* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Brandi Deptula Final Research Paper
Survey
Document related concepts
Horizontal gene transfer wikipedia , lookup
Phospholipid-derived fatty acids wikipedia , lookup
Magnetotactic bacteria wikipedia , lookup
Probiotics in children wikipedia , lookup
Bacterial cell structure wikipedia , lookup
Triclocarban wikipedia , lookup
Marine microorganism wikipedia , lookup
Bacterial morphological plasticity wikipedia , lookup
Metagenomics wikipedia , lookup
Bacterial taxonomy wikipedia , lookup
Transcript
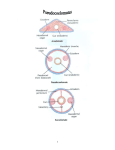
1 Diversity and function of bacteria related to the newly isolated organism JT5, a possible lignocellulose-degrading species from the gut of an evolutionarily higher termite Brandi Deptula University of Wisconsin Oshkosh Dr. Eric Matson, Mentor Department of Biology/Microbiology 2 Abstract Termites are one of the most successful groups of insect on the planet and are one of only two insects known capable of subsisting solely on lignocellulosic plant biomass. The remarkable digestive capability of termites is attributed to the highly complex microbial communities inhabiting their guts, and yet the vast majority of these microbes are not well understood due to an inability to cultivate them in the laboratory. Recently, a novel bacterium (strain JT5) related to the genus Dysgonomonas was isolated from an evolutionarily-higher termite, Gnathamitermes perplexus. This bacterium was determined to be an abundant member of the gut community and to possess genes for enzymes involved in lignocellulose degradation. We hypothesized that numerous bacteria related to JT5 would reside in G. perplexus and in a variety of other termite species. To test the hypothesis, we created inventories of SSU rRNA genes (a marker gene used for bacterial species identification) from G. perplexus gut community DNA. We targeted subsets of SSU rRNA genes in the gut bacterial population that are closely related to JT5 as well as subsets that are more broadly associated with the Dysgonomonas genus. We used similar methods to determine whether or not these SSU rRNA genes were present in gut community DNA samples from other termite species. Our investigations revealed a lower than expected diversity of bacteria predicted to share species-level relationships with JT5 in G. perplexus but a higher diversity at the genus level. We also found that many of the termite samples we analyzed contained bacteria predicted to be related to JT5. These results indicate that JT5 represents one of potentially several bacterial species within the genus Dysgonomonas that inhabit the guts of termite species. Thus, research on JT5 may broadly provide information on the role of bacterially-mediated lignocellulose degradation occurring in termites. 3 Introduction Investigating natural systems that break down plant biomass can provide us with new biotechnological applications (Okhuma, 2003). Due to the availability of large quantities of cellulosic plant materials, it is an attractive substrate for production of industrially important products. Most plant biomass is currently disposed of by burning (Gupta et al., 2011). The discovery of new approaches and systems for lignocellulose degradation would result in a cheaper and more favorable processing of plant materials into useful products. Producing biofuels out of woody crops, agricultural waste, and grasses poses the challenge of finding ways to breakdown the lignocellulose/xylan material. Hydrolytic enzymes are already being applied in industries to aid with agricultural and plant waste, and in chiral separation and ligand binding studies (Gupta et al., 2011). Recombinant organisms containing cellulose degrading and ethanol producing genes have been created, but as of yet they do not produce enough ethanol for commercial scale production (Vasan et al., 2010). New discoveries in this area would lead the way to more efficient renewable energies, and may also aid in better processing of paper materials in mills. When searching for ways to breakdown and extract plant polysaccharides, it is reasonable to look at the microbial systems found in the guts of organisms such as termites that flourish on a diet of lignocelluloses plant biomass (Gupta et al., 2011). Termites are subdivided into seven families. Six of these families are called lower termites and the seventh family, called higher termites, is made up of about 85% of the known species of termites (Matsui et al., 2009). Lower termites feed mainly on wood, utilizing the enzymes they make themselves, as well as those from bacteria, archaea, and protists in their guts to digest the wood (Matsui et al., 2009). Higher termites show a more diverse diet, including soil, grass, and self-cultivated fungi. Those subsisting on wood alone, are the minority of this 4 family. For this reason, higher termites are