* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download organic crystals: prediction of crystal structure from molecular structure
X-ray crystallography wikipedia , lookup
Pseudo Jahn–Teller effect wikipedia , lookup
Energy harvesting wikipedia , lookup
Geometrical frustration wikipedia , lookup
Metastable inner-shell molecular state wikipedia , lookup
Eigenstate thermalization hypothesis wikipedia , lookup
Colloidal crystal wikipedia , lookup
Energy applications of nanotechnology wikipedia , lookup
Crystal structure wikipedia , lookup
Heat transfer physics wikipedia , lookup
Jahn–Teller effect wikipedia , lookup
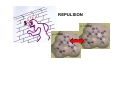
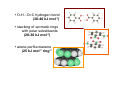
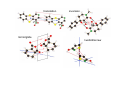
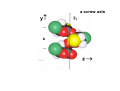
the crystal packing of organic molecules intermolecular energies determine crystal symmetry and intermolecular geometry angelo gavezzotti Dipartimento di Chimica Strutturale Università di Milano [email protected] p1 p2 pg pm p3 p4 pgg Part I. The physical nature and the computer simulation of the intermolecular potential a "modern" molecule is a collection of nuclei and of charge density points (~20000 significant ones for a medium-size molecule) the charge density box - a molecular orbital calculation - computer programs available - standard option for ρ(x,y,z) in e- per cubic angstrom V(i), ρ(i); q(i)= V(i) ρ(i) • In principle: The total intermolecular interaction energy is the expectation value of the Hamiltonian of a supramolecular system (just like the molecular energy is the expectation value of the molecular Hamiltonian) •In practice: for historical and practical reasons, one usually defines: - Coulombic terms - polarization terms - dispersion terms - repulsion terms stabilizing/destabilizing: ENERGIES < 0, > 0 attractive/repulsive: FORCES interactions can be: stabilizing and attractive stabilizing but repulsive destabilizing but attractive (activation) destabilizing and repulsive 5 4 3 2 1 0 -1 -2 3 3.5 4 4.5 dis tanza in angs tro m 5 Coulombic term ++, - - repulsive destabilizing + - attractive stabilizing strongly orientation-dependent + + + - + - + + - Coulombic potential energy • Point-charge model: a net charge at each atomic nucleus In terms of point-charge models, the coulombic energy is Ecoul =1/(4πε°) ΣΣ qi,A q j,B /Rij where do the q's come from? 1) Mulliken population analysis charges (variable with basis set) 2) ESP charges (more consistent) 3) re-scaled Extended Huckel (Gavezzotti and Filippini) 4) other recipes In terms of central multipoles The potential due to a charge distribution, at a distance R from the center of charges, is expanded in a series of multipoles: V(R) = A°/R +(1/R2) (Ax px+Ay py+Az pz) + (1/R3) (Az2 d z2+Axy dxy+Axz dxz+Ayz dyz+ Ax2-y2 dx2-y2) + higher terms over 1/R4 , 1/R5 ,... A coefficients: radial dependence of the potential p and d functions are the spherical harmonics. Central multipoles may be appropriate for very small molecules, but not for large organic molecules The distributed dipole model (A.J.Stone) distributed multipoles point charges? negative nuclei? • In terms of central or distributed dipoles, the coulombic energy is a sum of moment k to moment m terms, where each moment is a monopole, dipole, quadrupole moment. Coulombic potential energy: in terms of full density Ecoul = ∫ ∫ρA ((r1)) ρB ((r2) )/ |r1 -r2 | d 3r1d 3r2 + Σk ∫ Zk(A) ρB ((r2) )/ |rk -r2 | d 3r2 + + Σm ∫ Zm(B) ρA ((r1) )/ |rm -r1 | d 3r1 + Σk Σm Zk(A) Zm(B) |rm -rk | requires an analytical form for ρ Ecoul = Σk Σm qk,A ((r1)) qm,B ((r2) )/ |r1 -r2 | + Σk Zk(A) [Σm qm,B ((rm) )/ |rk -rm | ] + + Σk Zk(B) [Σm qm,A ((rm) )/ |rk -rm | ] + Σk Σm Zk(A) Zm(B) |rm -rk | replaces the integrals with a summation, ρ by points POLARIZATION attractive stabilizing + polarizer + + polarized Polarization (electrostatic induction) energy Polarizability is: a measure of the propensity of a given electronic environment to yield under the action of an external electric field : displacement/restraint the restraining force is coulombic attraction between the displaced charge and the nuclei polarizability is large when electrons are at a large distance from a weakly charged nucleus. electric field electron nucleus The energy involved in the polarization process is always stabilizing because the induced dipole µ always points in the stabilizing direction (compliance of the polarized medium to the polarizing field ε) The linear dipole polarization energy is given by E pol = - ∫ µ dε = -1/2 α ε2 classical polarization: analogy with molecular orbitals mixing, including virtual orbitals sometimes called charge-transfer energy. the electron game of chemistry.................. bond formation spin pairing charge transfer polarization Pauli repulsion avoidance of like spins NEUTRAL NEUTRAL ?? DISPERSION: conceptually similar to polarization it operates in any chemical system from NaCl to solid benzene the "glue of the world"!!! Dispersion energy fluctuating dipoles generate induced dipoles in a nearby electron distribution. The resulting quantum mechanical dipole-dipole coupling is the effect called dispersion. stabilization has to do with zero-point oscillator energies, and dispersion effects are a consequence of the uncertainty principle, like all zero-point energy effects. London model for dispersion energy α molecular polarizability, I ionization potential at a distance R: Edisp ≈ [ -I (α)2 ] (R)-6 The dispersion