestinal muscularis externa of embryos, newborn and adult germ-free mice. J Mol Histol. 2004;35:377–387. 37. Macpherson AJ, Geuking MB, McCoy KD. Immune responses that adapt the intestinal mucosa to commensal intestinal bacteria. Immunology. 2005;115:153–162. 38. Shulzhenko N, et al. Crosstalk between B lymphocytes, microbiota and the intestinal epithelium governs immunity versus metabolism in the gut. Nat Med. 2011;17:1585–1593. 39. El Aidy S, et al. The gut microbiota elicits a profound metabolic reorientation in the mouse jejunal mucosa during conventionalisation. Gut. 2013;62: 1306–1314. 40. El Aidy S, et al. Transient inflammatory-like state and microbial dysbiosis are pivotal in establishment of mucosal homeostasis during colonisation of germ-free mice. Benef Microbes. 2014;5:67–77. 41. Galindo-Villegas J, et al. Regulation of immunity and disease resistance by commensal microbes and chromatin modifications during zebrafish development. Proc Natl Acad Sci U S A. 2012;109:E2605–E2614. 42. Mazmanian SK, et al. An immunomodulatory molecule of symbiotic bacteria directs maturation of the host immune system. Cell. 2005;122:107–118. 43. Gaboriau-Routhiau V, et al. The key role of segmented filamentous May 2015 44. 45. 46. 47. 48. 49. 50. 51. 52. 53. bacteria in the coordinated maturation of gut helper T cell responses. Immunity. 2009;31: 677–689. Gaboriau-Routhiau V, Lecuyer E, Cerf-Bensussan N. Role of microbiota in postnatal maturation of intestinal T-cell responses. Curr Opin Gastroenterol. 2011;27: 502–508. Lee YK, et al. Proinflammatory T-cell responses to gut microbiota promote experimental autoimmune encephalomyelitis. Proc Natl Acad Sci U S A. 2011;108:4615–4622. El Aidy S, Kleerebezem M. Molecular signatures for the dynamic process of establishing intestinal host-microbial homeostasis: potential for disease diagnostics? Curr Opin Gastroenterol. 2013;29: 621–627. Costello EK, et al. The application of ecological theory toward an understanding of the human microbiome. Science. 2012;336: 1255–1262. Heumann D, et al. Gram-positive cell walls stimulate synthesis of tumor necrosis factor alpha and interleukin-6 by human monocytes. Infect Immun. 1994;62:2715–2721. Ulevitch RJ, Tobias PS. Receptordependent mechanisms of cell stimulation by bacterial endotoxin. Annu Rev Immunol. 1995;13:437–457. Bested AC, Logan AC, Selhub EM. Intestinal microbiota, probiotics and mental health: from Metchnikoff to modern advances: Part II - contemporary contextual research. Gut Pathog. 2013;5:3. Olszak T, et al. Microbial exposure during early life has persistent effects on natural killer T cell function. Science. 2012;336:489–493. Wieland Brown LC, et al. Production of alpha-galactosylceramide by a prominent member of the human gut microbiota. PLoS Biol. 2013;11: e1001610. Round JL, Mazmanian SK. The gut microbiota shapes intestinal 54. 55. 56. 57. 58. 59. 60. 61. 62. 63. immune responses during health and disease. Nat Rev Immunol. 2009;9:313–323. Atarashi K, et al. Induction of colonic regulatory T cells by indigenous Clostridium species. Science. 2011;331:337–341. Rook GA. Regulation of the immune system by biodiversity from the natural environment: an ecosystem service essential to health. Proc Natl Acad Sci U S A. 2013; 110:18360–18367. Rook GA, Raison CL, Lowry CA. Can we vaccinate against depression? Drug Discov Today. 2012;17: 451–458. Holzer P, Farzi A. Neuropeptides and the microbiota-gut-brain axis. Adv Exp Med Biol. 2014;817: 195–219. Rhee SH, Pothoulakis C, Mayer EA. Principles and clinical implications of the brain-gut-enteric microbiota axis. Nat Rev Gastroenterol Hepatol. 2009;6:306–314. Lyte M, Cryan JF. Dealing with ability of the microbiota to influence the brain, and ultimately cognition and behavioral. Adv Exp Med Biol. 2014;817:ix–xi. Diaz Heijtz R, et al. Normal gut microbiota modulates brain development and behavior. Proc Natl Acad Sci U S A. 2011;108: 3047–3052. Stilling RM, Dinan TG, Cryan JF. Microbial genes, brain & behaviour—epigenetic regulation of the gut-brain axis. Genes Brain Behav. 2014;13:69–86. Bercik P, et al. The intestinal microbiota affect central levels of brain-derived neurotropic factor and behavior in mice. Gastroenterology. 2011;141:599–609. 609.e1-3. Bravo JA, et al. Ingestion of Lactobacillus strain regulates emotional behavior and central GABA receptor expression in a mouse via the vagus nerve. Proc Natl Acad Sci U S A. 2011;108: 16050–16055. 