* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download BioSem
Survey
Document related concepts
Transcript
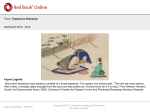
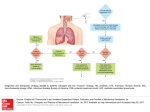
Service-enabling Biomedical Research Enterprise Chapter 5 B. Ramamurthy Page 1 5/22/2017 Introduction • Life sciences have witnessed a flurry of innovations triggered by sequencing of human genome as well as genomes of other genomes. • Area of transformational medicine aims to improve communication between basic and clinical science to allow more therapeutic and diagnostic insights. Page 2 5/22/2017 Translational medicine • From bench to bedside • Exchange ideas, information and knowledge across organizational, governance, sociocultural, political and national boundaries. • Currently mediated by the internet and exponentially-increasing resources • Digital resources: scientific literature, experimental data, curated annotation (metadata) human and machine generated. Ex: Blast Searches NCBI taxonomy Page 3 5/22/2017 Driving principles • • Key requirements: large volume of data to be managed. How? Transform to – – – – – – – Digital Machine readable Capable of being filtered Aggregated Transformed automatically Context information: use and meaning along with content Knowledge integration: combines data from research in mouse genetics, cell bilogy, animal neuropsychology, protein biology, neuropathology, and other areas. – Attention to drug discovery, systems bilogy and personalized medicine that rely heavily on integrating and interpreting data produced by experiments. – Heterogenious data Page 4 5/22/2017 BioSem Enterprise Architecture search Transform results Ex: integrate, generate metadata Dissemination Of results Clinical experiments Ex: drug discovery Diagnostic tools Research Knowledge Ex: Blast Clinical data Ex: JNI Academic Knowledge Ex: cell, psychology molecular Page 5 ontology Treatment methods 5/22/2017 Use case • Parkinson’s disease (PD): – System physiology perspective – Cellular and molecular biology perspective – Pharmacology relating to chemical compounds that bind to receptors – Example query: show me the neuronal components that bind to a ligand which is a therapeutic agent in Parkinson’s disease in reach of the dopaminergic neurons in the substania nigra. – Domain specific shared semantics and classifications – Ontologies can help map among the domains and support seamless integration and interoperation. Page 6 5/22/2017 Development of Ontologies • Manual interaction between ontologists in experts • Textual descriptions are used for adding to this base • Link pre-existing ontologies for extensive coverage Page 7 5/22/2017 Ontology design and creation Approach (fig. 5.1) Subject matter Knowledge (Text) Identify core terms And phrases Map phrases to Relationship between classes Model terms using ontological Constructs: classes, properties Arrange classes and relationships in subsumption hierarchies Information queries Pre-existing classifications And ontologies Page 8 Identify new classes and relationships Re-use classes and relationships Refine subsumption hierarchies Extenf subsumption hierarchies 5/22/2017 Identifying concepts and hierarchies • Text describing PD in p.105 • Study the analysis • Based on the analysis identify important ontological concepts relevant to PD: – – – – Genes Proteins Genetic mutations Diseases • See fig. 5.2 • Next step is to identify relationship among concepts Page 9 5/22/2017 Identifying and extracting relationships rdf:Res ource owl:Thing Gene Dis eas e LewyBody UCHL-1 Parkins onDis eas e Page 10 5/22/2017 Extending the ontology based on information queries • Consider various queries and identify concepts and relationships needed to be part of PD ontology. • These concepts are needed to retrieve information and knowledge from the system. • This lead to additional new concepts. See fig.5.4 Page 11 5/22/2017 PD: adding concepts to support information queries rdfs :Res ource owl:Thing Anatom icalEntity Protein Pathway Page 12 5/22/2017 Ontology Re-use • • • • • • • It is desirable to re-use the ontology and vocabulary developed in the healthcare and life-sciences fields. Diseases: PD information can be used in Huntington’s and Alzeimer’s. PD can reuse information from International classification of diseases ICD and its subset SNOMED. Genes: more genes and genomic concepts such as proteins, pathways are added to ontologies. Consider connecting to Gene Ontology. Neurological concepts: Consider using Neuro names 2007. Enzymes: concepts related to enzymes and other chemicals may be required; you may use Enzyme Nomenclature 2007 Be aware of inconsistencies and circularities. Multiple models may emerge; choice should be based on use cases and functional requirements. Page 13 5/22/2017 Data sources • Now answering the question that we posted in slide#6, three data sources need to be integrated: • Neuron database, PDSP KI database, PubChem Page 14 5/22/2017 Data Integration • A centralized approach where data available through web based interfaces is converted into RDF and stored in a centralized repository • A federated approach where data continues to reside in the existing repositories. RDF mediator converts underlying data into RDF format. • RDF allows for focus on logical structures of information in contrast to only representational format (XML) or storage format (relational). Page 15 5/22/2017 Mapping ontological concepts to RDF graphs • Sample query discussed earlier results in these concepts: – – – – Compartment located_on Neuron Receptor located_in Compartment Ligand binds_to Receptor Ligand associated_with Disease • Next task to map these into RDF maps in the underlying data sources. • Using ontological definitions, data sources, SPARQL queries, and name space, RDF graphs are extracted. Page 16 5/22/2017 Generation and merging of RDF graphs D_Neuron UR12 Neuron Database D1 UR14 Parkinson’s disease UR16 type binds_to associated_with Neuron UR12 D1 UR14 Located_in D_Dendrite UR12 5-H Tryptamine UR15 Located_in PDSPKI Database Page 17 5-H Tryptamine UR15 PubChem database 5/22/2017 Integrated RDF graph Parkinson’s disease UR16 D_Neuron UR12 type associated_with Neuron UR12 5-H Tryptamine UR15 Located_in D_Dendrite UR12 Page 18 binds_to Located_in D1 UR14 5/22/2017 Exam question? • 1. 2. 3. 4. Consider the PD case study that used ontological approach to querying distributed databases. Discuss 10 reasons of using this approach as opposed to common SQL query and relational database approach. Why is Google, Yahoo or MSN search not good enough for searching biological database? Discuss centralized and federated approach to data integration in the context of this case study. Submit a softcopy of the document in the digital drop box. How to do this? Read Chapter 5, read it again. The answers can be formed from the information provided there and from your experience with relational database systems. Page 19 5/22/2017 Summary • Semantic web technologies provide an attractive technological informatics foundation for enabling the Bench to Bedside Vision. • Many areas of biomedical research including drug discovery, systems biology, personalized medicine rely heavily on integrating and interpreting heterogeneous data set. • This is part of ongoing work in the framework of the work being performed in the Healthcare and Life Sciences Interest Group of W3C. Page 20 5/22/2017