* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download No Slide Title
Henipavirus wikipedia , lookup
Orthohantavirus wikipedia , lookup
Influenza A virus wikipedia , lookup
Hepatitis B wikipedia , lookup
Epidemiology of HIV/AIDS wikipedia , lookup
Microbicides for sexually transmitted diseases wikipedia , lookup
Diagnosis of HIV/AIDS wikipedia , lookup
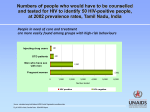
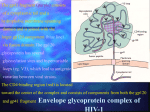
HIV Structure and Function Background: Basic Virology and Pathogenesis Structure: Virion structure, genomic structure, and accessory molecules Lifecycle: Infection and Expression Viruses Microscopic infectious agents that can infect the cells of a biological organism. Viruses can only replicate themselves by infecting a host cell and are incapable to reproduce on their own. A complete viral particle, known as a virion consists of nucleic acid surrounded by a protective coat of protein called a capsid. Types of Viruses (Baltimore Classification) I: Double-stranded DNA (Adenoviruses; Herpesviruses; Poxviruses, etc) Herpesviridae (Herpes, CMV, EBV), Poxviridae (Smallpox, Chickenpox, Vaccinia), Papilloma virus, Adenovirus II: Single-stranded (+) sense DNA (Parvoviruses) Erythema infectiosum, Phages III: Double-stranded RNA (Reoviruses; Birnaviruses) Rotavirus, Reovirus IV: Single-stranded (+) sense RNA (Picornaviruses; Togaviruses, etc) Polio, SARS, Hep A, Hep C, Rubella, Yellow fever V: Single-stranded (-) sense RNA (Orthomyxoviruses, Rhabdoviruses, etc) Rubella, Influenza, Rabies, Measles, Mumps, Ebola VI: Single-stranded (+) sense RNA with DNA intermediate (Retroviruses) HTLV, HIV VII: Double-stranded DNA with RNA intermediate (Hepadnaviruses) Hep B HIV Classification Group: Group VI (ssRNA-RT) Family: Retroviridae Genus: Lentivirus (Enveloped) Species: Human immunodeficiency virus 1 Species: Human immunodeficiency virus 2 Species Virulence Infectivity Prevalence Inferred origin HIV-1 HIV-2 High Low Global West Africa Chimpanzee Sooty Mangabey High Lower CRF14_BG HIV-1 Tropism www.hivmedicine.com/textbook/pathogen.htm Human immunodeficiency virus (HIV) is a retrovirus that causes acquired immunodeficiency syndrome (AIDS). AIDS has killed nearly 30 million people since it was first recognized in 1981. At the end of 2010, an estimated 34 million people were living with HIV worldwide. In 2010 alone, AIDS claimed an estimated 1.8 million lives, of which 250,000 were children. HIV primarily infects vital cells in the human immune system such as helper T cells (CD4+ T cells), macrophages and dendritic cells. HIV infection leads to low levels of CD4+ T cells. http://www.unaids.org/en/ Transmission Sexual route. Blood or blood product route. Mother-to-child transmission (MTCT). Viral RNA gp120 p24 RT & other virion proteins Fusion & Entry Binding Reverse transcription CD4 CXCR4 Nuclear localization & entry Integration Viral RNA gp120 p24 Cellular Activation Assembly RT & other virion prteins Post-translational processing Budding Translation Viral Gene Transcription HIV Structure Virion Genomic Proteomic HIV Structure SIV HIV HIV Structure Virion Genomic Proteomic Viral RNA gp120 p24 RT & other virion proteins Fusion & Entry Binding Reverse transcription CD4 CXCR4 Nuclear localization & entry Integration HIV-1 RNA Nature Reviews Microbiology 2, 461-472 (June 2004) Nature 460, 711-716 (6 August 2009) HIV-1 Integrated DNA Source: The AmFAR AIDS Handbook, D Ward, pp. 348 RNA Splicing Translational Frameshift Source: The Molecular Biology of HIV/AIDS, D Ward, pp. 19 Dr. Isabelle BOUALLAGA Institut Pasteur. http://www.123bio.net/ Source: Atlas of Infectious Diseases, Mandell & Mildvan (ed.), pp. 3.13 Source: Atlas of Infectious Diseases, Mandell & Mildvan (ed.), pp. 3.13 HIV Structure Virion Genomic Proteomic HIV Proteins Structural Proteins Gag: Matrix, Capsid, NC, p6 Pol: Protease, Reverse Transcriptase, Integrase Env: gp120, gp41 Regulatory Proteins Tat Rev Nef Accessory proteins Vif Vpr Vpu Source: The Molecular Biology of HIV/AIDS, D Ward, pp. 19 Source: BioAfrica Bioinformatics for HIV Research. http://bioafrica.mrc.ac.za/ Gene/Protein gag (Pr) 55gag Mass, KDa Function p55 Gag precursor protein MA - Matrix p17 Aids nuclear import and viral assembly CA - Capsid p24 HIV central core – contains HIV genome and enzymes NC - Nucleocapsid p15 (p6,p9) p6 – Precise location in virion unknown, not generally present in other retroviruses, may direct proteins to secretory path of cell. p9 – Assoc. with HIV RNA, involved in packaging of HIV RNA into virions myristate Association with membrane Matrix p17 Envelope Incorporation Viral Entry RNA binding RNA packing Formation of Gag multimers Capsid p24 p2 NC p9 p1 p6 Viral Budding Source: BioAfrica Bioinformatics for HIV