a good place to search for previously undiscovered gene activity. It is especially desirable to examine the worker class of termites, as they play the major role of foraging and breakdown of plant material (Matsui et al., 2009). The diversity of food sources utilized by higher termites is assumed to be related to the diversity of the prokaryotes they harbor in their gut. The variety of geographic habitats in which higher termites live was not found to change bacterial diversity among members of the same termite species (Husseneder et al., 2010). This suggests that termites need to retain a symbiotic gut community composed of a similar number and composition of species to fulfill their nutritional requirements. These species need not be identical in each termite; closely related microbial species or microbes with overlapping functions suffice. However, in laboratory termite colonies, a rapid shift in community structure was seen when varying the food source between high and low molecular weight carbon sources. In one such study, it was found that the gut population of a wood-feeding higher termite was dominated by spirochetes when fed wood or wood powder, Bacteroidetes when fed xylan, cellobiose, or glucose, and Firmicutes when fed xylose (Husseneder et al., 2010). While the activity of microbial species inhabiting the termite gut is largely responsible for the degradation of lignocellulose, and crucial to the ability to derive carbon and energy from plant biomass, the termite gut microbial community is responsible for many other important functions. For example, the gut microbiome is responsible for the production of acetate (the principal source of nutrition taken up by the termite) from hydrogen and carbon dioxide, as well as nitrogen fixation and recycling of uric acid (Ohkuma & Kudo, 1996). Wood is a poor source of nitrogen, thus, most termite species that live solely on woody material derive their nitrogen from nitrogen-fixing bacteria in their hindgut. Nitrogen fixing bacterial species have been 5 isolated from both higher and lower termites. The waste product, uric acid, is converted by bacteria into ammonia in order to recycle nitrogen containing compounds (Matsui et al., 2009). The termite gut is generally composed of three major compartments: the foregut, the midgut, and the hindgut or lumen. The hindgut is the largest section, comprising a large amount of the termite's weight, and contains the most dense and diverse quantity of symbionts (Matsui et al., 2009). Additional gut compartments vary based on species of termite. One example is that the digestive tracts of termites that belong to the genus Cubitermes, members of the higher termite family. In these species, the gut is further separated into distinct hindgut compartments containing variations in microbial community makeup (Schmitt-Wagner et al., 2003). The large number of potential niches allows for the possibility of species overlap between the sections. However, solid evidence on the role of possible functional redundancy of microbial communities in the termite gut is as of yet lacking (Yang et al., 2005). The hindgut paunch is an example of one gut compartment displaying varied microenvironments. Due to oxygen consumption by different prokaryotes within, there is a decrease in oxygen concentration inward from the gut wall towards a completely anoxic center (Brune et al., 1995). Hydrogen concentrations, consumption, and production are similarly stratified throughout the termite gut. The presence of oxygen impacts the hydrogen concentrations, which is explained by the presence of different species existing in different areas of the gut. Some species present are oxidizing hydrogen, while others are producing hydrogen as a byproduct of fermentative metabolism (Ebert & Brune, 1997). Studies using microsensors and radiotracers show that the gut compartments are separate functional zones for microbial activity. Such processes include hydrogen production, methanogenesis, and acetogenesis. It was noted that hydrogen accumulates in the anterior hindgut of all Cubitermes species investigated. The 6 population of bacteria related to clostridia, which are known fermentative bacteria, may explain the source of increased hydrogen (Schmitt-Wagner et al., 2003). The special organization of different bacteria in the termite guts have also been shown to control intestinal carbon flux (Yang et al., 2005). Such features of the gut underscore the variety of roles microbial species perform in the termite gut and their interdependence on each other (Ebert & Brune, 1997). The termite gut epithelium is the most probable location for oxygen diffusion into the gut. Oxygen concentration drops steadily on approaching the center of the gut which is anoxic. In between the wall and center is a microoxic environment of decreasing oxygen concentration. The anaerobic central portion allows