* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Struct Bio 101
Survey
Document related concepts
Transcript
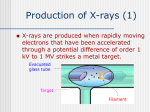
Structural Biology: What does 3D tell us? Stephen J Everse University of Vermont The life of a bio-chemist!! • Training – PhD & Postdoc with Russell F. Doolittle, UCSD • structure of fragment D of fibrinogen • structures of double-D of fibrin – Joined the faculty at UVM in 1998 • Structural biologist (crystallographer) • Current projects – factor Va – thioredoxin reductase – transferrin Everse Group Maria Cristina Bravo Brian Eckenroth, Ph.D. Fundamental Questions How do protein cofactors modulate enzymes? What determines and mediates protein-protein and protein-membrane interactions? How is a protein’s function defined by structure? How does structure prescribe the binding affinity of a metal? Coagulation Cascade Contact Activation Pathway Extrinsic Pathway Factor VIIa Tissue Factor Membrane Ca2+ Factor XIa HMW Kininogen Membrane Ca2+ Zn2+ IXa Factor IXa Factor VIIIa Membrane Ca2+ Prothrombinase Extrinsic Tenase X IX Xa Factor XII Prekallikrein HMW Kininogen “Surface” Factor Xa Factor Va Membrane Ca2+ II IIa “Thrombin” Intrinsic Tenase X Xa IX XI IXa Intrinsic Pathway XIa Relative Rate Prothrombinase of Prothrombin Activation Components “Prothrombinase” Ca2+FXa FVa HC Ca2+ FXa 1 Ca2+FXa 2 Ca2+FXa 20 FVa LC Ca2+ Prothrombin -Thrombin Ca2+FXa FVaCa HC2+ FVa LC W. Gould @2000 300,000 Bovine Factor Vai A3 Cu2+ A1 Ca2+ C1 Funded by: NIH American Society of Hematology C2 Prothrombinase (Va + Xa) A2 A1 A3 C2 C1 Hypothetical model Thioredoxin reductase DmTR Eckenroth et al. Biochemistry 2007 Outline • Determining a 3D structure – X-ray crystallography • Structural elements • Modeling a 3D structure Protein Structures Primary Secondary Tertiary Quaternary Arrangement Alpha helices & of secondary Packing of several polypeptide chains. Beta sheets, elements in 3D space. Loops. Given an amino acid sequence, we are interested in its secondary structures, and how they are arranged in higher structures. Amino acid sequence. Secondary Structure Helix • First predicted by Linus Pauling. Modeled on basis x-ray data which provided accurate geometries, bond lengths, and angles. Modeled before Kendrew’s structure; • 3.6 residues/ turn, 5.4Å/ turn; • The main chain forms a central cylinder with R-groups projecting out; • Variable lengths: from 4 to 40+ residues with the average helix length is 10 residues (3 turns). Secondary Structure The b Sheet • Unlike helix, b sheet composed of secondary structure elements distant in structure; • The b strands are located next to each other • Hydrogen bonds can form between C=O groups of one strand and NH groups of an adjacent strand. • Two different orientations – all strands run same direction: “parallel” – strands in alternating orientation: the “antiparallel”. b-Turns • Type I: Also referred to as a b turn: Hbond between Acyl O of AA1 and NH of AA4; • Type II, glycine must occupy the AA3 position due to steric effects; • Type III is equivalent to 310 helix; • Types I & III constitute some 70% of all b turns; • Proline is typically found in the second position, and most b turns have Asp, Asn, or Gly at the third position. Other Secondary Structural Elements • Random coil • Loop -turn – defined for 3 residues i, i+1, i+2 if a hydrogen bond exists between residues i and i+2 and the phi and psi angles of residue i+1 fall within 40 degrees of one of the following 2 classes turn type classic inverse phi(i+1) psi(i+1) 75.0 -64.0 -79.0 69.0 • Disordered structure Viewing Structures C or CA Ball-and-stick CPK Ribbon and Topology Diagrams Representations of Secondary Structures C -helix b-strand N Tools for Viewing Structures • Jmol – http://jmol.sourceforge.net • PyMOL – http://pymol.sourceforge.net • Swiss PDB viewer – http://www.expasy.ch/spdbv • Mage/KiNG – http://kinemage.biochem.duke.edu/software/mage.php – http://kinemage.biochem.duke.edu/software/king.php • Rasmol – http://www.umass.edu/microbio/rasmol/ RCSB http://www.rcsb.org/ GRASP Graphical Representation and Analysis of Structural Properties Red = negative surface charge Blue = positive surface charge Consurf • The ConSurf server enables the identification of functionally important regions on the surface of a protein or domain, of known three-dimensional (3D) structure, based on the phylogenetic relations between its close sequence homologues; • A multiple sequence alignment (MSA) is used to build a phylogenetic tree consistent with the MSA and calculates conservation scores with either an empirical Bayesian or the Maximum Likelihood method. QuickTime™ and a TIFF (Uncompressed) decompressor are needed to see this picture. http://consurf.tau.ac.il/ Movies QuickTime™ and a PNG decompressor are needed to see this picture. QuickTime™ and a Video decompressor are needed to see this picture. http://pymol.org Proteopedia Higher Level Structures: Motifs & Domains Motif is a simple combination of a few secondary structures, that appear in several different proteins in nature. A collection of motifs forms a domain. Domain is a more complex combination of secondary