Research. http://bioafrica.mrc.ac.za/ Gene/Protein Mass, KDa Function pol (Pr) 160gag-pol p160 Gag-Pol precursor protein PR - protease p10 Cleaves Gag and Gag-Pol IN - integrase p32 Integrates viral genome into host DNA RT – reverse transcriptase p66/p51 p66 – copies RNA genome. RNAse H p51 – regulatory? Gag p6 PR p10 RT p51 p66 IN p32 Anti-HIV drugs Nucleoside/Nucleotide Reverse Transcriptase Inhibitors lamivudine (3TC), zalcitabine (ddC), zidovudine (AZT), didanosine (ddI), stavudine (d4T) Non-Nucleoside Reverse Transcriptase Inhibitors Delavirdine, efavirenz, nevirapine Protease Inhibitors Amprenavir , indinavir, saquinavir, saquinavir, lopinavir/ritonavir, ritonavir, nelfinavir Integrase Inhibitors Isentress (Raltegravir or MK-0518) , JTK303/GS-9137 Fusion or Entry Inhibitors Enfurvitide (Fuzeon or T20), Maraviroc (Selzentry -CCR5 antagonist-) HAART (Highly Active Antiretroviral Therapy): three or more anti-HIV drugs (antiretrovirals) from different classes in combination allows them to work together to keep HIV levels down Gene/Protein env (Pr) 160 env Mass, KDa Function gp160 Env precursor protein SU – Envelope gp120 Surface glycoprotein that binds CD4 TM - Transmembrane gp41 Transmembrane protein that anchors gp120 to virus, responsible for fusion between virus and host cell membrane. Gene/Protein gag (Pr) 55gag Mass, KDa Function p55 Gag precursor protein MA - Matrix p17 Aids nuclear import and viral assembly CA - Capsid p24 HIV central core – contains HIV genome and enzymes NC - Nucleocapsid p15 (p6,p9) p6 – Precise location in virion unknown, not generally present in other retroviruses, may direct proteins to secretory path of cell. p9 – Assoc. with HIV RNA, involved in packaging of HIV RNA into virions pol (Pr) 160gag-pol p160 Gag-Pol precursor protein PR - protease p10 Cleaves Gag and Gag-Pol precursor proteins IN - integrase p31 Integrates viral genome into host DNA RT – reverse transcriptase p66/p51 p66 – copies RNA genome. RNAse H env (Pr) 160 env p51 – regulatory? gp160 Env precursor protein SU – Envelope gp120 Surface glycoprotein that binds CD4 TM - Transmembrane gp41 Transmembrane protein that anchors gp120 to virus, responsible for fusion between virus and host cell membrane. Source: The Molecular Biology of HIV/AIDS, D Ward, pp. 19 Regulatory Proteins: TAT: Trans-Activator of Transcription REV: Regulator of Virion protein expression NEF: Negative Regulatory Factor Accessory Proteins: VIF: Virion Infectivity Factor VPU: Viral Protein U VPR: Viral Protein R TAT: Trans-Activator of Transcription TAR Recruitment of CDK9 Poor transcription Tat Tat Tat Tat Tat CDK9 Release to extracellular medium: Apoptosis bystander T-cells CXCR4 negative interaction Enhanced transcription REV: Regulator of Virion protein expression RRE = rev-response element NEF: Negative Regulatory Factor * Downmodulation the expression of several surface molecules important in host immune function: MHC I, MHC II, CD4 * Activation from latency? Extracellular Nef might activate NF kB. T-cell activation (activates the production of MIP-1alpha and MIP-1beta in macrophages) VIF: Virion Infectivity Factor www.hivmedicine.com/textbook/pathogen.htm VPU: Viral Protein U * Involved in viral budding, enhancing virion release from the cell * Downregulation of CD4 VPR: Viral Protein R * Regulation of nuclear import of the HIV-1 pre-integration complex * Required for virus replication in non-dividing cells such as macrophages. * Induces cell cycle arrest and apoptosis in proliferating cells Gene/Protein tat Tat rev Rev Mass, KDa Function p14 Transactivates HIV transcription (kinase adaptor to RNAP2). p19 Nuclear export of unspliced & single-spliced RNA (via RRE). p27 Internalizes cell-surface CD4 & MHC Class I molecules. nef Nef May enhance cellular activation. vif Vif p23 Overcomes a post-entry block to infection. May influence virion assembly. vpu Vpu p16 Facilitates virion release via ion channel formation. Targets CD4 for destruction in ER (freeing bound env). vpr Vpr p15 Targets the HIV pre-integration complex to the nucleus. Arrests activated cells in G 2 phase of cell cycle. Take-home points: • Structure: – – – – – – – env is only exposed viral protein (neutralization resistant) infectivity mediated by gp120 & gp41 (CD4 & CCR5 or CXCR4) RNA genome -- requires HIV reverse transcription to DNA integration requires HIV integrase LTR/promoter requires cellular transcription factors productive viral gene expression requires HIV protease activity accessory genes modulate: 1) cellular function (e.g., nef, vpr) 2) viral gene expression (e.g., tat, rev) – gag (p24) represents the primary structural component of virion – small molecule inhibitors of these structure’s functional activity represent the primary current antiviral therapeutic strategy