for the presence of strict anaerobes to inhabit this niche. Many bacterial inhabitants of the oxygenic gut wall are aerobes or facultative anaerobes, however, members of Bacteroidetes and Clostridiales, which are often strict anaerobes, are found to be abundant along the gut wall. This suggests they may be at least partially tolerant to microoxic conditions (Nakajima et al, 2005). These findings highlight the importance of the challenging task of investigating the locations of different bacterial species in situ, in a way that doesn't disturb the internal structure of the gut community. Studies have been conducted in an attempt to categorize the microbial diversity from a variety of different termites. Very few numerically significant bacteria from higher termites have ever been grown in pure culture (Okhuma, 2003). Most of the isolated organisms belong to one of four phylogenetic divisions within the domain Bacteria. These included Proteobacteria, Spirochetes, Bacteriodetes, and low G-C content gram positive bacteria. More current studies state that the prokaryote inhabitants of termite guts are represented by organisms belonging to the phyla Actinobacteria, Firmicutes, Bacteroides, Proteobacteria, and Spirochetes, as well as various Archea (Matteotti et al., 2012; Yang et al., 2005; Schmitt-Wagner et al. ,2003). One 7 study identified species that couldn't be assigned to any known major groups of bacteria. This study also found that two thirds of the distinct species identified had less than 90% sequence identity to any known 16s rDNA sequence data of cultured organisms while another 10 species had no close similarity to any recognized bacterial phylum in rRNA databases (Ohkuma & Kudo, 1996). Additionally, analysis of bacterial clone lineages showed they were most closely related to bacteria isolated from closely related termites, suggesting co-evolution between termites and their gut microbes (Hongoh, 2010; Schmitt-Wagner et al., 2003; Yang et al., 2005). A study of the phylogeny and termite gut niche location in Reticulitermes santonensis analyzed fractions of gut sections to compare the diversity and community structure of each section. More than 200 different restriction patterns were found by analyzing the gut of this wood-feeding termite using restriction fragment length polymorphism analysis. Species richness of hindgut fluid was highest, while in contrast, it was lowest in the midgut. Of all the sections sampled, it was implied that only in the hindgut would further sampling increase the number of new species identified (Yang et al., 2005). Regarding phylogenic location studies, it has been seen that the steep increase in gut pH in the midgut to hindgut transition correlates with a steep drop in microbial density. This evidence of changing physiochemical conditions between the mid and hindgut represents a probable barrier for microbes, preventing passage between gut compartments (Schmitt-Wagner et al., 2003). Figure 1 shows the phylogenic placement of isolate JT5. Of all species studies, Reticulitermes speratus, a lower termite, has been the most studied in regards to bacterial diversity in the gut. More than 300 bacterial phylotypes have been reported, and has an estimated species per termite as high as up to 700 different species per individual termite. This study especially focused on the microbial community niche of the gut 8 epithelium. In this species, methanogenic Archaea exclusively colonized the gut wall, whereas, in other species they are found within the cells of gut protists. Due to the oxygen and hydrogen gradients originating at the gut epithelium, bacteria living in this niche are considered unlike ones that are able to freely move throughout the gut. Findings such as these also show proof of the formation of distinct niches within the termite guts (Nakajima et al., 2005). Abundant groups found along the gut epithelium, including Actinobacteria, Bacteroidetes, Clostridiales, and Lactococcus, are non-motile species. However, Bacteroidetes were found in significant numbers in all gut segments analyzed, except for the protozoan fraction (Nakajima et al., 2005; Yang et al., 2005). Groups found in the gut lumen, such as Spirochaetes and Desulfovibrio, have a high level of motility. This may be necessary to counteract the fluid dynamics of the gut and stay in the niche they're adapted to (Nakajima et al., 2005). Comparisons of two soil-feeding higher termites of the genus Cubitermes were undertaken to compare the abundance and distribution of bacterial species in their gut tract with the intent to discover changes in microbiota along the passage through the gut and to compare the different bacterial species in the homologous gut compartments of the two species (SchmittWagner et al., 2003). Cocci and short rods were found to be dominant in all gut sections. Spirochetes were also found in significant