structures. It has a very specific function (contains an active site). A protein may contain more than one domain. Super-secondary Structures or Motifs Domains "Within a single subunit [polypeptide chain], contiguous portions of the polypeptide chain frequently fold into compact, local semiindependent units called domains." - Richardson, 1981 Domains are: • can be built from structural motifs; • independently folding elements; • functional units; • separable by proteases. Typically, globular proteins are organized into one or more domains. EGF domain from p-selectin Evolutionarily Conserved Domains Often certain structural themes (domains) repeat themselves, but not always in proteins that have similar biological functions. This phenomenon of repeating structures is consistent with the notion that the proteins are genetically related, and that they arose from one another or from a common ancestor. In looking at the amino acid sequences, sometimes there are obvious homologies, and you could predict that the 3-D structures would be similar. But sometimes virtually identical 3-D structures have no sequence similarities at all! Rates of Change • Not all proteins change at the same rate; • Why? • Functional pressures – Surface residues are observed to change most frequently; – Interior less frequently; SequenceStructureFunction Many sequences can give same structure Side chain pattern more important than sequence When homology is high (>50%), likely to have same structure and function (Structural Genomics) Cores conserved Surfaces and loops more variable *3-D shape more conserved than sequence* *There are a limited number of structural frameworks* W. Chazin © 2003 Degree of Evolutionary Conservation Less conserved Information poor DNA seq Protein seq ACAGTTACAC CGGCTATGTA CTATACTTTG HDSFKLPVMS KFDWEMFKPC GKFLDSGKLG S. Lovell © 2002 More conserved Information rich Structure Function How is a 3D structure determined ? 1. Experimental methods (Best approach): • X-rays crystallography - stable fold, good quality crystals. • NMR - stable fold, not suitable for large molecule. 2. In-silico methods (partial solutions based on similarity): • Sequence or profile alignment - uses similar sequences, limited use of 3D information. • Threading - needs 3D structure, combinatorial complexity. • Ab-initio structure prediction - not always successful. Experimental Determination of Atomic Resolution Structures X-ray X-rays Diffraction Pattern NMR RF Resonance RF H0 Direct detection of atom positions Crystals Indirect detection of H-H distances In solution Resolving Power Signal • d • Position Resolving Power: The ability to see two points that are separated by a given distance as distinct Resolution of two points separated by a distance d requires radiation with a wavelength on the order of d or shorter: wavelength Mark Rould © 2007 X-ray Microscopes? nair nair nglass •Lenses require a difference in refractive index between the air and lens material in order to 'bend' and redirect light (or any other form of electromagnetic radiation.) •The refractive index for x-rays is almost exactly 1.00 for all materials. ∆ There are no lenses for xrays. Mark Rould © 2007 Light Scattering and Lenses are Described by Fourier Transforms Scattering = Fourier Transform of specimen Lens applies a second Fourier Transform to the scattered rays to give the image Since X-rays cannot be focused by lenses and refractive index of X-rays in all materials is very close to 1.0 how do we get an atomic image? Mark Rould © 2007 X-ray Diffraction with “The Fourier Duck” The molecule Images by Kevin Cowtan http://www.yorvic.york.ac.uk/~cowtan The diffraction pattern Animal Magic The diffraction pattern Images by Kevin Cowtan http://www.yorvic.york.ac.uk/~cowtan The CAT (molecule) Solution: Measure Scattered Rays, Use Fourier Transform to Mimic Lens Transforms Computer X-Ray Detector Mark Rould © 2007 A Problem… A single molecule is a very weak scatterer of X-rays. Most of the X-rays will pass through the molecule without being diffracted. Those rays which are diffracted are too weak to be detected. Solution: Analyzing diffraction from crystals instead of single molecules. A crystal is made of a three-dimensional repeat of ordered molecules (1014) whose signals reinforce each other. The resulting diffracted rays are strong enough to be detected. A Crystal • • • 3D repeating lattice; Unit cell is the smallest unit of the lattice; Come in all shapes and sizes. Sylvie Doublié © 2000 Crystals come from slowly precipitating the biological molecule out of solution under conditions that will not damage or denature it (sometimes). Putting it all together: X-ray diffraction Electron density map QuickTime™ and a TIFF (Uncompressed) decompressor are needed to see this picture. Rubisco diffraction pattern Crystallographer Detector Computer Scattered rays Object X-rays Diffraction pattern is a collection of diffraction spots (reflections) Sylvie Doublié © 2000 Model What information does structure give you? 3-D view of macromolecules at near atomic resolution. The result of a successful structural project is a “structure” or model of the macromolecule in the crystal. You can assign: - secondary structure elements - position and conformation of side chains - position of ligands, inhibitors, metals etc. A model allows you: - to understand biochemical and genetic data (i.e., structural basis of functional changes in mutant or modified macromolecule). - generate hypotheses regarding the roles of particular residues or domains Sylvie Doublié © 2000 What did I just say????!!! • A structure is a “MODEL”!! • What does that mean? – It is someone’s interpretation of the primary data!!! So what happens when we can’t get an NMR or X-ray structure? 