965 Clinical Therapeutics 64. Wikoff WR, et al. Metabolomics analysis reveals large effects of gut microflora on mammalian blood metabolites. Proc Natl Acad Sci U S A. 2009;106:3698–3703. 65. Matsumoto M, et al. Cerebral low-molecular metabolites influenced by intestinal microbiota: a pilot study. Front Syst Neurosci. 2013;7:9. 66. Clarke G, et al. The microbiomegut-brain axis during early life regulates the hippocampal serotonergic system in a sex-dependent manner. Mol Psychiatry. 2013;18: 666–673. 67. El Aidy S, et al. The microbiota and the gut-brain axis: insights from the temporal and spatial mucosal alterations during colonisation of the germfree mouse intestine. Benef Microbes. 2012;3:251–259. 68. Hsiao EY, et al. Microbiota modulate behavioral and physiological abnormalities associated with neurodevelopmental disorders. Cell. 2013;155:1451–1463. 69. Holmes E, et al. Human metabolic phenotype diversity and its association with diet and blood pressure. Nature. 2008;453:396–400. 70. Arroyo DS, et al. Toll-like receptors are key players in neurodegeneration. Int Immunopharmacol. 2011;11:1415–1421. 71. Flint HJ, et al. Microbial degradation of complex carbohydrates in the gut. Gut Microbes. 2012;3: 289–306. 72. Burcelin R, Cani PD, Knauf C. Glucagon-like peptide-1 and energy homeostasis. J Nutr. 2007; 137(11 Suppl):2534s–2538s. 73. Musso G, Gambino R, Cassader M. Interactions between gut microbiota and host metabolism predisposing to obesity and diabetes. Annu Rev Med. 2011;62:361–380. 74. Schroeder FA, et al. Antidepressant-like effects of the histone deacetylase inhibitor, sodium butyrate, in the mouse. Biol Psychiatry. 2007;62:55–64. 966 75. Grider JR, Piland BE. The peristaltic reflex induced by shortchain fatty acids is mediated by sequential release of 5-HT and neuronal CGRP but not BDNF. Am J Physiol Gastrointest Liver Physiol. 2007;292:G429–G437. 76. Macfabe DF. Short-chain fatty acid fermentation products of the gut microbiome: implications in autism spectrum disorders. Microb Ecol Health Dis. 2012:23. 77. Gundersen BB, Blendy JA. Effects of the histone deacetylase inhibitor sodium butyrate in models of depression and anxiety. Neuropharmacology. 2009;57: 67–74. 78. Thomas SP, Nandhra HS, Singh SP. Pharmacologic treatment of first-episode schizophrenia: a review of the literature. Prim Care Companion CNS Disord. 2012:14. 79. Wang L, et al. Elevated fecal short chain fatty acid and ammonia concentrations in children with autism spectrum disorder. Dig Dis Sci. 2012;57:2096–2102. 80. De Filippo C, et al. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc Natl Acad Sci U S A. 2010;107:14691–14696. 81. Williams BB, et al. Discovery and characterization of gut microbiota decarboxylases that can produce the neurotransmitter tryptamine. Cell Host Microbe. 2014; 16:495–503. 82. O’Mahony SM, et al. Serotonin, tryptophan metabolism and the brain-gut-microbiome axis. Behav Brain Res. 2015;277c:32–48. 83. Takaki M, et al. Physiological responses of guinea-pig myenteric neurons secondary to the release of endogenous serotonin by tryptamine. Neuroscience. 1985; 16:223–240. 84. Turvill JL, Connor P, Farthing MJ. The inhibition of cholera toxin-induced 5-HT release by 85. 86. 87. 88. 89. 90. 91. 92. 93. 94. the 5-HT(3) receptor antagonist, granisetron, in the rat. Br J Pharmacol. 2000;130:1031–1036. Reigstad CS, et al. Gut microbes promote colonic serotonin production through an effect of short-chain fatty acids on enterochromaffin cells. Faseb J. 2014. [E-pub ahead of print]. Gershon MD, Liu MT. Serotonin and neuroprotection in functional bowel disorders. Neurogastroenterol Motil. 2007;19 (Suppl 2):19–24. Furness JB, et al. The gut as a sensory organ. Nat Rev Gastroenterol Hepatol. 2013;10:729–740. Samuel BS, et al. Effects of the gut microbiota on host adiposity are modulated by the shortchain fatty-acid binding G protein-coupled receptor, Gpr41. Proc Natl Acad Sci U S A. 2008; 105:16767–16772. Cummings DE, Overduin J. Gastrointestinal regulation of food intake. J Clin Invest. 2007;117:13–23. Okano-Matsumoto S, et al. Electrophysiological evidence for distinct vagal pathways mediating CCK-evoked motor effects in the proximal versus distal stomach. J Physiol. 2011;589(Pt 2): 371–393. Bohorquez DV, et al. Neuroepithelial circuit formed by innervation of sensory enteroendocrine cells. J Clin Invest. 