number in the P 1 , P3, and P4 hindgut compartments. The first section of the hindgut, P 1, contained mainly low G+C content gram positive bacteria. The amount of this type of bacteria decreased progressively through the posterior hindgut sections, while more diversity of species in the Proteobacteria, Cytophaga-FlexibacterBacteriodetes, and spirochetes was noted. Highest species diversity was observed in the P4 hindgut compartment. Despite the decrease in low G+C content gram positive bacteria upon moving toward the posterior gut, one half of the clones in the P5 section were found to be in this 9 category. One third of the total from the P5 section were associated with Proteobacteria, Cytophaga-Flexibacter, and Bacteriodetes. The increase in these phyla along the hindgut correlates with the increase of acetate, lactate, and succinate in the P3 segment of the Cubitermes gut. Finally, it was indicated that Actinomycetes did not play a major role in the guts of these termites, as no clones of this type were found in this study. This is curious because Actinomycetes showing cellulolytic and lignin-solubilizing activity, have been isolated in the past from soil feeding termites (Schmitt-Wagner et al. 2003). While it has been proven that termites produce their own celluloltytic enzymes in their salivary glands or midgut epithelium to initiate the digestive process, microbial flora in the digestive tract is necessary for termite survival (Matsui et al., 2009; Scharf et al., 2010; Yang et al., 2005). It is well known that phylogenetically lower termites contain symbiotic protozoan flagelletes that are contributing a significant amount of cellulases, as several cellulase genes have been found in termite gut protozoa (Matsui et al., 2009; Wenzel et al., 2002). Termites that cultivate fungus are known to consume fungal cellulases to aid in the breakdown of lignocellulose. It is presumed that in higher termites, which lack flagellate protozoa, the contribution of lignocelluloses-degrading enzymes from bacteria must be higher (Matsui et al., 2009; Warneke et al., 2009). Cellulose is broken down by a series of reactions involving three separate cellulases: endo-beta-1,4-glucanses (EG), cellobiohydrolases (CBH), and beta-glucosidases (BGL). Hemicellulose is broken down by hemicellulases: endo-beta-1-4-xylanase, beta-xylosidase, and alphaglucuronidase (Matteotti et al., 2010). Endo-beta-1,4-glucanase is an essential enzyme in the breakdown of cellulose because it begins the breakdown of cellulose into smaller cellulose molecules (Cho et al., 2010). This is the enzyme at the beginning of the chain of cellulose 10 breakdown, and is the enzyme most often found produced by the termite itself. Endo-beta-1-4glucanase is found in highest concentrations in the salivary glands of lower termites, and in the mid-gut of higher termites (Matsui et al., 2009). This provides reaction sites for CBH to break cellulose down further into oligosaccharides or cellobiose. Finally, the products of the CBH reaction are hydrolyzed by BGL to yield glucose (Cho et al., 2010). Investigations on termite gut communities have identified over 700 genes predicted to encode enzymes likely contributing to the breakdown of lignocellulose, spanning at least 45 different carbohydrate active enzyme families (Warnecke et al., 2007). For example, two genera of lower termites have yielded diverse species of Gram-positive and Gram-negative bacteria that were able to be cultivated, and possessed beta-glucosidase, cellobiohydrolase, beta-xylosidase, and Carboxymethylcellulase enzyme activity (Matteotti et al., 2012). Cellulose-degrading bacteria from the gut of Zootemopsis angusticollis have been identified and quantified. Serial dilutions were preformed from the gut contents of termites that were fed either wood or fiber paper. After enrichment on carboxylmethylcellulose or filter paper, 119 cellulolytic bacterial strains belonging to 23 different groups were isolated. Many of the isolates hydrolyzed both carboxylmethylcellulose and the filter paper (Wenzel et al., 2002). Many other species are unable to be cultivated as of yet, and while attempts to sequence the entire genome of uncultivable bacteria from termite guts have been successful, enzyme activity is often presumed by genomic analysis (Hongoh, 2010). The guts of termites contain a wealth of highly specialized co-evolved symbionts. Further research on culturing these species and analyzing their functional roles in the termite gut will likely divulge a tremendous amount of information on how these gut communities function. Combined with cultivation-independent approaches, this research will lead to a better 11 understanding of termite ecology. Such pursuits advance our understanding of complex symbiosis found in the guts of various termite species and may ultimately lead to new design approaches to biomaterial processing inspired by termite gut systems. The termite gut systems may even provide us with superior organisms and enzymes for biofuels production, nitrogen processing, and methane generation. Methods Bacterial Cultivation To examine the growth rate of isolate JT5 on four different carbon sources, anaerobic tubes were amended with 50 ul of vitamin/cofactor mix (Leadbetter et al., 1999). Two tubes were amended with 400 ul of 250 mM Cellobiose, two tubes with 200 ul of 0.5 M Glucose, two tubes with 200 ul of 0.5M Xylose, and two tubes with 400u1 Carboxymethylcellulose(CMC) to equal 36mg CMC per tube, providing a carbon source equivalent to the other tubes . Absorption readings were taken using a Spectronic 21D (Milton Roy) Spectrophotometer. 