* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
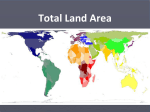
Download Microbial Source Tracking • Pathogen Contamination –What is it?
Phospholipid-derived fatty acids wikipedia , lookup
Bacterial cell structure wikipedia , lookup
Marine microorganism wikipedia , lookup
Disinfectant wikipedia , lookup
Bacterial morphological plasticity wikipedia , lookup
Metagenomics wikipedia , lookup
Human microbiota wikipedia , lookup
Microbial Source Tracking • Pathogen Contamination –What is it? –Where is it coming from? –Who is making it? –How can we address it? Tiered Approach Weight of Evidence Microbial Source Tracking Intensive sampling using standard FIB measurements; Infrared thermography Sanitary Survey Identification of impaired areas based on long-term monitoring Traditional Culturing Methods EPA Rapid Methods • Quantitative Polymerase Chain Reaction (qPCR) • 16S ribosomal RNA (16S rRNA) markers Method A targets Enterococcus Method B targets Bacteroidales qPCR in Microbial Source Tracking • 16S rRNA gene found in nearly all bacteria and Archaea • Small changes in genes allow for identification of hosts • qPCR allows for quantification of specific host inputs qPCR Basics • Low detection limit – Pro: very sensitive – Con: low levels are not likely to be health risks • Detects living and dead cells – Pro: conveys source information – Con: might be outdated source information • Can detect lots of sources – Pro: potentially unlimited host tracking ability – Con: each host detection requires a separate assay • Provides quantitative information – Pro: enables evaluation of relative contribution of host animals – Con: requires innovative calibration work Year OneMethod Validation Year Two Pilot Project: Withers Swash Source Tracking Strategies • Space and Time – Targeted wet and dry weather sampling • Tracers – Bacteria concentrations – Chemical concentrations • Water mass indicators • Optical brighteners • Caffeine – Host animals • Multiple antibiotic resistance of fecal bacteria • Genetic markers of fecal bacteria Illicit Discharge Detection and Elimination: A Guidance Manual for Program Development and Technical Assessments (CWP and Pitt, 2004) Sampling • 2 dry • 3 wet Nalgene stormwater sampler – First flush sampling Hobo Site-Specific Storm Hydrographs Hobo water level loggers Figure 3.3.1-1: Hydrograph showing the response to a rain event at Site 1 as recorded by an in12 situ Hobo™ water level logger. Figure 3.4.1-1: Concentration of Enterococcus (MPN/100 mL) as a function of sampling site. Black circles represent dry samples and white circles represent wet samples. Figure 3.4.1-4: Enterococcus concentrations (MPN/100 mL) in Withers Basin sub-watersheds. Figure 3.4.1-2: Enterococcus and E. coli concentrations Figure 3.4.2.2-1: BacHum concentrations (genome copies/100 mL) in Withers Basin sub-watersheds. Results below the reporting limit are in the dark green-shaded grouping. Figure 3.4.2.3-1: BacCan concentrations (genome copies/100 mL) in Withers Swash subwatersheds. Concentrations grouped into quartiles. Weight-of-evidence approach using indices • By Parameter – Data grouped into quartiles with the bottom bin set at regulatory threshold – Each data group assigned an integer value aka a “grade” Table 3.4.1-1: Concentration ranges used to assign rank orders to E. coli and Enterococcus concentrations. These values were used to create Table 3.4.1-2. Weight-of-evidence approach using indices • • By Parameter By Site – Each parameter assigned a “grade” – Parameter grades summed to determine site “grade” • Averaged Wet vs Dry • Averaged Overall Table 3.4.1-2. Fecal Indicator Bacteria (E. coli + Enterococcus) “Grades” Weight-of-evidence approach using indices • • • By Parameter By Site Visualizing sites to prioritize for remediation – Color coded matrixes – Qualitative rankings Table 3.4.3-3: Qualitative descriptors Weight-of-evidence approach using indices • • • By Parameter By Site Visualizing which sites to prioritize for remediation – Color coded matrixes – Qualitative rankings – Every site with a “very strong” detection of FIB had a “strong” or “very strong” detection for at least one of the qPCR assays Table 3.4.3-4: Summary of the results of the qualitative descriptors for regulatory FIB and qPCR markers. Sites with very strong qualitative rankings are in red font. Table 3.5.1-1: Sites with “strong” and “very strong” levels of BacHum concentrations. Percentages determined by dividing the average rank order by the maximum rank order for each tracer. Percentages greater than 50% are shaded red. Table 3.5.1-2: Sites with “very strong” levels of BacCan. Percentages determined by dividing the average rank order by the maximum rank order for each tracer. Table 3.6.4-1: Sites with strong evidence for either human or canine sourced Bacteroides. Citation: Wood, J., J.M. Trapp, S.M. Libes, and E.J. Burge. 2013. Watershed Assessment Report: Stormwater Management Planning: Development of a Pilot Investigative Approach to Remediate Bacterial Source Impairments along the Grand Strand. Final Report, prepared under the authority of Section 22 of the Water Resources Development Act of 1974 for the US Army Corps of Engineers, Charleston District; Horry County, SC; Georgetown County, SC; City of Myrtle Beach, SC; and City of North Myrtle Beach, SC. 100 pgs + Appendices. http://www.coastal.edu/envsci/projects/pollution/documents.html Thank you to our Partners Figure 3.4.2.1-1: GenBac concentrations (genome copies/100 mL) in Withers Basin sub-watersheds. Concentrations are grouped into quartiles. What we want our MST data to be Sources of bacterial pollution in Withers Swash “Universal (GenBac) Wildlife Human (BacHum) Dog (BacCan) Goose (CGO-1F) What our MST data actually are Goose (CGO-1F) Wildlife Dog (BacCan) “Universal” (GenBac) Human (BacHum) Sources of bacterial pollution in Withers Swash