* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Student project proposal
Survey
Document related concepts
Transcript
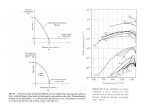
15.10.15 Student project proposal Submitted by: Nadia Opara PhD student at C-CINA (group of Prof. Henning Stahlberg, Dr. Thomas Braun) e-mail: [email protected] Topic: High quality sample of protein nanocrystals from nanoliter volumes for electron diffraction data collection. Main aim of the project – establish reproducible conditions for protein nanocrystal delivery using minimal sample volumes. Fast and lossless sample delivery system to the electron microscope (EM) grid is a prerequisite for optimal exploitation of biological material - usually available in limited amounts - in structural analysis. Additionally, advent of the new generation of high-speed cameras allows collection diffraction data of high resolution. In the view of recent advances in structural analysis well-established path for nanocrystal protein sample preparation is of the high importance. Trehalose embedding technique was reported as a successful method to preserve 2D protein crystals especially for high-resolution transmission electron microscopy (TEM) studies [1, 2]. Recently it was proven that, this method can be also adapted for adequately small (sufficiently thin) 3D protein crystals for electron diffraction data collection (Fig. 1) [3]. Nanoliter volumes of protein crystals solution initially sugar embedded were deposited on EM grid. A) B) Fig. 1 A) Nanocrystals deposited on TEM grid, B) Electron diffraction pattern of lysozyme The lysozyme protein was used as model system since is readily forming 3D crystals in broad spectrum of chemical conditions. Final dimensions of the grown protein crystal depend on physical and biochemical system parameters like: protein concentration, pH, temperature or concentration of precipitat [4]. However, unlike lysozyme, many biologically important proteins are difficult to obtain in large quantities and forms very small crystals (nanocrystals) useless for other methods exploited in structural biology. 1 Protein crystals in size (thickness) optimal for diffraction and tomography studies (100-200 nm) can be obtained by crystallization experiment using vapor diffusion method on either standard sitting drop plate with 96 wells or silicon chip with anisotropically etched cavities (Fig. 2). Both systems are compatible with nanovolume dispensing unit together with dew point stage developed at C-CINA [3], which allows deposition at low temperatures limiting evaporation of the low-volume sample. Fig. 2 4x4 array of wells on silicon chip for protein crystallization. Main advantage of pristine silicon single crystal material is its very good heat conductivity. Protein crystallization chambers made of silicon can be used for tests, which require conditions of different temperatures. Application of the thermal cyclic changes can lead to melting and regrowth of the crystalline material and improve crystal quality [5]. Nonetheless, in this method protein material is prone to clustering in the edges of the cavities (Fig. 2). During realization of the project evaluation of different geometries of protein growth – sitting drop plate geometry vs silicon chip with cavities using selected physico-chemical conditions will lead to determination parameters for growth the well-defined protein crystal size diffracting to the high resolution. Available at C-CINA electron microscopy equipment will be used to examine of the quality of the delivered nanocrystalline material. References: [1] The use of trehalose in the preparation of specimens for molecular electron microscopy, Chiu et al., Micron 42, 762-772 (2011) [2] Low-dose electron diffraction of catalase crystals dried with a matrix of the disaccharide, trehalose: Is a “dry protein crystallography” feasible?, W. H. Massover, Microsc. Microanal. 8 (Suppl.2) (2002) [3] S. Arnold et al., Electron microscopy sample preparation from nanoliter volumes – poster presented at SNI annual meeting, (September 2015) [4] Generation of Size-Controlled, Submicrometer Protein Crystals, Falkner et al, Chem. Mater., 17, 2679-2686 (2005) [5] Growth of crystals in optical tweezers, U. Gibson et al, Proc. SPIE 5930, Optical Trapping and Optical Micromanipulation II, 593014 (2005) 2