200 ul of a log phase culture of JT5 was added to each tube and another absorption reading was taken. Tubes were incubated at 27°C. Daily readings were taken for 14 days. Logarithmic Growth curves were plotted and generation time during log-phase growth was calculated using Microsoft excel. PCR Amplification Whole gut community DNA samples were diluted to equal 1Ong/u1 concentrations. Each sample was combined with 10 ul fail-safe Buffer D (Epicentre, Madison,WI), 0.25 ul Taq polymerase (New England Biolabs, Ipswich, MA), 7.25u1 molecular grade water, and 1 ul each 12 forward and reverse primer, into a 20 ul reaction volume. The samples were amplified by Polymerase Chain Reaction (PCR) using 27F/1492R universal bacterial primers with the ability to amplify all bacterial DNA present (Lane et al., 1991). A second PCR reaction was prepared using 591F/1392R specific dysgonomonas primers to amplify only DNA similar to isolate JT5 (5'-ACTGCGGGTCTTGAGTATGG-3' and 5'-GCGGTTACGCACTTCAGATA-3', respectively). A third PCR reaction was prepared using 627F/1492R to amplify DNA representative of the genus Dsygonomonas, which includes isolate JT5. A control tube of PCR water was created for the universal primer set as a negative control. PCR was performed using a T100 Thermocycler (Bio-rad, Hercules, CA). PCR conditions were 95°C denaturation, 58°C annealing, and 72°C elongation temperatures for 30 cycles. PCR products were visualized using gel electrophoresis through 0.7% agarose gels. Gel was post stained for visualization with 0.15mg/m1 Ethidium Bromide for 20 minutes. The image was viewed using Biorad Gel Doc set to UV filter. Cellular transformation & Clone library amplification Topo TA Cloning Kits were used to clone PCR products from 591F/1392R and 627F/1492R. Cells were plated on LB agar containing 10Oug/m1 Ampicillian too select for transformants. After growth at 37°C for 24 hours, 96 transformant clones were selected. PCR was performed on each colony that resulted in successful transformation, using T7 and T3 primers designed to target Topo vector. A denaturation temperature of 95°C, annealing temperature of 55°C, and elongation temperature of 72°C were used for 30 cycles. Restriction Digest 13 For each set of PCR products, a restriction digest was performed using 1.6u1 molecular grade water, 0.2 ul MSPI enzyme (New England Biolabs, Ipswich, MA), Buffer 4 (New England Biolabs, Ipswich, MA), and BSA (New England Biolabs, Ipswich, MA) to equal a 2 ul reaction mixture. To the mixture, 5 ul of each PCR product was added, and the reaction proceeded for 3 hours at 37°C. Restriction digest products were visualized using gel electrophoresis through 2.5% agarose gels. Gels were post stained for visualization with Ethidium Bromide for 30 minutes. The image was viewed using Biorad molecular imaging FX scanner to visualize restriction fragment length polymorphism (RFLP) patterns Vector extraction Unique RFLP types were cultured from the stored libraries in LB broth containing 10Oug/m1 Ampicillian and shaken at 190 rpm for 17 hours. Each culture was divided into sterile ependorf microfuge tubes and centrifuged for 3 minutes at 8,000 x g. Supernatent was removed and the remaining pellet was processed using QIA prep Spin Miniprep Kit (QIAGEN). Final extracted vector samples were sent for genome sequencing by The University of Wisconsin Biotechnology Center, Madison, WI. Termite gut comparisons A whole gut sample °Ong/up from 8 termite species had DNA amplified using PCR using JT5 DNA Ong/0 as a positive control and water as a negative control. Each sample was prepared in triplicate using 27F/1492R, 627F/1492R, and 591F/1392R primers. Temperatures were 95°C denaturation, 58°C annealing, and 68°C elongation for 28 cycles. Products were visualized using gel electrophoresis through 1% agarose gel. Gel was post stained for 14 visualization with Ethidium Bromide for 20 minutes. The image was viewed using Biorad molecular imaging FX scanner and Biorad GelDoc. Results and Discussion Analysis of generation times was performed by measuring optical density in three separate experiments for 13 days, 10 days, and 12 days respectively. For