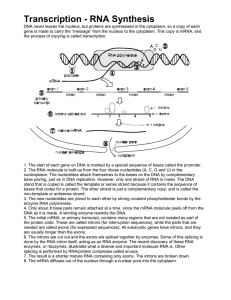
Transcription
... nucleoplasm. The nucleotides attach themselves to the bases on the DNA by complementary base pairing, just as in DNA replication. However, only one strand of RNA is made. The DNA stand that is copied is called the template or sense strand because it contains the sequence of bases that codes for a pr ...
... nucleoplasm. The nucleotides attach themselves to the bases on the DNA by complementary base pairing, just as in DNA replication. However, only one strand of RNA is made. The DNA stand that is copied is called the template or sense strand because it contains the sequence of bases that codes for a pr ...
Ex vivo analysis of splicing assays
... alternative splicing patterns (Ward et al., 2009; López-Bigas et al., 2005). The question is how to improve our understanding of new disease-causing mutations, that do not affect the coding potential of a gene, but that affect pre-mRNA splicing. In order to analyse precise mechanism of alternative s ...
... alternative splicing patterns (Ward et al., 2009; López-Bigas et al., 2005). The question is how to improve our understanding of new disease-causing mutations, that do not affect the coding potential of a gene, but that affect pre-mRNA splicing. In order to analyse precise mechanism of alternative s ...
Lecture 10 Biol302 Spring 2011
... Removal of introns must be very precise. Conserved sequences for removal of the introns of nuclear mRNA genes are minimal. – Dinucleotide sequences at the 5’ and 3’ ends of introns. – An A residue about 30 nucleotides upstream from the 3’ splice site is needed for lariat formation. ...
... Removal of introns must be very precise. Conserved sequences for removal of the introns of nuclear mRNA genes are minimal. – Dinucleotide sequences at the 5’ and 3’ ends of introns. – An A residue about 30 nucleotides upstream from the 3’ splice site is needed for lariat formation. ...
Trnascription in eucaryotes
... regions called introns that interrupt the coding regions. A gene can contain as many as 500 introns that vary from 50-20,000 base pairs in length. The primary transcript must be edited to remove the introns before translation can occur. ...
... regions called introns that interrupt the coding regions. A gene can contain as many as 500 introns that vary from 50-20,000 base pairs in length. The primary transcript must be edited to remove the introns before translation can occur. ...
talk_splicing - Columbia University
... Binding can be directly to the silenced splicing factor (U2F65, SF1, …). Splicing factors have been shown to bind mature mRNA in human cells (Carmo-Fonseca et. al, 2006). Alternatively, binding can be to some other factor which is affected by the silencing (secondary effect). Binding can induce both ...
... Binding can be directly to the silenced splicing factor (U2F65, SF1, …). Splicing factors have been shown to bind mature mRNA in human cells (Carmo-Fonseca et. al, 2006). Alternatively, binding can be to some other factor which is affected by the silencing (secondary effect). Binding can induce both ...
Pre-mRNA splicing: life at the centre of the central dogma
... Alternative splicing enables a single gene to increase its coding capacity, allowing the synthesis of structurally and functionally distinct protein isoforms. Usually, alternative exons have suboptimal splicing signals, and their inclusion is modulated by trans-acting factors that recognize an arran ...
... Alternative splicing enables a single gene to increase its coding capacity, allowing the synthesis of structurally and functionally distinct protein isoforms. Usually, alternative exons have suboptimal splicing signals, and their inclusion is modulated by trans-acting factors that recognize an arran ...
Answers questions chapter 14
... huge challenge for those that include large numbers of introns (for example, well over a hundred in some cases), and/or introns of vast dimensions (for example, up to 500,000 nucleotides). This challenge is compounded by the relatively weak sequence requirements for splice sites, as this increases t ...
... huge challenge for those that include large numbers of introns (for example, well over a hundred in some cases), and/or introns of vast dimensions (for example, up to 500,000 nucleotides). This challenge is compounded by the relatively weak sequence requirements for splice sites, as this increases t ...
NisimNaim-AdiPotok
... factories not involved in pre-mRNA splicing מר יהודה ברודי טל-ד"ר ירון שב ...
... factories not involved in pre-mRNA splicing מר יהודה ברודי טל-ד"ר ירון שב ...