2015;125:782–786. Rousseaux C, et al. Lactobacillus acidophilus modulates intestinal pain and induces opioid and cannabinoid receptors. Nat Med. 2007;13:35–37. Verdu EF, et al. Specific probiotic therapy attenuates antibiotic induced visceral hypersensitivity in mice. Gut. 2006;55:182–190. Lyte M. Probiotics function mechanistically as delivery vehicles for neuroactive compounds: Microbial endocrinology in the design and use of probiotics. Bioessays. 2011;33:574–581. Volume 37 Number 5 S. El Aidy et al. 95. Dinan TG, Stanton C, Cryan JF. Psychobiotics: a novel class of psychotropic. Biol Psychiatry. 2013; 74:720–726. 96. Asano Y, et al. Critical role of gut microbiota in the production of biologically active, free catecholamines in the gut lumen of mice. Am J Physiol Gastrointest Liver Physiol. 2012;303:G1288–G1295. 97. Wall R, et al. Bacterial neuroactive compounds produced by psychobiotics. Adv Exp Med Biol. 2014;817:221–239. 98. Wilson ID, Nicholson JK. Topics in Xenobiochemistry: do metabolic pathways exist for xenobiotics? The micro-metabolism hypothesis. Xenobiotica. 2003;33:887–901. 99. Nicholson JK, Holmes E, Wilson ID. Gut microorganisms, mammalian metabolism and personalized health care. Nat Rev Microbiol. 2005;3:431–438. 100. Davey KJ, et al. Antipsychotics and the gut microbiome: olanzapine-induced metabolic dysfunction is attenuated by antibiotic administration in the rat. Transl Psychiatry. 2013;3:e309. 101. Morgan AP, et al. The antipsychotic olanzapine interacts with the gut microbiome to cause weight gain in mouse. PLoS One. 2014;9:e115225. 102. Olofsson PS, et al. Rethinking inflammation: neural circuits in the regulation of immunity. Immunol Rev. 2012;248:188–204. 103. El Aidy S, Dinan TG, Cryan JF. Immune modulation of the brain-gut-microbe axis. Front Microbiol. 2014;5:146. 104. Maestroni GJ. Dendritic cell migration controlled by alpha 1badrenergic receptors. J Immunol. 2000;165:6743–6747. 105. Furness JB. The enteric nervous system and neurogastroenterology. Nat Rev Gastroenterol Hepatol. 2012; 9:286–294. 106. Kunze WA, Furness JB. The enteric nervous system and May 2015 107. 108. 109. 110. 111. 112. 113. 114. regulation of intestinal motility. Annu Rev Physiol. 1999;61:117–142. Muller PA, et al. Crosstalk between muscularis macrophages and enteric neurons regulates gastrointestinal motility. Cell. 2014;158:300–313. Borovikova LV, et al. Vagus nerve stimulation attenuates the systemic inflammatory response to endotoxin. Nature. 2000;405:458–462. Wang H, et al. Nicotinic acetylcholine receptor alpha7 subunit is an essential regulator of inflammation. Nature. 2003;421:384–388. Rosas-Ballina M, Tracey KJ. Cholinergic control of inflammation. J Intern Med. 2009;265:663–679. Rosas-Ballina M, et al. Acetylcholine-synthesizing T cells relay neural signals in a vagus nerve circuit. Science. 2011;334:98–101. Reardon C, et al. Lymphocytederived ACh regulates local innate but not adaptive immunity. Proc Natl Acad Sci U S A. 2013;110:1410–1415. Reardon C, et al. Thymic stromal lymphopoetin-induced expression of the endogenous inhibitory enzyme SLPI mediates recovery from colonic inflammation. Immunity. 2011;35:223–235. Cho I, Blaser MJ. The human microbiome: at the interface of health and disease. Nat Rev Genet. 2012;13:260–270. 115. Ruddick JP, et al. Tryptophan metabolism in the central nervous system: medical implications. Expert Rev Mol Med. 2006;8:1–27. 116. Galland L. The gut microbiome and the brain. J Med Food. 2014; 17:1261–1272. 117. Braniste V, et al. The gut microbiota influences blood-brain barrier permeability in mice. Sci Transl Med. 2014;6. 263ra158. 118. Grigoleit JS, et al. Dose-dependent effects of endotoxin on neurobehavioral functions in humans. PLoS One. 2011;6:e28330. 119. Bauer TM, et al. Small intestinal bacterial overgrowth in human cirrhosis is associated with systemic endotoxemia. Am J Gastroenterol. 2002;97:2364–2370. 120. Severance EG, et al. Discordant patterns of bacterial translocation markers and implications for innate immune imbalances in schizophrenia. Schizophr Res. 2013;148:130–137. 121. Sathyabama S, Khan N, Agrewala JN. Friendly pathogens: prevent or provoke autoimmunity. Crit Rev Microbiol. 2014;40:273–280. 122. Berer K, et al. Commensal microbiota and myelin autoantigen cooperate to trigger autoimmune demyelination. Nature. 2011;479: 538–541. Address correspondence to: John F. Cryan, PhD, Department of Anatomy and Neuroscience, University College Cork, Cork, Ireland. E-mail: j.cryan@ ucc.ie 967