the first experiment, cellobiose readings were recorded for 13 days, whereas the glucose samples showed growth up to 21 days. Generation time was calculated by plugging the formula (LOGI° (reading 2) — LOGio( reading 1))/(0.301 * (time 2-time 1)) into Microsoft Excell. For each substrate, 4 separate generation times were calculated, and averaged together. Generation times were as follows: Glucose 28.56 hours, Cellobiose 25.78 hours, Xylose 32.2 hours, and CMC 30.72 hours. JT5 is capable of growth on all carbon sources we provided. Readings for CMC did not appear to follow the standard bacterial growth curve as did the others. It appeared to have a short rapid log phase, followed by a steady decline in optical density. It is unsure if this is due to bacterial cell death or decrease in substrate due to bacterial consumption. As CMC is used in experiments to induce expression of cellulolytic enzymes, possible upregulation of these enzymes could be increasing consumption of the carbon source used. After an initial log phase growth, demonstrated by increased optical density readings, the readings showed more fluctuation up and down before decreasing steadily. The CMC media appeared to lose viscosity over the course of bacterial growth. Regardless, we have shown that JT5 can be cultured on various carbon sources, which suggests a variety of enzymes required for the breakdown of plant-based sugars. PCR amplification of a whole gut sample was run against 27F/1492R, 591F/1392R, and 627F/1492R. Product from each reaction was cloned into E. coli and plated out on LB agar 15 containing 10Oug/m1 Ampicillian, followed by the selection of 96 individual colonies. Restriction digest was performed, resulting in an 84% yield for the universal bacteria library, 89% yield for the Dysgonomonas library, and 66% yield for the JT5-specific library. (Table 1) The universal bacterial library was made up of 70 unique clones. The Dysgonomonas library contained 17 unique clones. The JT5-specific library showed only 2 unique types of clones. A universal bacterial library of 70 different clones, reinforces our expectations that the termite gut habitat is comprised of a wide variety of bacteria. Finding 17 unique clones in the Dysgonomonas library suggests that members of the genus Dysgonomonas are abundant and diverse members of G. perplexus' gut community. (Table 1) JT5-like species were found to be present in only 2 unique clones, dismissing our hypothesis that they are diverse in termite guts. However, it still appears that they are abundant in the gut of G. perplexus. PCR amplification of a whole gut sample from 8 different termites was run against 27F/1492R, 591F/1392R, and 627F/1492R. Higher termites sampled were: Gnathamitermes perplexus, Nasutitermes sp., Rhynchotermes sp., Microcerotermes sp., and Amitermes sp. Lower termites sampled were: Insisitermes minor, Reticulitermes hesperus, & Zootermopsis nevadensis. Positive results were seen in all specimens for the universal 27F/1492R bacterial primer set as expected. Positive results for Dysgonomonas specific primers (627F/1492R) were present in all 5 higher termites, and the lower termite Reticulitermes hesperus, as well as a faintly positive result in the lower termite Insisitermes minor. Dysgonomonas presence in higher termites matches with our expectation that they are abundant and play a role in lignocellulose breakdown. A low positive signal in the two lower termites shows that this genus is not only contained in members of higher termites, but is also present to a lesser degree in some lower termites. A positive result for presence of JT5 was found only in the higher termite Gnathamitermes perplexus. According 16 to these results, Dsygonomonas appears to be widespread in the guts of numerous termites, to a higher degree in higher termites, and a lower degree in lower termites. In contrast, JT5 was only identified in G. perplexus gut habitat, suggesting that it is likely unique to this termite. (Figure 3) PCR amplification of a sample from the proctedal segment 2A (P2A), diverticulum (DV), and proctedal segment3 (P3P) sections of G. perplexus gut was run against 27F/1492R, 591F/1392R, and 627F/1492R. All 3 sections showed positive results for the universal bacterial primer set as expected and for Dysgonomonas specific primers. Dysgonomonas signal was weakest in the P2A segment and strongest in the DV segment. Indication of Dysgonomonas in all three tested gut sections supports our idea that the genus is widespread and abundant. A positive result for JT5 specific primer was seen only in the P3P segment (Figure 3) indicating that the P3P section is this bacterium's specialized niche. As expected, universal bacterial presence was seen in all three gut segments. Dysgonomonas was also present in all three sections, suggesting the genus is an important player in the digestion of lignocellulose