2˚ & 3˚ Structure Prediction Secondary (2o) Structure Table 10 Phi & Psi angles for Regular Secondary Structure Conformations Structure Antiparallel b-sheet Parallel b-Sheet Right-handed -helix 310 helix p helix Polyproline I Polyproline II Polyglycine II Phi (F) -139 -119 -+64 -49 -57 -83 -78 -80 Psi(Y) +135 +113 +40 -26 -70 +158 +149 +150 Secondary Structure Prediction • One of the first fields to emerge in bioinformatics (~1967) • Grew from a simple observation that certain amino acids or combinations of amino acids seemed to prefer to be in certain secondary structures • Subject of hundreds of papers and dozens of books, many methods… Simplified C-F Algorithm • Select a window of 7 residues • Calculate average P over this window and assign that value to the central residue • Repeat the calculation for Pb and Pc • Slide the window down one residue and repeat until sequence is complete • Analyze resulting “plot” and assign secondary structure (H, B, C) for each residue to highest value Limitations of Chou-Fasman • Does not take into account long range information (>3 residues away) • Does not take into account sequence content or probable structure class • Assumes simple additive probability (not true in nature) • Does not include related sequences or alignments in prediction process • Only about 55% accurate (on good days) Protein Principles • Proteins reflect millions of years of evolution. • Most proteins belong to large evolutionary families. • 3D structure is better conserved than sequence during evolution. • Similarities between sequences or between structures may reveal information about shared biological functions of a protein family. The PhD Algorithm • Search the SWISS-PROT database and select high scoring homologues • Create a sequence “profile” from the resulting multiple alignment • Include global sequence info in the profile • Input the profile into a trained two-layer neural network to predict the structure and to “clean-up” the prediction http://www.predictprotein.org/ PHD ZHANG GOR III JASEP7 PTIT LEVIN LIM GOR I CF Scores (%) Prediction Performance 75 70 65 60 55 50 45 Best of the Best • PredictProtein-PHD (72%) – http://www.predictprotein.org/ • Jpred (73-75%) – http://www.compbio.dundee.ac.uk/wwwjpred/index.html • SAM-T08 (75%) – http://compbio.soe.ucsc.edu/SAM_T08/T08query.html • PSIpred (77%) – http://bioinf.cs.ucl.ac.uk/psipred/psiform.html Structure Prediction • Threading • A protein fold recognition technique that involves incrementally replacing the sequence of a known protein structure with a query sequence of unknown structure. • Why threading? • Secondary structure is more conserved than primary structure • Tertiary structure is more conserved TH than secondary structure R E A D An Approach SAS Calculations • DSSP - Database of Secondary Structures for Proteins – http://swift.cmbi.ru.nl/gv/start/index.html • VADAR - Volume Area Dihedral Angle Reporter – http://redpoll.pharmacy.ualberta.ca/vadar/ • GetArea – http://curie.utmb.edu/getarea.html • Naccess - Atomic Solvent Accessible Area Calculations – http://www.bioinf.msnchester.ac.uk/naccess 3D Threading Servers Generate 3D models or coordinates of possible models based on input sequence • PredictProtein-PHDacc – http://www.predictprotein.org • PredAcc – http://mobyle.rpbs.univ-paris-diderot.fr/cgibin/portal.py?form=PredAcc • Loopp (version 2) – http://cbsuapps.tc.cornell.edu/loopp.aspx • Phyre – http://www.sbg.bio.ic.ac.uk/~phyre/ • SwissModel – http://swissmodel.expasy.org/ • All require email addresses since the process may take hours to complete Ab Initio Folding • Two Central Problems – Sampling conformational space (10100) – The energy minimum problem • The Sampling Problem (Solutions) – Lattice models, off-lattice models, simplified chain methods, parallelism • The Energy Problem (Solutions) – Threading energies, packing assessment, topology assessment Lattice Folding http://folding.stanford.edu/ For the gamers out there… http://fold.it/portal/ Print & Online Resources Crystallography Made Crystal Clear, by Gale Rhodes http://www.usm.maine.edu/~rhodes/CMCC/index.html http://ruppweb.dyndns.org/Xray/101index.html Online tutorial with interactive applets and quizzes. http://www.ysbl.york.ac.uk/~cowtan/fourier/fourier.html Nice pictures demonstrating Fourier transforms http://ucxray.berkeley.edu/~jamesh/movies/ Cool movies demonstrating key points about diffraction, resolution, data quality, and refinement. http://www-structmed.cimr.cam.ac.uk/course.html Notes from a macromolecular crystallography course taught in Cambridge