Self-Quiz Questions Activity 1: When is a Genome
... Match the correct term with each definition or select the best answer for each question. 1. A series of codons from a single strand of DNA sequence which can be "read" in three different ways, depending on whether one starts at the first nucleotide position, the second or third Reading Frame (RF) Al ...
... Match the correct term with each definition or select the best answer for each question. 1. A series of codons from a single strand of DNA sequence which can be "read" in three different ways, depending on whether one starts at the first nucleotide position, the second or third Reading Frame (RF) Al ...
When Is a Genome Project Finished?
... Match the correct term with each definition or select the best answer for each question. 1. A series of codons from a single strand of DNA sequence which can be "read" in three different ways, depending on whether one starts at the first nucleotide position, the second or third Reading Frame (RF) Al ...
... Match the correct term with each definition or select the best answer for each question. 1. A series of codons from a single strand of DNA sequence which can be "read" in three different ways, depending on whether one starts at the first nucleotide position, the second or third Reading Frame (RF) Al ...
U1-snRNA–mediated rescue of mRNA processing in
... Modified small nuclear RNAs (snRNAs) have been shown to promote changes in mRNA splicing in cellular and animal models of human diseases. However, these approaches were mainly aimed at inducing skipping of defective exons,10,11 an approach that would not produce functional coagulation proteins, or m ...
... Modified small nuclear RNAs (snRNAs) have been shown to promote changes in mRNA splicing in cellular and animal models of human diseases. However, these approaches were mainly aimed at inducing skipping of defective exons,10,11 an approach that would not produce functional coagulation proteins, or m ...
outline File - selu moodle
... Begins at a promoter transcribes the transcription unit ends at the terminator Promoter – sequence within DNA Elongation uses RNA polymerase to add ribonucleotides that are complementary to the template strand Most common mechanism for termination is the formation of a hairpin structure In proka ...
... Begins at a promoter transcribes the transcription unit ends at the terminator Promoter – sequence within DNA Elongation uses RNA polymerase to add ribonucleotides that are complementary to the template strand Most common mechanism for termination is the formation of a hairpin structure In proka ...
INTERVENING SEQUENCES IN EUKARYOTES
... enzyme(s), but rather is simply a series of phosphoester bond transfers. ...
... enzyme(s), but rather is simply a series of phosphoester bond transfers. ...
M1 - Biochemistry Transcription III / mRNA Processing
... Why are there introns???? Don’t know, but an evolutionary advantage may be that splicing errors potentially can lead to functionally different but useful proteins that come about without having to make a new gene. Alternative (differential) splicing of a precursor-RNA from the same genes can, for so ...
... Why are there introns???? Don’t know, but an evolutionary advantage may be that splicing errors potentially can lead to functionally different but useful proteins that come about without having to make a new gene. Alternative (differential) splicing of a precursor-RNA from the same genes can, for so ...
Alternative Splicing
... • mutations in the Alu can create a 5’ or 3’ site in an intron causing it to be an exon • This mutation doesn’t impact existing exons • It only has effect when it is alternatively spliced in ...
... • mutations in the Alu can create a 5’ or 3’ site in an intron causing it to be an exon • This mutation doesn’t impact existing exons • It only has effect when it is alternatively spliced in ...
When Is a Genome Project Finished?
... Match the correct term with each definition or select the best answer for each question. 1. A series of codons from a single strand of DNA sequence which can be "read" in three different ways, depending on whether one starts at the first nucleotide position, the second or third Reading Frame (RF) Al ...
... Match the correct term with each definition or select the best answer for each question. 1. A series of codons from a single strand of DNA sequence which can be "read" in three different ways, depending on whether one starts at the first nucleotide position, the second or third Reading Frame (RF) Al ...
ASviewer: Visualizing the transcript structure and functional
... Summary: Alternative splicing (AS) produces diverse transcript structures by differential use of splice sites. Comparing the gene structure and functional domains of splice variants is an essential but nontrivial task with numerous gene predictions available publicly. We developed a novel viewer (AS ...
... Summary: Alternative splicing (AS) produces diverse transcript structures by differential use of splice sites. Comparing the gene structure and functional domains of splice variants is an essential but nontrivial task with numerous gene predictions available publicly. We developed a novel viewer (AS ...
LECT34 RNAproc
... Q: How is this accomplished? A: Cells have mechanism that recognize introns. The most common is a spliceosome that recognizes the boundaries of intron-exon junctions and knows were to cleave and splice Q: What is involved in the recognition? A: Small RNA molecules that work with spliceosomes, The RN ...