by termites. Likely it plays more of a role in the DV segment, as the signal was strongest there, and a lesser role in the P2A segment, where the signal was weakest. JT5 presence was only identified in the P3P segment, suggesting its role in lignocellulose breakdown occurs after other processing has occurred in the fore- and mid-gut. Conclusions We have observed that representatives of the genus Dysgonomonas are abundant, diverse, and widespread in the gut tracts of termites. This is based on diversity analysis of the G. perplexus whole gut, and comparison testing of various termite guts for the presence of Dysgonomonas and JT5. We also found, contrary to our belief, that JT5-like species are not 17 diverse, but it is likely they are abundant based on our studies. It has also possible that JT5 may be unique to the termite G. perplexus since we only obtained a positive result for JT5-like species in this termite. We identified that JT5-like species are specifically located in the hindgut (posterior) portion of the G. perplexus gut. JT5 is could play a significant role in lignocellulose degradation and warrants additional research into its metabolic capabilities. We are interested in further studies of glycohydrolyase gene expression. We would like to understand if the genes are being expressed in the termite, and under what conditions they are being expressed. There is a lot of potential for future studies in cloning these genes to develop better processing of plant materials in the creation of biofuels. 18 References Brune, A., Emerson, D., & Breznak, J. (1995). The termite gut microflora as an oxygen sink: Burnum, microelectrode determination of oxygen and pH gradients in guts of lower and higher termites. Applied and Environmental Microbiology, 61, 2681-2687. Burnum, K.E., Callister, S.J., Nicora, C.D., Purvine, S.O., Hugenholtz, P., Warnecke, F., . . . Lipton, M.S. (2011). Proteome insights into the symbiotic relationship between a captive colony of Nasutitermes corniger and its hindgut microbiome. The ISME Journal, 5, 161- 164. Cho, M., Kim, Y., Shin, K., Kim, Y., Kim, Y., Kim, T. (2010). Symbiotic adaptation of bacteria in the gut of Reticulitermes speratus: Low endo-beta-1,4-glucanase activity. Biochemical and Biophysical Research Communications. 395, 432-435. Ebert, A. & Brune, A. (1997). Hydrogen Concentration Profiles at the Oxic-Anoxic Interface: a Microsensor Study of the Hindgut of the Wood-Feeding Lower Termite Reticulitermes flavipes (Kollar). Appl. Environ.MicrobioL 63, 4039 — 4046. Gupta, P., Samant, K., & Sahu, A. (2011). Isolation of Cellulose-Degrading Bacteria and Determination of Their Cellulolytic Potential. International Journal of Microbiology. Husseneder, C., Ho, H., & Blackwell, M. (2010). Comparison of the Bacterial Symbiont Composition of The Formosan Subterranean Termite from its Native and Introduced Range. The Open Microbiology Journal, 4, 53-66. Hongoh, Yuichi. (2010). Diversity and Genomes of Uncultured Microbial Symbionts in the Termite Gut. Bioscience, Biotechnology, Biochemistry. 74, 1145-1151. Mathew, G.M., Ju, Y., Lai, C., Mathew, D., & Huang, C. (2011). Microbial community analysis in thetermite gut and fungus comb of Odontotermes formosanus: the implication of Bacillus as mutualists. FEMS Microbial Ecology. 79, 504-517. Lane, D.J. (1991). 16S/23 rRNA sequencing, p. 115-175. In E. Stackebrant and M. Goodfellow (ed), Nucleic acid techniques in bacterial systematics. John Wiley & Sons, Inc., New York. Leadbetter, J.R., Schmidt, T.M., Graber, J.R., and Breznak, J.A. (1999) Acetogenesis from H2 plus CO2 19 by spirochetes from termite guts. Science 283: 686-689. Matsui, T., Tokuda, G., & Shinzato, N. (2009). Termites as Functional Gene Resources. Recent Patents on Biotechnology. 3, 10-18. Matteotti, C., Haubruge, E., Thonart, P., Francis, F., De Pauw, E., Portelle, D., & Vandenbol, M. (2010). Characterization of a new Beta-glucosidase/Beta-xylosidase from the gut microbiota of the Termite (Reticulitermes santonensis). FEMS Microbiology Letters. 314, 147-157. Matteotii, C., Bauwens, J., Brasseur, C., Tarayre, C., Thonart, P., Destain, J.,....Vandenbol, M. (2012). Identification and characterization of a new xylanase from Gram-positive bacteria isolate from termite gut (Reticulitermes santonensis). Protein Expression and Purification. 83, 117-127. Nakajima, H., Hongoh, Y., Usami, R., Kudo, T., & Ohkuma, M. (2005). Spatial distribution of bacterial phylotypes in the gut of the termite Reticulitermes speratus and the bacterial community colonizing the gut epithelium. FEMS Microbiology Ecology. 54, 247-255. Ohkuma, M. (2003). Termite symbiotic systems: efficient bio-recycling of lignocellulose. AppL Microbiol. Biotechnol., 61, 1-9. Okhuma, M. & Kudo, T. (1996). Phylogenic Diversity of the Intestinal Bacterial Community in the Termite Reticulitermes speratus. Applied and Environmental Microbiology. 