energy is always stabilizing, like polarization energy is. Fritz London REPULSION Exchange-overlap-Pauli repulsion Short-range repulsion has no classical analogy. Pauli principle:no two electrons with same quantum numbers in the same region of space mathematical requirement: proper wavefunctions must be antisymmetrical Approximate models for the repulsion energy: SAB = electron density overlap Erep = A SABγ Pauli: avoid overlap reasonable: energy proportional to overlap, with A and γ parameters Etot = Ecoul + Epol + Edisp + Erep chemical interpretations; for example, coulombic end polarization energies are large in molecules with a permanent charge separation the definition of partitioned energies is not unique. partitioning schemes: Morokuma-Stone IMPT-etc... One often speaks of "van der Waals" energies more appropriately one should speak of Coulomb-London-Pauli energies Etot = Ecoul + Epol + Edisp + Erep • full quantum chemical methods approximate methods: • the 'atom-atom' potential E = A exp(-BRkm) - C(Rkm)6 E = A (Rkm)-12 - B (Rkm)-6 • semiempirical methods ('PIXEL') building a model for calculating interaction energies a molecule at X' = MX°+ t rotation and translation the electric field a molecule at X° if M and t are space group operations, calculate lattice energy inspecting the nature of the intermolecular chemical bond exercise parallel offset benzene dimer 80 Ecoul Epol energies, kJ/mol 60 Edisp 40 Erep 20 Etot 0 -20 -40 -60 -80 2 3 4 5 interplanar distance 6 benzoic acid dimer 300 E(coul) E(pol) 200 energies, kJ/mol E(disp) E(rep) 100 E(tot), n=3 0 -100 -200 -300 1 2 3 R(O...H) 4 The energetic glossary of intermolecular interactions very strong interactions in charged species +H NCH CH=CHCOO3 2 , energies in kJ/mol - + + -325 -178 + +83 - + - • O-H...O=C hydrogen bond (30-40 kJ mol-1) • stacking of aromatic rings with polar substituents (20-30 kJ mol-1) • arene-perfluoroarene (25 kJ mol-1 ring-1) • very acidic C-H to π-clouds (10 kJ mol-1) acetylene-acetone • very acidic C-H to basic O, N (<10 kJ mol-1) benzoquinone • aliphatic C-H to O,N,π (<5 kJ mol-1) Part II Molecular organization in crystals Calculation of crystal energies CRYSTAL: condensed phase in which matter is organized with periodic translational symmetry chemical entities are related by symmetry that repeats itself periodically in 3D space the basic chemical unit an inversion center translation inversion mirror/glide twofold/screw P1 or P1space groups inversion center plus translation a screw axis 21 __ b __ z→ a screw axis plus perpendicular translation: monoclinic __ ↑ b ↓__ ← c → inversion center + screw axis = glide plane 21 →G →G exercise: space group P21/c b c basic facts about molecular crystals molecular materials ▪ non-linear optics ▪ pigments ▪ explosives ▪ pharmaceutical dosage forms ▪ mesophases (LCD displays) molecular materials vs. inorganics, ceramics, etc. ▪ organic chemistry much more versatile than inorganic chemistry ▪ less weight ▪ more pollution (solvents, etc.) ▪ very little mechanical resistence sublimation enthalpy, kJ/mole... organic crystals sublimation heat, kJ/mol 250 200 150 100 50 0 0 50 100 150 200 250 molecular weight 300 350 400 problem: a quantitative estimate of the lattice energy solution: 1) give the computer the co-ordinates of the asymmetric unit, the cell dimensions and space group; these come from X-ray diffraction 2) generate a cluster of molecules and calculate the lattice sums alternative solution: periodicize atomic orbitals (band structure) and calculate the electronic wavefunction on crystal orbitals (Bloch methods) Φ(r+R, k) = exp(ik·R) Φ(r,k) k=a*/2 k=0 Bloch function 160 calculated lattice energy 140 a 10 kJ/mol difference between obs and calc is considered a good result (experimental values often wrong by much more than that) 120 100 80 60 40 20 20 40 60 80 100 120 sublimation enthalpy 140 160 Ecoul Epol Edisp Erep Etot argon -2 -1 -12 8 -7 DTE -21 -9 -123 60 -93 SUCROSE -273 -126 -116 315 -200 Ecoul NaCl > -800 Epol Edisp Erep -108 -29 169 -9 254 forsterite (one ion pair) [2Mg]4+[SiO4]4- -8156 -1009 weak forces require close contact of molecular objects: close packing in organic crystals organic crystals: prediction of crystal structure from molecular structure • generate compact structures (geometry) • compute lattice energies (physics of the interaction) • ranking generated structures in order of stability (a problem in polymorphism) ...a largely unsolved problem! close packing is a necessary but not a sufficient condition anhydrous caffeine computational crystal structures -40 Coul. -60 -80 total -100 -120 -140 1.35 disp. 1.4 1.45 1.5 1.55 1.6 density many good crystal structures, all close packed... ...and yet no one has ever been able to have a good crystal of anhydrous caffeine! for any given molecular structure, many crystal structures can always be derived by a simple simulation experimental ∆H's between polymorphs are a few kJ/mol about 10 generated structures have lattice energies withing a range of a few kJ mol-1 crystallization is not only a matter of equilibrium thermodynamics, otherwise one would always crystallize a Boltzmann distribution of many polymorphs aspirin, naphthalene and sucrose have been crystallized in tons without ever observing a second polymorph "soft matter" great flexibility in phase behaviour the good side: many options the bad side: difficult to control a slab of crystalline succinic anhydride ?? liquid the problem.... kinetics