... Q: How is this accomplished? A: Cells have mechanism that recognize introns. The most common is a spliceosome that recognizes the boundaries of intron-exon junctions and knows were to cleave and splice Q: What is involved in the recognition? A: Small RNA molecules that work with spliceosomes, The RN ...
Eukaryotic Gene Regulation
... cells because of a silencer that binds a cellular factor which repress transcription. However, in cells that are required to produce the hormone the effect of the silencer is itself neutralised by an enhancer located 1.2 kb upstream of the promoter of the gene and is only “activated” in the cells [t ...
... cells because of a silencer that binds a cellular factor which repress transcription. However, in cells that are required to produce the hormone the effect of the silencer is itself neutralised by an enhancer located 1.2 kb upstream of the promoter of the gene and is only “activated” in the cells [t ...
Chapt. 14 Eukaryotic mRNA processing I: splicing 14.1 Genes are in
... • Many eukaryotic transcripts have alternative splicing – can have profound effects on protein products: • Secreted or membrane-bound protein • Activity and inactivity ...
... • Many eukaryotic transcripts have alternative splicing – can have profound effects on protein products: • Secreted or membrane-bound protein • Activity and inactivity ...
Gene Section LPHN2 (latrophilin 2) Atlas of Genetics and Cytogenetics
... DNA/RNA Description LPHH1 consists of 19 commonly used coding exons. A further seven exons have been identified which may be alternatively spliced into the core backbone with variable frequencies and tissue specificities. At least a number of these additional exons are highly conserved in mammalian ...
... DNA/RNA Description LPHH1 consists of 19 commonly used coding exons. A further seven exons have been identified which may be alternatively spliced into the core backbone with variable frequencies and tissue specificities. At least a number of these additional exons are highly conserved in mammalian ...
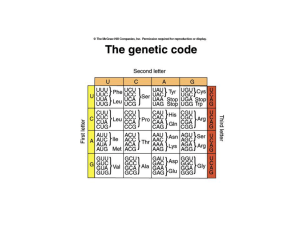
Features of the genetic code
... • Open reading frame starting at the initiation codon (AUG) • Each codon has 5’ base and a 3’ base e.g. 5’CGU3’ • Mutations that modify the genetic code are of 3 types: frameshift (include deletions and insertions), missense (lead to an amino acid replacement) and nonsense (mutation that generates a ...
... • Open reading frame starting at the initiation codon (AUG) • Each codon has 5’ base and a 3’ base e.g. 5’CGU3’ • Mutations that modify the genetic code are of 3 types: frameshift (include deletions and insertions), missense (lead to an amino acid replacement) and nonsense (mutation that generates a ...
Alternative splicing

Alternative splicing is a regulated process during gene expression that results in a single gene coding for multiple proteins. In this process, particular exons of a gene may be included within or excluded from the final, processed messenger RNA (mRNA) produced from that gene. Consequently the proteins translated from alternatively spliced mRNAs will contain differences in their amino acid sequence and, often, in their biological functions (see Figure). Notably, alternative splicing allows the human genome to direct the synthesis of many more proteins than would be expected from its 20,000 protein-coding genes. Alternative splicing is sometimes termed differential splicing.Alternative splicing occurs as a normal phenomenon in eukaryotes, where it greatly increases the biodiversity of proteins that can be encoded by the genome; in humans, ~95% of multi-exonic genes are alternatively spliced. There are numerous modes of alternative splicing observed, of which the most common is exon skipping. In this mode, a particular exon may be included in mRNAs under some conditions or in particular tissues, and omitted from the mRNA in others.The production of alternatively spliced mRNAs is regulated by a system of trans-acting proteins that bind to cis-acting sites on the primary transcript itself. Such proteins include splicing activators that promote the usage of a particular splice site, and splicing repressors that reduce the usage of a particular site. Mechanisms of alternative splicing are highly variable, and new examples are constantly being found, particularly through the use of high-throughput techniques. Researchers hope to fully elucidate the regulatory systems involved in splicing, so that alternative splicing products from a given gene under particular conditions could be predicted by a ""splicing code"".Abnormal variations in splicing are also implicated in disease; a large proportion of human genetic disorders result from splicing variants. Abnormal splicing variants are also thought to contribute to the development of cancer.