62,461 -468. Scharf, M.E., Kovaleva, E.S., Jadhao, S., Campbell, J.H., Buchman, G.W., & Boucias, D.G. (2010). Functional and translational analyses of beta-glucosidase gene (glycosyl hydrolase family 1) Isolated from the gut of the lower termite Reticulitermes flavipes. Insect Biochemistry and Molecular Biology. 40, 611-620. Schmitt-Wagner, D., Friedrich, M.W., Wagner, B., & Brune, A. (2003). Phylogenetic Diversity, Abundance, and Axial Distribution of Bacteria in the Intestinal Tract of Two Soil-Feeding Termites (Cubitermes spp). Applied Environmental Microbiology. 69(10), 6007-6017 Vasan, P.T., Piriya, P.S., Prabhu, D., & Vennison, S.J. (2010). Cellulosic ethanol production by Zymomonas mobilis harboring an endoglucanase gene from Enterobacter cloacae. Bioresource Technology. 102, 2585 — 2589. 20 Warneke, F., Luginbuhl, P., Ivanova, N., Ghassemian, M., Richardson, T.H., Stege, J.T., Cayouette, M...Leadbetter, J.R. (2007). Metagenomic and functional analysis of hindgut microbiota of a wood-feeding higher termite. Nature. 450, 560-564. Wenzel, M., Schonig, I., Berchtold, M., Kampfer, P., & Konig, H. (2002). Aerobic and facultatively Anaerobic cellulolytic bacteria from the gut of the termite Zootermopsis angusticollis. Journal of Applied Microbiology. 92, 32-40. Yang, H., Schmitt-Wagner, D., Stingl, U., & Brune, A. (2005). Niche heterogeneity determines bacterial community structure in the termite gut (Reticulitermes santonensis). Environmental Microbiology. 7(7), 916-932. 21 Figure 1. Isolate JT5 is a member of phylum Bacteroidetes and groups with genus Dysgonomonas within the family Porphyromonadacaea. The DNA distance tree was constructed in ARB using the Kimura algorithm and was based on bacterial SSU rRNA sequences withl, 204 aligned nucleotides for each sequence. Bootstrap values (based on 100 replicates) are given when >75%. Scale bar represents 10% difference in nucleotide sequences. Bacteria from the phyla Proteobacteria and Spirochaetes are used as outgroups. 22 Figure 2. RFLP analysis of Dysgonomonas library. PCR products and restriction digested PCR products are shown in the upper panel. The lower panel shows the analysis of the library, indicating 17 different RFLP types. The most abundant of which (DI) comprised over 25% of the inventory. 23 Figure 3. Isolate JT5 inhabits the anterior portion of the G. perplexus gut. PCR amplifications of different gut sections show a JT5 signal only in the posterior section using JT5-specific primers (J) while a strong signal was observed for the posterior section, the P2 section and, to a lesser extent, the anterior gut section using Dysgonomonas-specific primers. As expected, all sections produced a strong signal when universal bacterial primers (U) were used. 24 Figure 1. Dysgonomonas gadei uncultured Bacteroidetes bacterium Dysgonomonas hoistadii Dysgonomonas mossii Dysgonomonas wimpennyt Isolate JT5 Dysgonomonas capnocytophagoides uncultured Bacteroidetes bacterium uncultured bacterium 1. r Bacteroides cellulosilyticus Parabacteroides goldsteinii Famil y:Porphyromonadaceae uncultured Bacteroidetes bacterium Porphyromonas gingival's Cytophaga xylanolytica Marinifilum fragile Rikenella microfusus Sphingobacterium multivorum Jo" IOU Cellulophaga lytica Flavobacteriumjohnsoniae Flexibacter polymorphus Kiebsiella pneumoniae subsp. pneumoniae Pseudomonas fluorescens Ralstonsa pickethi Variovorax paradoxus Pelagibactenum halotolerans Rhodopseudomonas rhenobacensrs Desulfobulbus propionicus Campylobactercoh Helicobacter pylon 1 00 Borreha burgdorien Spirochaeta aura nha subsp aurantia Treponema azotonutricium Treponema succinifaciens Leptospira interrogans serovar Icterohaemorrhagiae 0.1 25 Figure 2. ****** -0** * * * **.*************.* ***** **0 -*** 1***** -** 4***** * w 01* .. YIP . W. MWM M. Qu a ntity of e ac h 1 %..4 140 w . • M 11:koniftpos.,I, : 8 11401 411 X- RR 25 20 a) 15 a 10 5 0 1 I 2 V=41 I r 3 4 5 6 7 8 as 4■■■4 1 41.4 4iwp- 9 10 11 12 13 14 15 16 17 Number of unique clones 26 Figure 3. P2 Sample Posterior Sample Anterior Sample 1 mm Anterior Sample P2 Sample Posterior Sample 27 Table 1. Table 1 Termite Species Ecosystem Food Warm temperate desert Dry grass, soil Soil Contact Universal Bacterial Dysgonomonas High +++ +++ +++ +++ +++ ++ ++ Higher Termites Gnathamitermes perplexus Rhynchotermes sp. Cost004 Wet rainforest transition Leaf litter Wet rainforest transition Roots, Soil Amitermes sp. Cost010 Nasutitermes sp. Cost003 Wet rainforest transition Wood Microcerotermes sp. Cost008 Lowland moist forest Palm High High Low Low Lower Termites lnsisitermes minor Low-altitude residential Wood High Reticulitermes hesperus Zoo termopsis nevadensis High-altitude pine forrest Wood High High-altitude pine forrest Wood Low +++ +++ +++ Relative signal intensity given by ( + ) symbols. Where the signal is indicated as ( - ), no PCR amplification was observed. JTS-Specific